Tianbin Li
Can Large Language Models Function as Qualified Pediatricians? A Systematic Evaluation in Real-World Clinical Contexts
Nov 17, 2025Abstract:With the rapid rise of large language models (LLMs) in medicine, a key question is whether they can function as competent pediatricians in real-world clinical settings. We developed PEDIASBench, a systematic evaluation framework centered on a knowledge-system framework and tailored to realistic clinical environments. PEDIASBench assesses LLMs across three dimensions: application of basic knowledge, dynamic diagnosis and treatment capability, and pediatric medical safety and medical ethics. We evaluated 12 representative models released over the past two years, including GPT-4o, Qwen3-235B-A22B, and DeepSeek-V3, covering 19 pediatric subspecialties and 211 prototypical diseases. State-of-the-art models performed well on foundational knowledge, with Qwen3-235B-A22B achieving over 90% accuracy on licensing-level questions, but performance declined ~15% as task complexity increased, revealing limitations in complex reasoning. Multiple-choice assessments highlighted weaknesses in integrative reasoning and knowledge recall. In dynamic diagnosis and treatment scenarios, DeepSeek-R1 scored highest in case reasoning (mean 0.58), yet most models struggled to adapt to real-time patient changes. On pediatric medical ethics and safety tasks, Qwen2.5-72B performed best (accuracy 92.05%), though humanistic sensitivity remained limited. These findings indicate that pediatric LLMs are constrained by limited dynamic decision-making and underdeveloped humanistic care. Future development should focus on multimodal integration and a clinical feedback-model iteration loop to enhance safety, interpretability, and human-AI collaboration. While current LLMs cannot independently perform pediatric care, they hold promise for decision support, medical education, and patient communication, laying the groundwork for a safe, trustworthy, and collaborative intelligent pediatric healthcare system.
TCM-5CEval: Extended Deep Evaluation Benchmark for LLM's Comprehensive Clinical Research Competence in Traditional Chinese Medicine
Nov 17, 2025Abstract:Large language models (LLMs) have demonstrated exceptional capabilities in general domains, yet their application in highly specialized and culturally-rich fields like Traditional Chinese Medicine (TCM) requires rigorous and nuanced evaluation. Building upon prior foundational work such as TCM-3CEval, which highlighted systemic knowledge gaps and the importance of cultural-contextual alignment, we introduce TCM-5CEval, a more granular and comprehensive benchmark. TCM-5CEval is designed to assess LLMs across five critical dimensions: (1) Core Knowledge (TCM-Exam), (2) Classical Literacy (TCM-LitQA), (3) Clinical Decision-making (TCM-MRCD), (4) Chinese Materia Medica (TCM-CMM), and (5) Clinical Non-pharmacological Therapy (TCM-ClinNPT). We conducted a thorough evaluation of fifteen prominent LLMs, revealing significant performance disparities and identifying top-performing models like deepseek\_r1 and gemini\_2\_5\_pro. Our findings show that while models exhibit proficiency in recalling foundational knowledge, they struggle with the interpretative complexities of classical texts. Critically, permutation-based consistency testing reveals widespread fragilities in model inference. All evaluated models, including the highest-scoring ones, displayed a substantial performance degradation when faced with varied question option ordering, indicating a pervasive sensitivity to positional bias and a lack of robust understanding. TCM-5CEval not only provides a more detailed diagnostic tool for LLM capabilities in TCM but aldso exposes fundamental weaknesses in their reasoning stability. To promote further research and standardized comparison, TCM-5CEval has been uploaded to the Medbench platform, joining its predecessor in the "In-depth Challenge for Comprehensive TCM Abilities" special track.
A Survey of Scientific Large Language Models: From Data Foundations to Agent Frontiers
Aug 28, 2025



Abstract:Scientific Large Language Models (Sci-LLMs) are transforming how knowledge is represented, integrated, and applied in scientific research, yet their progress is shaped by the complex nature of scientific data. This survey presents a comprehensive, data-centric synthesis that reframes the development of Sci-LLMs as a co-evolution between models and their underlying data substrate. We formulate a unified taxonomy of scientific data and a hierarchical model of scientific knowledge, emphasizing the multimodal, cross-scale, and domain-specific challenges that differentiate scientific corpora from general natural language processing datasets. We systematically review recent Sci-LLMs, from general-purpose foundations to specialized models across diverse scientific disciplines, alongside an extensive analysis of over 270 pre-/post-training datasets, showing why Sci-LLMs pose distinct demands -- heterogeneous, multi-scale, uncertainty-laden corpora that require representations preserving domain invariance and enabling cross-modal reasoning. On evaluation, we examine over 190 benchmark datasets and trace a shift from static exams toward process- and discovery-oriented assessments with advanced evaluation protocols. These data-centric analyses highlight persistent issues in scientific data development and discuss emerging solutions involving semi-automated annotation pipelines and expert validation. Finally, we outline a paradigm shift toward closed-loop systems where autonomous agents based on Sci-LLMs actively experiment, validate, and contribute to a living, evolving knowledge base. Collectively, this work provides a roadmap for building trustworthy, continually evolving artificial intelligence (AI) systems that function as a true partner in accelerating scientific discovery.
S2-UniSeg: Fast Universal Agglomerative Pooling for Scalable Segment Anything without Supervision
Aug 09, 2025Abstract:Recent self-supervised image segmentation models have achieved promising performance on semantic segmentation and class-agnostic instance segmentation. However, their pretraining schedule is multi-stage, requiring a time-consuming pseudo-masks generation process between each training epoch. This time-consuming offline process not only makes it difficult to scale with training dataset size, but also leads to sub-optimal solutions due to its discontinuous optimization routine. To solve these, we first present a novel pseudo-mask algorithm, Fast Universal Agglomerative Pooling (UniAP). Each layer of UniAP can identify groups of similar nodes in parallel, allowing to generate both semantic-level and instance-level and multi-granular pseudo-masks within ens of milliseconds for one image. Based on the fast UniAP, we propose the Scalable Self-Supervised Universal Segmentation (S2-UniSeg), which employs a student and a momentum teacher for continuous pretraining. A novel segmentation-oriented pretext task, Query-wise Self-Distillation (QuerySD), is proposed to pretrain S2-UniSeg to learn the local-to-global correspondences. Under the same setting, S2-UniSeg outperforms the SOTA UnSAM model, achieving notable improvements of AP+6.9 on COCO, AR+11.1 on UVO, PixelAcc+4.5 on COCOStuff-27, RQ+8.0 on Cityscapes. After scaling up to a larger 2M-image subset of SA-1B, S2-UniSeg further achieves performance gains on all four benchmarks. Our code and pretrained models are available at https://github.com/bio-mlhui/S2-UniSeg
MedITok: A Unified Tokenizer for Medical Image Synthesis and Interpretation
May 25, 2025Abstract:Advanced autoregressive models have reshaped multimodal AI. However, their transformative potential in medical imaging remains largely untapped due to the absence of a unified visual tokenizer -- one capable of capturing fine-grained visual structures for faithful image reconstruction and realistic image synthesis, as well as rich semantics for accurate diagnosis and image interpretation. To this end, we present MedITok, the first unified tokenizer tailored for medical images, encoding both low-level structural details and high-level clinical semantics within a unified latent space. To balance these competing objectives, we introduce a novel two-stage training framework: a visual representation alignment stage that cold-starts the tokenizer reconstruction learning with a visual semantic constraint, followed by a textual semantic representation alignment stage that infuses detailed clinical semantics into the latent space. Trained on the meticulously collected large-scale dataset with over 30 million medical images and 2 million image-caption pairs, MedITok achieves state-of-the-art performance on more than 30 datasets across 9 imaging modalities and 4 different tasks. By providing a unified token space for autoregressive modeling, MedITok supports a wide range of tasks in clinical diagnostics and generative healthcare applications. Model and code will be made publicly available at: https://github.com/Masaaki-75/meditok.
Building a Human-Verified Clinical Reasoning Dataset via a Human LLM Hybrid Pipeline for Trustworthy Medical AI
May 11, 2025Abstract:Despite strong performance in medical question-answering, the clinical adoption of Large Language Models (LLMs) is critically hampered by their opaque 'black-box' reasoning, limiting clinician trust. This challenge is compounded by the predominant reliance of current medical LLMs on corpora from scientific literature or synthetic data, which often lack the granular expert validation and high clinical relevance essential for advancing their specialized medical capabilities. To address these critical gaps, we introduce a highly clinically relevant dataset with 31,247 medical question-answer pairs, each accompanied by expert-validated chain-of-thought (CoT) explanations. This resource, spanning multiple clinical domains, was curated via a scalable human-LLM hybrid pipeline: LLM-generated rationales were iteratively reviewed, scored, and refined by medical experts against a structured rubric, with substandard outputs revised through human effort or guided LLM regeneration until expert consensus. This publicly available dataset provides a vital source for the development of medical LLMs that capable of transparent and verifiable reasoning, thereby advancing safer and more interpretable AI in medicine.
GMAI-VL-R1: Harnessing Reinforcement Learning for Multimodal Medical Reasoning
Apr 02, 2025Abstract:Recent advances in general medical AI have made significant strides, but existing models often lack the reasoning capabilities needed for complex medical decision-making. This paper presents GMAI-VL-R1, a multimodal medical reasoning model enhanced by reinforcement learning (RL) to improve its reasoning abilities. Through iterative training, GMAI-VL-R1 optimizes decision-making, significantly boosting diagnostic accuracy and clinical support. We also develop a reasoning data synthesis method, generating step-by-step reasoning data via rejection sampling, which further enhances the model's generalization. Experimental results show that after RL training, GMAI-VL-R1 excels in tasks such as medical image diagnosis and visual question answering. While the model demonstrates basic memorization with supervised fine-tuning, RL is crucial for true generalization. Our work establishes new evaluation benchmarks and paves the way for future advancements in medical reasoning models. Code, data, and model will be released at \href{https://github.com/uni-medical/GMAI-VL-R1}{this link}.
Towards Interpretable Counterfactual Generation via Multimodal Autoregression
Mar 29, 2025Abstract:Counterfactual medical image generation enables clinicians to explore clinical hypotheses, such as predicting disease progression, facilitating their decision-making. While existing methods can generate visually plausible images from disease progression prompts, they produce silent predictions that lack interpretation to verify how the generation reflects the hypothesized progression -- a critical gap for medical applications that require traceable reasoning. In this paper, we propose Interpretable Counterfactual Generation (ICG), a novel task requiring the joint generation of counterfactual images that reflect the clinical hypothesis and interpretation texts that outline the visual changes induced by the hypothesis. To enable ICG, we present ICG-CXR, the first dataset pairing longitudinal medical images with hypothetical progression prompts and textual interpretations. We further introduce ProgEmu, an autoregressive model that unifies the generation of counterfactual images and textual interpretations. We demonstrate the superiority of ProgEmu in generating progression-aligned counterfactuals and interpretations, showing significant potential in enhancing clinical decision support and medical education. Project page: https://progemu.github.io.
OphCLIP: Hierarchical Retrieval-Augmented Learning for Ophthalmic Surgical Video-Language Pretraining
Nov 23, 2024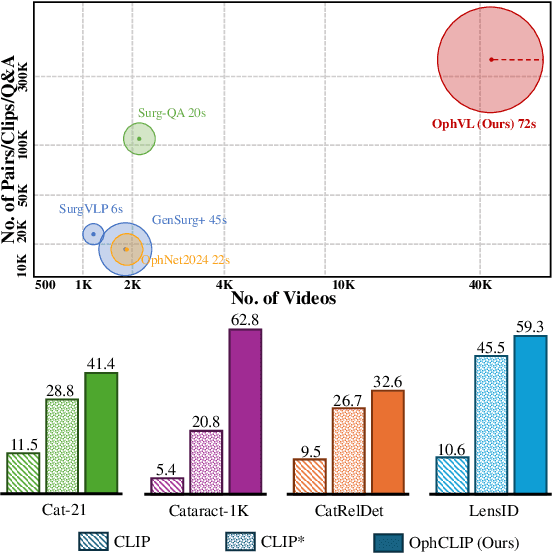
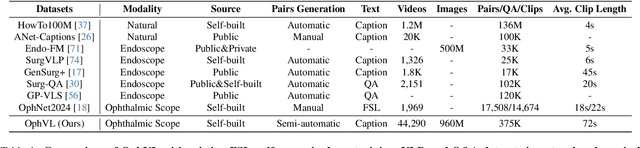
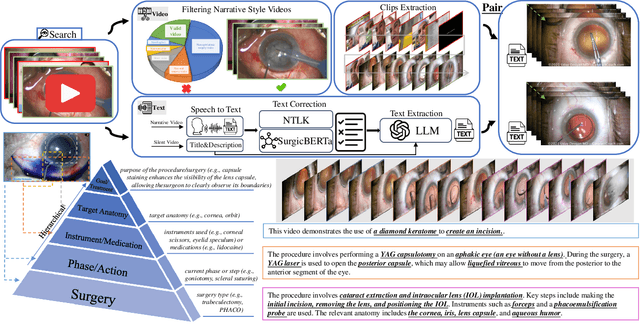
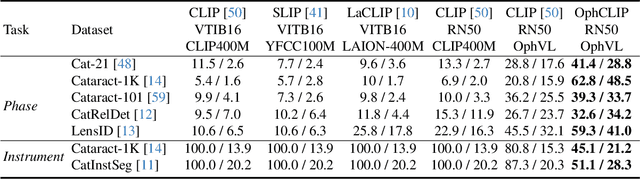
Abstract:Surgical practice involves complex visual interpretation, procedural skills, and advanced medical knowledge, making surgical vision-language pretraining (VLP) particularly challenging due to this complexity and the limited availability of annotated data. To address the gap, we propose OphCLIP, a hierarchical retrieval-augmented vision-language pretraining framework specifically designed for ophthalmic surgical workflow understanding. OphCLIP leverages the OphVL dataset we constructed, a large-scale and comprehensive collection of over 375K hierarchically structured video-text pairs with tens of thousands of different combinations of attributes (surgeries, phases/operations/actions, instruments, medications, as well as more advanced aspects like the causes of eye diseases, surgical objectives, and postoperative recovery recommendations, etc). These hierarchical video-text correspondences enable OphCLIP to learn both fine-grained and long-term visual representations by aligning short video clips with detailed narrative descriptions and full videos with structured titles, capturing intricate surgical details and high-level procedural insights, respectively. Our OphCLIP also designs a retrieval-augmented pretraining framework to leverage the underexplored large-scale silent surgical procedure videos, automatically retrieving semantically relevant content to enhance the representation learning of narrative videos. Evaluation across 11 datasets for phase recognition and multi-instrument identification shows OphCLIP's robust generalization and superior performance.
GMAI-VL & GMAI-VL-5.5M: A Large Vision-Language Model and A Comprehensive Multimodal Dataset Towards General Medical AI
Nov 21, 2024Abstract:Despite significant advancements in general artificial intelligence, such as GPT-4, their effectiveness in the medical domain (general medical AI, GMAI) remains constrained due to the absence of specialized medical knowledge. To address this challenge, we present GMAI-VL-5.5M, a comprehensive multimodal medical dataset created by converting hundreds of specialized medical datasets into meticulously constructed image-text pairs. This dataset features comprehensive task coverage, diverse modalities, and high-quality image-text data. Building upon this multimodal dataset, we propose GMAI-VL, a general medical vision-language model with a progressively three-stage training strategy. This approach significantly enhances the model's ability by integrating visual and textual information, thereby improving its ability to process multimodal data and support accurate diagnosis and clinical decision-making. Experimental evaluations demonstrate that GMAI-VL achieves state-of-the-art results across a wide range of multimodal medical tasks, such as visual question answering and medical image diagnosis. Our contributions include the development of the GMAI-VL-5.5M dataset, the introduction of the GMAI-VL model, and the establishment of new benchmarks in multiple medical domains. Code and dataset will be released at https://github.com/uni-medical/GMAI-VL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge