Yu Guang Wang
Linear Causal Representation Learning by Topological Ordering, Pruning, and Disentanglement
Sep 26, 2025Abstract:Causal representation learning (CRL) has garnered increasing interests from the causal inference and artificial intelligence community, due to its capability of disentangling potentially complex data-generating mechanism into causally interpretable latent features, by leveraging the heterogeneity of modern datasets. In this paper, we further contribute to the CRL literature, by focusing on the stylized linear structural causal model over the latent features and assuming a linear mixing function that maps latent features to the observed data or measurements. Existing linear CRL methods often rely on stringent assumptions, such as accessibility to single-node interventional data or restrictive distributional constraints on latent features and exogenous measurement noise. However, these prerequisites can be challenging to satisfy in certain scenarios. In this work, we propose a novel linear CRL algorithm that, unlike most existing linear CRL methods, operates under weaker assumptions about environment heterogeneity and data-generating distributions while still recovering latent causal features up to an equivalence class. We further validate our new algorithm via synthetic experiments and an interpretability analysis of large language models (LLMs), demonstrating both its superiority over competing methods in finite samples and its potential in integrating causality into AI.
How Particle System Theory Enhances Hypergraph Message Passing
May 24, 2025Abstract:Hypergraphs effectively model higher-order relationships in natural phenomena, capturing complex interactions beyond pairwise connections. We introduce a novel hypergraph message passing framework inspired by interacting particle systems, where hyperedges act as fields inducing shared node dynamics. By incorporating attraction, repulsion, and Allen-Cahn forcing terms, particles of varying classes and features achieve class-dependent equilibrium, enabling separability through the particle-driven message passing. We investigate both first-order and second-order particle system equations for modeling these dynamics, which mitigate over-smoothing and heterophily thus can capture complete interactions. The more stable second-order system permits deeper message passing. Furthermore, we enhance deterministic message passing with stochastic element to account for interaction uncertainties. We prove theoretically that our approach mitigates over-smoothing by maintaining a positive lower bound on the hypergraph Dirichlet energy during propagation and thus to enable hypergraph message passing to go deep. Empirically, our models demonstrate competitive performance on diverse real-world hypergraph node classification tasks, excelling on both homophilic and heterophilic datasets.
Score-matching-based Structure Learning for Temporal Data on Networks
Dec 10, 2024



Abstract:Causal discovery is a crucial initial step in establishing causality from empirical data and background knowledge. Numerous algorithms have been developed for this purpose. Among them, the score-matching method has demonstrated superior performance across various evaluation metrics, particularly for the commonly encountered Additive Nonlinear Causal Models. However, current score-matching-based algorithms are primarily designed to analyze independent and identically distributed (i.i.d.) data. More importantly, they suffer from high computational complexity due to the pruning step required for handling dense Directed Acyclic Graphs (DAGs). To enhance the scalability of score matching, we have developed a new parent-finding subroutine for leaf nodes in DAGs, significantly accelerating the most time-consuming part of the process: the pruning step. This improvement results in an efficiency-lifted score matching algorithm, termed Parent Identification-based Causal structure learning for both i.i.d. and temporal data on networKs, or PICK. The new score-matching algorithm extends the scope of existing algorithms and can handle static and temporal data on networks with weak network interference. Our proposed algorithm can efficiently cope with increasingly complex datasets that exhibit spatial and temporal dependencies, commonly encountered in academia and industry. The proposed algorithm can accelerate score-matching-based methods while maintaining high accuracy in real-world applications.
A Survey for Large Language Models in Biomedicine
Aug 29, 2024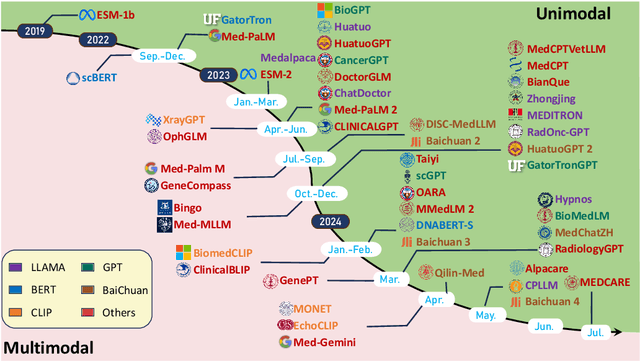
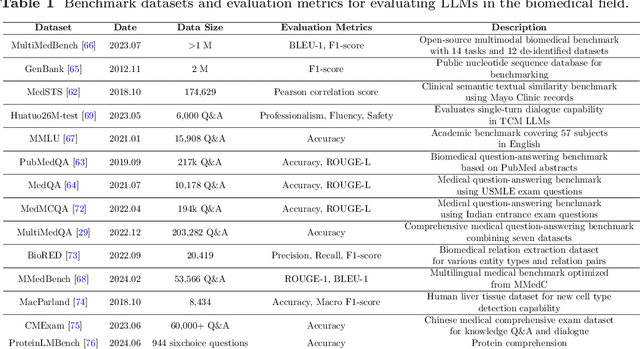
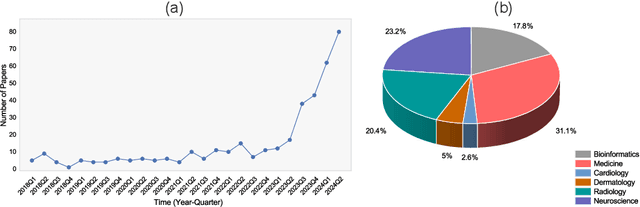
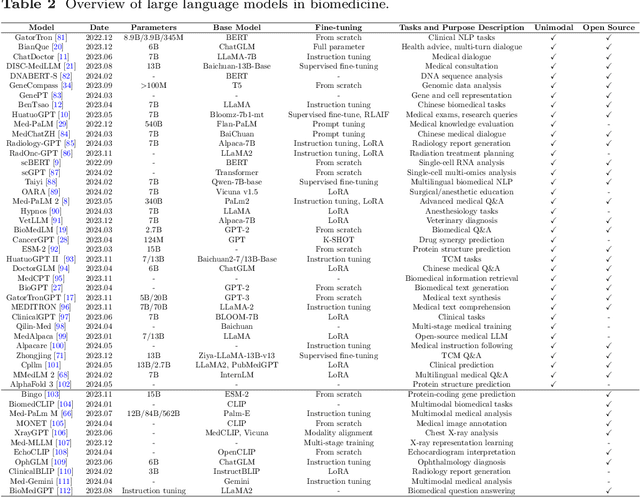
Abstract:Recent breakthroughs in large language models (LLMs) offer unprecedented natural language understanding and generation capabilities. However, existing surveys on LLMs in biomedicine often focus on specific applications or model architectures, lacking a comprehensive analysis that integrates the latest advancements across various biomedical domains. This review, based on an analysis of 484 publications sourced from databases including PubMed, Web of Science, and arXiv, provides an in-depth examination of the current landscape, applications, challenges, and prospects of LLMs in biomedicine, distinguishing itself by focusing on the practical implications of these models in real-world biomedical contexts. Firstly, we explore the capabilities of LLMs in zero-shot learning across a broad spectrum of biomedical tasks, including diagnostic assistance, drug discovery, and personalized medicine, among others, with insights drawn from 137 key studies. Then, we discuss adaptation strategies of LLMs, including fine-tuning methods for both uni-modal and multi-modal LLMs to enhance their performance in specialized biomedical contexts where zero-shot fails to achieve, such as medical question answering and efficient processing of biomedical literature. Finally, we discuss the challenges that LLMs face in the biomedicine domain including data privacy concerns, limited model interpretability, issues with dataset quality, and ethics due to the sensitive nature of biomedical data, the need for highly reliable model outputs, and the ethical implications of deploying AI in healthcare. To address these challenges, we also identify future research directions of LLM in biomedicine including federated learning methods to preserve data privacy and integrating explainable AI methodologies to enhance the transparency of LLMs.
TourSynbio: A Multi-Modal Large Model and Agent Framework to Bridge Text and Protein Sequences for Protein Engineering
Aug 27, 2024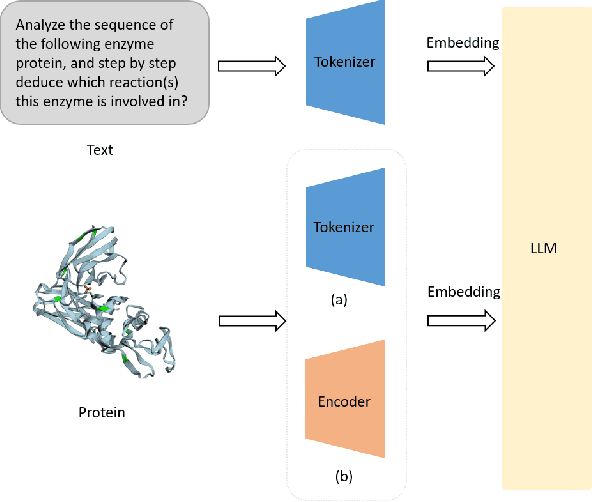
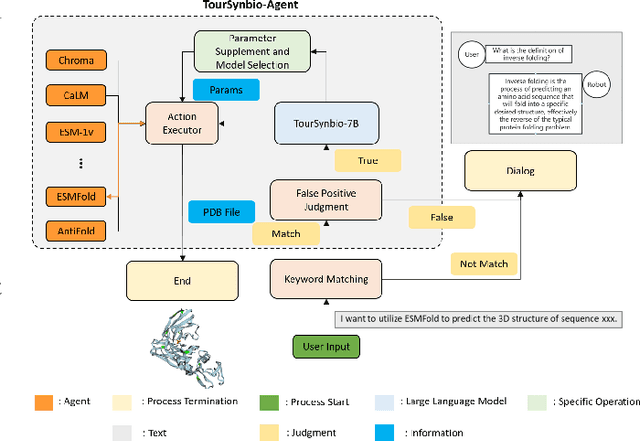
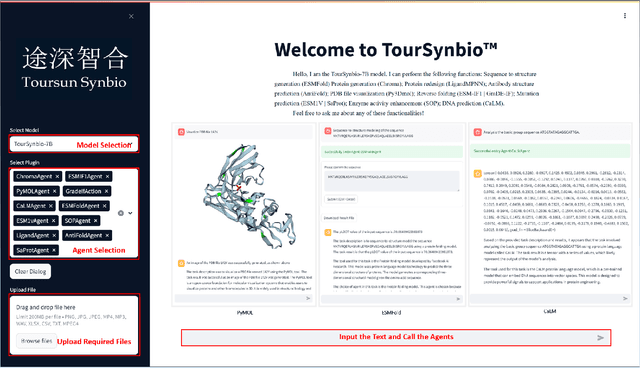
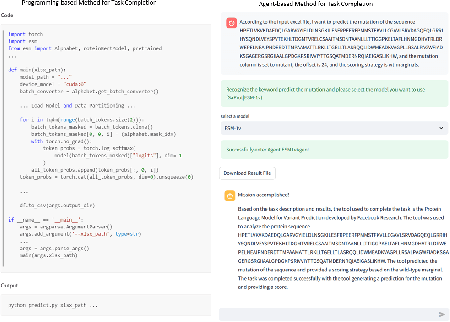
Abstract:The structural similarities between protein sequences and natural languages have led to parallel advancements in deep learning across both domains. While large language models (LLMs) have achieved much progress in the domain of natural language processing, their potential in protein engineering remains largely unexplored. Previous approaches have equipped LLMs with protein understanding capabilities by incorporating external protein encoders, but this fails to fully leverage the inherent similarities between protein sequences and natural languages, resulting in sub-optimal performance and increased model complexity. To address this gap, we present TourSynbio-7B, the first multi-modal large model specifically designed for protein engineering tasks without external protein encoders. TourSynbio-7B demonstrates that LLMs can inherently learn to understand proteins as language. The model is post-trained and instruction fine-tuned on InternLM2-7B using ProteinLMDataset, a dataset comprising 17.46 billion tokens of text and protein sequence for self-supervised pretraining and 893K instructions for supervised fine-tuning. TourSynbio-7B outperforms GPT-4 on the ProteinLMBench, a benchmark of 944 manually verified multiple-choice questions, with 62.18% accuracy. Leveraging TourSynbio-7B's enhanced protein sequence understanding capability, we introduce TourSynbio-Agent, an innovative framework capable of performing various protein engineering tasks, including mutation analysis, inverse folding, protein folding, and visualization. TourSynbio-Agent integrates previously disconnected deep learning models in the protein engineering domain, offering a unified conversational user interface for improved usability. Finally, we demonstrate the efficacy of TourSynbio-7B and TourSynbio-Agent through two wet lab case studies on vanilla key enzyme modification and steroid compound catalysis.
Improving Antibody Design with Force-Guided Sampling in Diffusion Models
Jun 09, 2024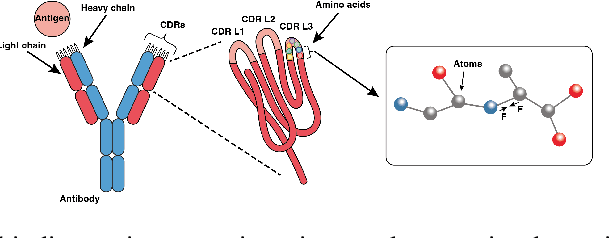
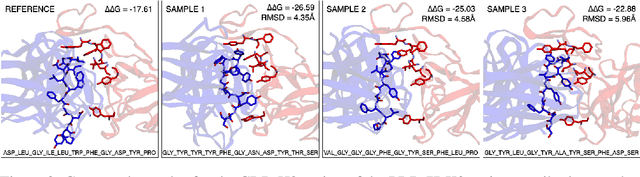
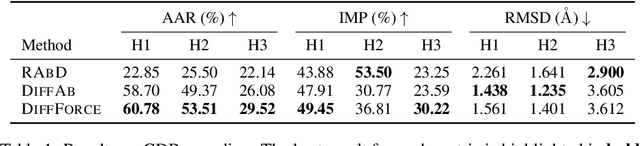
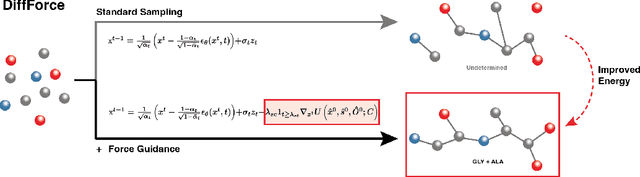

Abstract:Antibodies, crucial for immune defense, primarily rely on complementarity-determining regions (CDRs) to bind and neutralize antigens, such as viruses. The design of these CDRs determines the antibody's affinity and specificity towards its target. Generative models, particularly denoising diffusion probabilistic models (DDPMs), have shown potential to advance the structure-based design of CDR regions. However, only a limited dataset of bound antibody-antigen structures is available, and generalization to out-of-distribution interfaces remains a challenge. Physics based force-fields, which approximate atomic interactions, offer a coarse but universal source of information to better mold designs to target interfaces. Integrating this foundational information into diffusion models is, therefore, highly desirable. Here, we propose a novel approach to enhance the sampling process of diffusion models by integrating force field energy-based feedback. Our model, DiffForce, employs forces to guide the diffusion sampling process, effectively blending the two distributions. Through extensive experiments, we demonstrate that our method guides the model to sample CDRs with lower energy, enhancing both the structure and sequence of the generated antibodies.
A Fine-tuning Dataset and Benchmark for Large Language Models for Protein Understanding
Jun 08, 2024Abstract:The parallels between protein sequences and natural language in their sequential structures have inspired the application of large language models (LLMs) to protein understanding. Despite the success of LLMs in NLP, their effectiveness in comprehending protein sequences remains an open question, largely due to the absence of datasets linking protein sequences to descriptive text. Researchers have then attempted to adapt LLMs for protein understanding by integrating a protein sequence encoder with a pre-trained LLM. However, this adaptation raises a fundamental question: "Can LLMs, originally designed for NLP, effectively comprehend protein sequences as a form of language?" Current datasets fall short in addressing this question due to the lack of a direct correlation between protein sequences and corresponding text descriptions, limiting the ability to train and evaluate LLMs for protein understanding effectively. To bridge this gap, we introduce ProteinLMDataset, a dataset specifically designed for further self-supervised pretraining and supervised fine-tuning (SFT) of LLMs to enhance their capability for protein sequence comprehension. Specifically, ProteinLMDataset includes 17.46 billion tokens for pretraining and 893,000 instructions for SFT. Additionally, we present ProteinLMBench, the first benchmark dataset consisting of 944 manually verified multiple-choice questions for assessing the protein understanding capabilities of LLMs. ProteinLMBench incorporates protein-related details and sequences in multiple languages, establishing a new standard for evaluating LLMs' abilities in protein comprehension. The large language model InternLM2-7B, pretrained and fine-tuned on the ProteinLMDataset, outperforms GPT-4 on ProteinLMBench, achieving the highest accuracy score. The dataset and the benchmark are available at https://huggingface.co/datasets/tsynbio/ProteinLMBench.
How Universal Polynomial Bases Enhance Spectral Graph Neural Networks: Heterophily, Over-smoothing, and Over-squashing
May 21, 2024Abstract:Spectral Graph Neural Networks (GNNs), alternatively known as graph filters, have gained increasing prevalence for heterophily graphs. Optimal graph filters rely on Laplacian eigendecomposition for Fourier transform. In an attempt to avert prohibitive computations, numerous polynomial filters have been proposed. However, polynomials in the majority of these filters are predefined and remain fixed across different graphs, failing to accommodate the varying degrees of heterophily. Addressing this gap, we demystify the intrinsic correlation between the spectral property of desired polynomial bases and the heterophily degrees via thorough theoretical analyses. Subsequently, we develop a novel adaptive heterophily basis wherein the basis vectors mutually form angles reflecting the heterophily degree of the graph. We integrate this heterophily basis with the homophily basis to construct a universal polynomial basis UniBasis, which devises a polynomial filter based graph neural network - UniFilter. It optimizes the convolution and propagation in GNN, thus effectively limiting over-smoothing and alleviating over-squashing. Our extensive experiments, conducted on a diverse range of real-world and synthetic datasets with varying degrees of heterophily, support the superiority of UniFilter. These results not only demonstrate the universality of UniBasis but also highlight its proficiency in graph explanation.
Layer-diverse Negative Sampling for Graph Neural Networks
Mar 18, 2024



Abstract:Graph neural networks (GNNs) are a powerful solution for various structure learning applications due to their strong representation capabilities for graph data. However, traditional GNNs, relying on message-passing mechanisms that gather information exclusively from first-order neighbours (known as positive samples), can lead to issues such as over-smoothing and over-squashing. To mitigate these issues, we propose a layer-diverse negative sampling method for message-passing propagation. This method employs a sampling matrix within a determinantal point process, which transforms the candidate set into a space and selectively samples from this space to generate negative samples. To further enhance the diversity of the negative samples during each forward pass, we develop a space-squeezing method to achieve layer-wise diversity in multi-layer GNNs. Experiments on various real-world graph datasets demonstrate the effectiveness of our approach in improving the diversity of negative samples and overall learning performance. Moreover, adding negative samples dynamically changes the graph's topology, thus with the strong potential to improve the expressiveness of GNNs and reduce the risk of over-squashing.
Dirichlet Energy Enhancement of Graph Neural Networks by Framelet Augmentation
Nov 09, 2023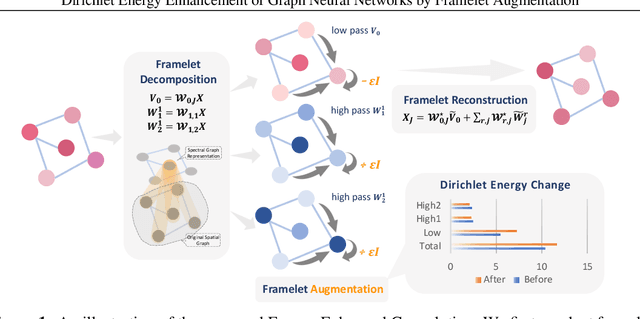
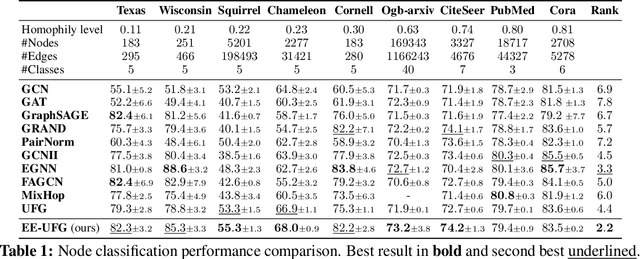
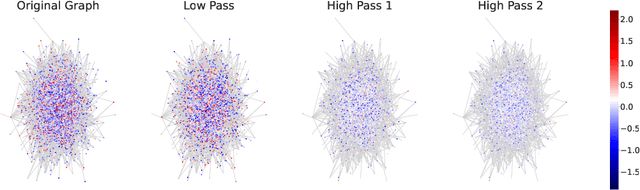
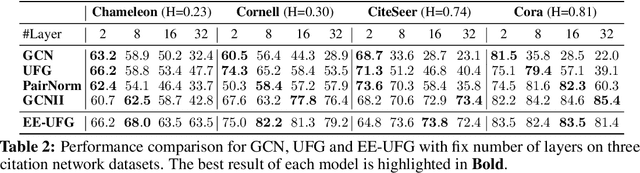
Abstract:Graph convolutions have been a pivotal element in learning graph representations. However, recursively aggregating neighboring information with graph convolutions leads to indistinguishable node features in deep layers, which is known as the over-smoothing issue. The performance of graph neural networks decays fast as the number of stacked layers increases, and the Dirichlet energy associated with the graph decreases to zero as well. In this work, we introduce a framelet system into the analysis of Dirichlet energy and take a multi-scale perspective to leverage the Dirichlet energy and alleviate the over-smoothing issue. Specifically, we develop a Framelet Augmentation strategy by adjusting the update rules with positive and negative increments for low-pass and high-passes respectively. Based on that, we design the Energy Enhanced Convolution (EEConv), which is an effective and practical operation that is proved to strictly enhance Dirichlet energy. From a message-passing perspective, EEConv inherits multi-hop aggregation property from the framelet transform and takes into account all hops in the multi-scale representation, which benefits the node classification tasks over heterophilous graphs. Experiments show that deep GNNs with EEConv achieve state-of-the-art performance over various node classification datasets, especially for heterophilous graphs, while also lifting the Dirichlet energy as the network goes deeper.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge