Shixiang Tang
Owl-AuraID 1.0: An Intelligent System for Autonomous Scientific Instrumentation and Scientific Data Analysis
Mar 31, 2026Abstract:Scientific discovery increasingly depends on high-throughput characterization, yet automation is hindered by proprietary GUIs and the limited generalizability of existing API-based systems. We present Owl-AuraID, a software-hardware collaborative embodied agent system that adopts a GUI-native paradigm to operate instruments through the same interfaces as human experts. Its skill-centric framework integrates Type-1 (GUI operation) and Type-2 (data analysis) skills into end-to-end workflows, connecting physical sample handling with scientific interpretation. Owl-AuraID demonstrates broad coverage across ten categories of precision instruments and diverse workflows, including multimodal spectral analysis, microscopic imaging, and crystallographic analysis, supporting modalities such as FTIR, NMR, AFM, and TGA. Overall, Owl-AuraID provides a practical, extensible foundation for autonomous laboratories and illustrates a path toward evolving laboratory intelligence through reusable operational and analytical skills. The code are available at https://github.com/OpenOwlab/AuraID.
Project Imaging-X: A Survey of 1000+ Open-Access Medical Imaging Datasets for Foundation Model Development
Mar 29, 2026Abstract:Foundation models have demonstrated remarkable success across diverse domains and tasks, primarily due to the thrive of large-scale, diverse, and high-quality datasets. However, in the field of medical imaging, the curation and assembling of such medical datasets are highly challenging due to the reliance on clinical expertise and strict ethical and privacy constraints, resulting in a scarcity of large-scale unified medical datasets and hindering the development of powerful medical foundation models. In this work, we present the largest survey to date of medical image datasets, covering over 1,000 open-access datasets with a systematic catalog of their modalities, tasks, anatomies, annotations, limitations, and potential for integration. Our analysis exposes a landscape that is modest in scale, fragmented across narrowly scoped tasks, and unevenly distributed across organs and modalities, which in turn limits the utility of existing medical image datasets for developing versatile and robust medical foundation models. To turn fragmentation into scale, we propose a metadata-driven fusion paradigm (MDFP) that integrates public datasets with shared modalities or tasks, thereby transforming multiple small data silos into larger, more coherent resources. Building on MDFP, we release an interactive discovery portal that enables end-to-end, automated medical image dataset integration, and compile all surveyed datasets into a unified, structured table that clearly summarizes their key characteristics and provides reference links, offering the community an accessible and comprehensive repository. By charting the current terrain and offering a principled path to dataset consolidation, our survey provides a practical roadmap for scaling medical imaging corpora, supporting faster data discovery, more principled dataset creation, and more capable medical foundation models.
Intern-S1-Pro: Scientific Multimodal Foundation Model at Trillion Scale
Mar 26, 2026Abstract:We introduce Intern-S1-Pro, the first one-trillion-parameter scientific multimodal foundation model. Scaling to this unprecedented size, the model delivers a comprehensive enhancement across both general and scientific domains. Beyond stronger reasoning and image-text understanding capabilities, its intelligence is augmented with advanced agent capabilities. Simultaneously, its scientific expertise has been vastly expanded to master over 100 specialized tasks across critical science fields, including chemistry, materials, life sciences, and earth sciences. Achieving this massive scale is made possible by the robust infrastructure support of XTuner and LMDeploy, which facilitates highly efficient Reinforcement Learning (RL) training at the 1-trillion parameter level while ensuring strict precision consistency between training and inference. By seamlessly integrating these advancements, Intern-S1-Pro further fortifies the fusion of general and specialized intelligence, working as a Specializable Generalist, demonstrating its position in the top tier of open-source models for general capabilities, while outperforming proprietary models in the depth of specialized scientific tasks.
Charting Empirical Laws for LLM Fine-Tuning in Scientific Multi-Discipline Learning
Feb 11, 2026Abstract:While large language models (LLMs) have achieved strong performance through fine-tuning within individual scientific domains, their learning dynamics in multi-disciplinary contexts remains poorly understood, despite the promise of improved generalization and broader applicability through cross-domain knowledge synergy. In this work, we present the first systematic study of multi-disciplinary LLM fine-tuning, constructing a five-discipline corpus and analyzing learning patterns of full fine-tuning, LoRA, LoRA-MoE, and LoRA compositions. Particularly, our study shows that multi-disciplinary learning is substantially more variable than single-discipline training and distills four consistent empirical laws: (1) Balance-then-Diversity: low-resource disciplines degrade performance unless mitigated via diversity-aware upsampling; (2) Merge-then-Align: restoring instruction-following ability is critical for cross-discipline synergy; (3) Optimize-then-Scale: parameter scaling offers limited gains without prior design optimization; and (4) Share-then-Specialize: asymmetric LoRA-MoE yields robust gains with minimal trainable parameters via shared low-rank projection. Together, these laws form a practical recipe for principled multi-discipline fine-tuning and provide actionable guidance for developing generalizable scientific LLMs.
SciIF: Benchmarking Scientific Instruction Following Towards Rigorous Scientific Intelligence
Jan 08, 2026Abstract:As large language models (LLMs) transition from general knowledge retrieval to complex scientific discovery, their evaluation standards must also incorporate the rigorous norms of scientific inquiry. Existing benchmarks exhibit a critical blind spot: general instruction-following metrics focus on superficial formatting, while domain-specific scientific benchmarks assess only final-answer correctness, often rewarding models that arrive at the right result with the wrong reasons. To address this gap, we introduce scientific instruction following: the capability to solve problems while strictly adhering to the constraints that establish scientific validity. Specifically, we introduce SciIF, a multi-discipline benchmark that evaluates this capability by pairing university-level problems with a fixed catalog of constraints across three pillars: scientific conditions (e.g., boundary checks and assumptions), semantic stability (e.g., unit and symbol conventions), and specific processes(e.g., required numerical methods). Uniquely, SciIF emphasizes auditability, requiring models to provide explicit evidence of constraint satisfaction rather than implicit compliance. By measuring both solution correctness and multi-constraint adherence, SciIF enables finegrained diagnosis of compositional reasoning failures, ensuring that LLMs can function as reliable agents within the strict logical frameworks of science.
SciEvalKit: An Open-source Evaluation Toolkit for Scientific General Intelligence
Dec 30, 2025Abstract:We introduce SciEvalKit, a unified benchmarking toolkit designed to evaluate AI models for science across a broad range of scientific disciplines and task capabilities. Unlike general-purpose evaluation platforms, SciEvalKit focuses on the core competencies of scientific intelligence, including Scientific Multimodal Perception, Scientific Multimodal Reasoning, Scientific Multimodal Understanding, Scientific Symbolic Reasoning, Scientific Code Generation, Science Hypothesis Generation and Scientific Knowledge Understanding. It supports six major scientific domains, spanning from physics and chemistry to astronomy and materials science. SciEvalKit builds a foundation of expert-grade scientific benchmarks, curated from real-world, domain-specific datasets, ensuring that tasks reflect authentic scientific challenges. The toolkit features a flexible, extensible evaluation pipeline that enables batch evaluation across models and datasets, supports custom model and dataset integration, and provides transparent, reproducible, and comparable results. By bridging capability-based evaluation and disciplinary diversity, SciEvalKit offers a standardized yet customizable infrastructure to benchmark the next generation of scientific foundation models and intelligent agents. The toolkit is open-sourced and actively maintained to foster community-driven development and progress in AI4Science.
Omni-Weather: Unified Multimodal Foundation Model for Weather Generation and Understanding
Dec 25, 2025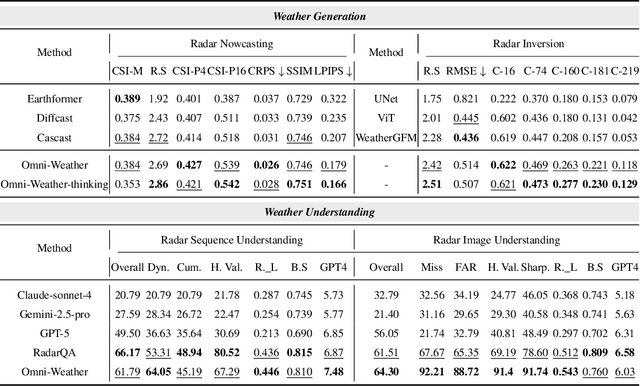
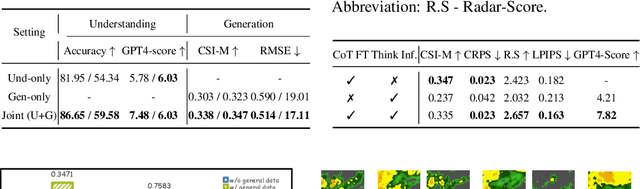
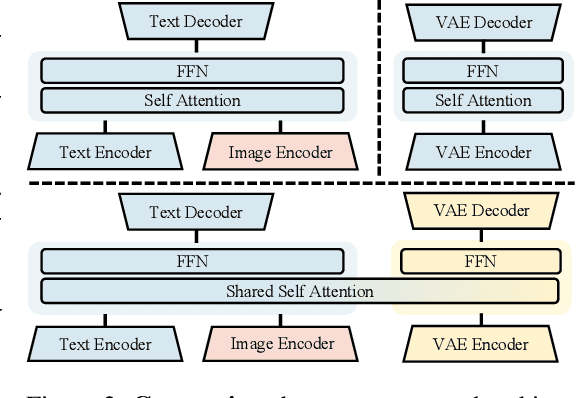
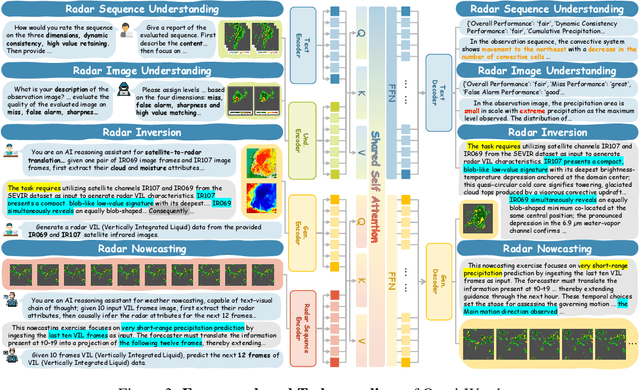

Abstract:Weather modeling requires both accurate prediction and mechanistic interpretation, yet existing methods treat these goals in isolation, separating generation from understanding. To address this gap, we present Omni-Weather, the first multimodal foundation model that unifies weather generation and understanding within a single architecture. Omni-Weather integrates a radar encoder for weather generation tasks, followed by unified processing using a shared self-attention mechanism. Moreover, we construct a Chain-of-Thought dataset for causal reasoning in weather generation, enabling interpretable outputs and improved perceptual quality. Extensive experiments show Omni-Weather achieves state-of-the-art performance in both weather generation and understanding. Our findings further indicate that generative and understanding tasks in the weather domain can mutually enhance each other. Omni-Weather also demonstrates the feasibility and value of unifying weather generation and understanding.
Probing Scientific General Intelligence of LLMs with Scientist-Aligned Workflows
Dec 18, 2025Abstract:Despite advances in scientific AI, a coherent framework for Scientific General Intelligence (SGI)-the ability to autonomously conceive, investigate, and reason across scientific domains-remains lacking. We present an operational SGI definition grounded in the Practical Inquiry Model (PIM: Deliberation, Conception, Action, Perception) and operationalize it via four scientist-aligned tasks: deep research, idea generation, dry/wet experiments, and experimental reasoning. SGI-Bench comprises over 1,000 expert-curated, cross-disciplinary samples inspired by Science's 125 Big Questions, enabling systematic evaluation of state-of-the-art LLMs. Results reveal gaps: low exact match (10--20%) in deep research despite step-level alignment; ideas lacking feasibility and detail; high code executability but low execution result accuracy in dry experiments; low sequence fidelity in wet protocols; and persistent multimodal comparative-reasoning challenges. We further introduce Test-Time Reinforcement Learning (TTRL), which optimizes retrieval-augmented novelty rewards at inference, enhancing hypothesis novelty without reference answer. Together, our PIM-grounded definition, workflow-centric benchmark, and empirical insights establish a foundation for AI systems that genuinely participate in scientific discovery.
Visionary: The World Model Carrier Built on WebGPU-Powered Gaussian Splatting Platform
Dec 09, 2025Abstract:Neural rendering, particularly 3D Gaussian Splatting (3DGS), has evolved rapidly and become a key component for building world models. However, existing viewer solutions remain fragmented, heavy, or constrained by legacy pipelines, resulting in high deployment friction and limited support for dynamic content and generative models. In this work, we present Visionary, an open, web-native platform for real-time various Gaussian Splatting and meshes rendering. Built on an efficient WebGPU renderer with per-frame ONNX inference, Visionary enables dynamic neural processing while maintaining a lightweight, "click-to-run" browser experience. It introduces a standardized Gaussian Generator contract, which not only supports standard 3DGS rendering but also allows plug-and-play algorithms to generate or update Gaussians each frame. Such inference also enables us to apply feedforward generative post-processing. The platform further offers a plug in three.js library with a concise TypeScript API for seamless integration into existing web applications. Experiments show that, under identical 3DGS assets, Visionary achieves superior rendering efficiency compared to current Web viewers due to GPU-based primitive sorting. It already supports multiple variants, including MLP-based 3DGS, 4DGS, neural avatars, and style transformation or enhancement networks. By unifying inference and rendering directly in the browser, Visionary significantly lowers the barrier to reproduction, comparison, and deployment of 3DGS-family methods, serving as a unified World Model Carrier for both reconstructive and generative paradigms.
ARCHE: A Novel Task to Evaluate LLMs on Latent Reasoning Chain Extraction
Nov 16, 2025



Abstract:Large language models (LLMs) are increasingly used in scientific domains. While they can produce reasoning-like content via methods such as chain-of-thought prompting, these outputs are typically unstructured and informal, obscuring whether models truly understand the fundamental reasoning paradigms that underpin scientific inference. To address this, we introduce a novel task named Latent Reasoning Chain Extraction (ARCHE), in which models must decompose complex reasoning arguments into combinations of standard reasoning paradigms in the form of a Reasoning Logic Tree (RLT). In RLT, all reasoning steps are explicitly categorized as one of three variants of Peirce's fundamental inference modes: deduction, induction, or abduction. To facilitate this task, we release ARCHE Bench, a new benchmark derived from 70 Nature Communications articles, including more than 1,900 references and 38,000 viewpoints. We propose two logic-aware evaluation metrics: Entity Coverage (EC) for content completeness and Reasoning Edge Accuracy (REA) for step-by-step logical validity. Evaluations on 10 leading LLMs on ARCHE Bench reveal that models exhibit a trade-off between REA and EC, and none are yet able to extract a complete and standard reasoning chain. These findings highlight a substantial gap between the abilities of current reasoning models and the rigor required for scientific argumentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge