Daguang Xu
Surg$Σ$: A Spectrum of Large-Scale Multimodal Data and Foundation Models for Surgical Intelligence
Mar 17, 2026Abstract:Surgical intelligence has the potential to improve the safety and consistency of surgical care, yet most existing surgical AI frameworks remain task-specific and struggle to generalize across procedures and institutions. Although multimodal foundation models, particularly multimodal large language models, have demonstrated strong cross-task capabilities across various medical domains, their advancement in surgery remains constrained by the lack of large-scale, systematically curated multimodal data. To address this challenge, we introduce Surg$Σ$, a spectrum of large-scale multimodal data and foundation models for surgical intelligence. At the core of this framework lies Surg$Σ$-DB, a large-scale multimodal data foundation designed to support diverse surgical tasks. Surg$Σ$-DB consolidates heterogeneous surgical data sources (including open-source datasets, curated in-house clinical collections and web-source data) into a unified schema, aiming to improve label consistency and data standardization across heterogeneous datasets. Surg$Σ$-DB spans 6 clinical specialties and diverse surgical types, providing rich image- and video-level annotations across 18 practical surgical tasks covering understanding, reasoning, planning, and generation, at an unprecedented scale (over 5.98M conversations). Beyond conventional multimodal conversations, Surg$Σ$-DB incorporates hierarchical reasoning annotations, providing richer semantic cues to support deeper contextual understanding in complex surgical scenarios. We further provide empirical evidence through recently developed surgical foundation models built upon Surg$Σ$-DB, illustrating the practical benefits of large-scale multimodal annotations, unified semantic design, and structured reasoning annotations for improving cross-task generalization and interpretability.
Generalized Recognition of Basic Surgical Actions Enables Skill Assessment and Vision-Language-Model-based Surgical Planning
Mar 13, 2026Abstract:Artificial intelligence, imaging, and large language models have the potential to transform surgical practice, training, and automation. Understanding and modeling of basic surgical actions (BSA), the fundamental unit of operation in any surgery, is important to drive the evolution of this field. In this paper, we present a BSA dataset comprising 10 basic actions across 6 surgical specialties with over 11,000 video clips, which is the largest to date. Based on the BSA dataset, we developed a new foundation model that conducts general-purpose recognition of basic actions. Our approach demonstrates robust cross-specialist performance in experiments validated on datasets from different procedural types and various body parts. Furthermore, we demonstrate downstream applications enabled by the BAS foundation model through surgical skill assessment in prostatectomy using domain-specific knowledge, and action planning in cholecystectomy and nephrectomy using large vision-language models. Multinational surgeons' evaluation of the language model's output of the action planning explainable texts demonstrated clinical relevance. These findings indicate that basic surgical actions can be robustly recognized across scenarios, and an accurate BSA understanding model can essentially facilitate complex applications and speed up the realization of surgical superintelligence.
SAW: Toward a Surgical Action World Model via Controllable and Scalable Video Generation
Mar 13, 2026Abstract:A surgical world model capable of generating realistic surgical action videos with precise control over tool-tissue interactions can address fundamental challenges in surgical AI and simulation -- from data scarcity and rare event synthesis to bridging the sim-to-real gap for surgical automation. However, current video generation methods, the very core of such surgical world models, require expensive annotations or complex structured intermediates as conditioning signals at inference, limiting their scalability. Other approaches exhibit limited temporal consistency across complex laparoscopic scenes and do not possess sufficient realism. We propose Surgical Action World (SAW) -- a step toward surgical action world modeling through video diffusion conditioned on four lightweight signals: language prompts encoding tool-action context, a reference surgical scene, tissue affordance mask, and 2D tool-tip trajectories. We design a conditional video diffusion approach that reformulates video-to-video diffusion into trajectory-conditioned surgical action synthesis. The backbone diffusion model is fine-tuned on a custom-curated dataset of 12,044 laparoscopic clips with lightweight spatiotemporal conditioning signals, leveraging a depth consistency loss to enforce geometric plausibility without requiring depth at inference. SAW achieves state-of-the-art temporal consistency (CD-FVD: 199.19 vs. 546.82) and strong visual quality on held-out test data. Furthermore, we demonstrate its downstream utility for (a) surgical AI, where augmenting rare actions with SAW-generated videos improves action recognition (clipping F1-score: 20.93% to 43.14%; cutting: 0.00% to 8.33%) on real test data, and (b) surgical simulation, where rendering tool-tissue interaction videos from simulator-derived trajectory points toward a visually faithful simulation engine.
Surg-R1: A Hierarchical Reasoning Foundation Model for Scalable and Interpretable Surgical Decision Support with Multi-Center Clinical Validation
Mar 12, 2026Abstract:Surgical scene understanding demands not only accurate predictions but also interpretable reasoning that surgeons can verify against clinical expertise. However, existing surgical vision-language models generate predictions without reasoning chains, and general-purpose reasoning models fail on compositional surgical tasks without domain-specific knowledge. We present Surg-R1, a surgical Vision-Language Model that addresses this gap through hierarchical reasoning trained via a four-stage pipeline. Our approach introduces three key contributions: (1) a three-level reasoning hierarchy decomposing surgical interpretation into perceptual grounding, relational understanding, and contextual reasoning; (2) the largest surgical chain-of-thought dataset with 320,000 reasoning pairs; and (3) a four-stage training pipeline progressing from supervised fine-tuning to group relative policy optimization and iterative self-improvement. Evaluation on SurgBench, comprising six public benchmarks and six multi-center external validation datasets from five institutions, demonstrates that Surg-R1 achieves the highest Arena Score (64.9%) on public benchmarks versus Gemini 3.0 Pro (46.1%) and GPT-5.1 (37.9%), outperforming both proprietary reasoning models and specialized surgical VLMs on the majority of tasks spanning instrument localization, triplet recognition, phase recognition, action recognition, and critical view of safety assessment, with a 15.2 percentage point improvement over the strongest surgical baseline on external validation.
Humanoid Robots as First Assistants in Endoscopic Surgery
Feb 27, 2026Abstract:Humanoid robots have become a focal point of technological ambition, with claims of surgical capability within years in mainstream discourse. These projections are aspirational yet lack empirical grounding. To date, no humanoid has assisted a surgeon through an actual procedure, let alone performed one. The work described here breaks this new ground. Here we report a proof of concept in which a teleoperated Unitree G1 provided endoscopic visualization while an attending otolaryngologist performed a cadaveric sphenoidectomy. The procedure was completed successfully, with stable visualization maintained throughout. Teleoperation allowed assessment of whether the humanoid form factor could meet the physical demands of surgical assistance in terms of sustenance and precision; the cognitive demands were satisfied -- for now -- by the operator. Post-procedure analysis identified engineering targets for clinical translation, alongside near-term opportunities such as autonomous diagnostic scoping. This work establishes form-factor feasibility for humanoid surgical assistance while identifying challenges for continued development.
LUMEN: Longitudinal Multi-Modal Radiology Model for Prognosis and Diagnosis
Feb 24, 2026Abstract:Large vision-language models (VLMs) have evolved from general-purpose applications to specialized use cases such as in the clinical domain, demonstrating potential for decision support in radiology. One promising application is assisting radiologists in decision-making by the analysis of radiology imaging data such as chest X-rays (CXR) via a visual and natural language question-answering (VQA) interface. When longitudinal imaging is available, radiologists analyze temporal changes, which are essential for accurate diagnosis and prognosis. The manual longitudinal analysis is a time-consuming process, motivating the development of a training framework that can provide prognostic capabilities. We introduce a novel training framework LUMEN, that is optimized for longitudinal CXR interpretation, leveraging multi-image and multi-task instruction fine-tuning to enhance prognostic and diagnostic performance. We conduct experiments on the publicly available MIMIC-CXR and its associated Medical-Diff-VQA datasets. We further formulate and construct a novel instruction-following dataset incorporating longitudinal studies, enabling the development of a prognostic VQA task. Our method demonstrates significant improvements over baseline models in diagnostic VQA tasks, and more importantly, shows promising potential for prognostic capabilities. These results underscore the value of well-designed, instruction-tuned VLMs in enabling more accurate and clinically meaningful radiological interpretation of longitudinal radiological imaging data.
VISTA-PATH: An interactive foundation model for pathology image segmentation and quantitative analysis in computational pathology
Jan 23, 2026Abstract:Accurate semantic segmentation for histopathology image is crucial for quantitative tissue analysis and downstream clinical modeling. Recent segmentation foundation models have improved generalization through large-scale pretraining, yet remain poorly aligned with pathology because they treat segmentation as a static visual prediction task. Here we present VISTA-PATH, an interactive, class-aware pathology segmentation foundation model designed to resolve heterogeneous structures, incorporate expert feedback, and produce pixel-level segmentation that are directly meaningful for clinical interpretation. VISTA-PATH jointly conditions segmentation on visual context, semantic tissue descriptions, and optional expert-provided spatial prompts, enabling precise multi-class segmentation across heterogeneous pathology images. To support this paradigm, we curate VISTA-PATH Data, a large-scale pathology segmentation corpus comprising over 1.6 million image-mask-text triplets spanning 9 organs and 93 tissue classes. Across extensive held-out and external benchmarks, VISTA-PATH consistently outperforms existing segmentation foundation models. Importantly, VISTA-PATH supports dynamic human-in-the-loop refinement by propagating sparse, patch-level bounding-box annotation feedback into whole-slide segmentation. Finally, we show that the high-fidelity, class-aware segmentation produced by VISTA-PATH is a preferred model for computational pathology. It improve tissue microenvironment analysis through proposed Tumor Interaction Score (TIS), which exhibits strong and significant associations with patient survival. Together, these results establish VISTA-PATH as a foundation model that elevates pathology image segmentation from a static prediction to an interactive and clinically grounded representation for digital pathology. Source code and demo can be found at https://github.com/zhihuanglab/VISTA-PATH.
SurgWorld: Learning Surgical Robot Policies from Videos via World Modeling
Dec 30, 2025Abstract:Data scarcity remains a fundamental barrier to achieving fully autonomous surgical robots. While large scale vision language action (VLA) models have shown impressive generalization in household and industrial manipulation by leveraging paired video action data from diverse domains, surgical robotics suffers from the paucity of datasets that include both visual observations and accurate robot kinematics. In contrast, vast corpora of surgical videos exist, but they lack corresponding action labels, preventing direct application of imitation learning or VLA training. In this work, we aim to alleviate this problem by learning policy models from SurgWorld, a world model designed for surgical physical AI. We curated the Surgical Action Text Alignment (SATA) dataset with detailed action description specifically for surgical robots. Then we built SurgeWorld based on the most advanced physical AI world model and SATA. It's able to generate diverse, generalizable and realistic surgery videos. We are also the first to use an inverse dynamics model to infer pseudokinematics from synthetic surgical videos, producing synthetic paired video action data. We demonstrate that a surgical VLA policy trained with these augmented data significantly outperforms models trained only on real demonstrations on a real surgical robot platform. Our approach offers a scalable path toward autonomous surgical skill acquisition by leveraging the abundance of unlabeled surgical video and generative world modeling, thus opening the door to generalizable and data efficient surgical robot policies.
SDUM: A Scalable Deep Unrolled Model for Universal MRI Reconstruction
Dec 19, 2025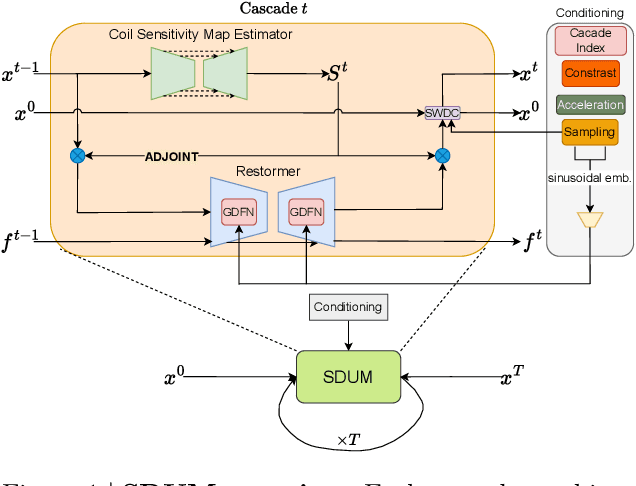
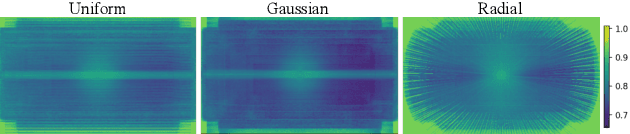

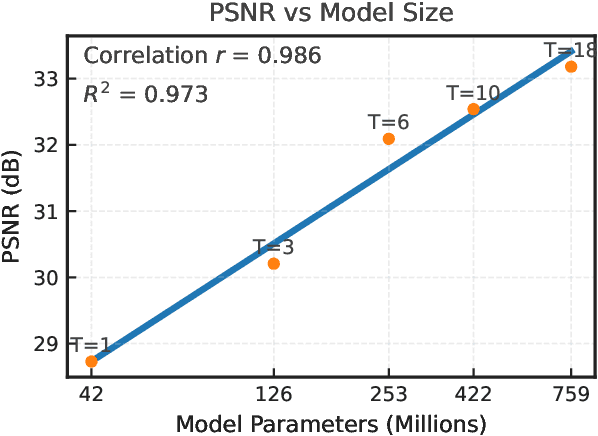
Abstract:Clinical MRI encompasses diverse imaging protocols--spanning anatomical targets (cardiac, brain, knee), contrasts (T1, T2, mapping), sampling patterns (Cartesian, radial, spiral, kt-space), and acceleration factors--yet current deep learning reconstructions are typically protocol-specific, hindering generalization and deployment. We introduce Scalable Deep Unrolled Model (SDUM), a universal framework combining a Restormer-based reconstructor, a learned coil sensitivity map estimator (CSME), sampling-aware weighted data consistency (SWDC), universal conditioning (UC) on cascade index and protocol metadata, and progressive cascade expansion training. SDUM exhibits foundation-model-like scaling behavior: reconstruction quality follows PSNR ${\sim}$ log(parameters) with correlation $r{=}0.986$ ($R^2{=}0.973$) up to 18 cascades, demonstrating predictable performance gains with model depth. A single SDUM trained on heterogeneous data achieves state-of-the-art results across all four CMRxRecon2025 challenge tracks--multi-center, multi-disease, 5T, and pediatric--without task-specific fine-tuning, surpassing specialized baselines by up to ${+}1.0$~dB. On CMRxRecon2024, SDUM outperforms the winning method PromptMR+ by ${+}0.55$~dB; on fastMRI brain, it exceeds PC-RNN by ${+}1.8$~dB. Ablations validate each component: SWDC ${+}0.43$~dB over standard DC, per-cascade CSME ${+}0.51$~dB, UC ${+}0.38$~dB. These results establish SDUM as a practical path toward universal, scalable MRI reconstruction.
See More, Change Less: Anatomy-Aware Diffusion for Contrast Enhancement
Dec 08, 2025Abstract:Image enhancement improves visual quality and helps reveal details that are hard to see in the original image. In medical imaging, it can support clinical decision-making, but current models often over-edit. This can distort organs, create false findings, and miss small tumors because these models do not understand anatomy or contrast dynamics. We propose SMILE, an anatomy-aware diffusion model that learns how organs are shaped and how they take up contrast. It enhances only clinically relevant regions while leaving all other areas unchanged. SMILE introduces three key ideas: (1) structure-aware supervision that follows true organ boundaries and contrast patterns; (2) registration-free learning that works directly with unaligned multi-phase CT scans; (3) unified inference that provides fast and consistent enhancement across all contrast phases. Across six external datasets, SMILE outperforms existing methods in image quality (14.2% higher SSIM, 20.6% higher PSNR, 50% better FID) and in clinical usefulness by producing anatomically accurate and diagnostically meaningful images. SMILE also improves cancer detection from non-contrast CT, raising the F1 score by up to 10 percent.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge