Can Zhao
TouchGuide: Inference-Time Steering of Visuomotor Policies via Touch Guidance
Jan 28, 2026Abstract:Fine-grained and contact-rich manipulation remain challenging for robots, largely due to the underutilization of tactile feedback. To address this, we introduce TouchGuide, a novel cross-policy visuo-tactile fusion paradigm that fuses modalities within a low-dimensional action space. Specifically, TouchGuide operates in two stages to guide a pre-trained diffusion or flow-matching visuomotor policy at inference time. First, the policy produces a coarse, visually-plausible action using only visual inputs during early sampling. Second, a task-specific Contact Physical Model (CPM) provides tactile guidance to steer and refine the action, ensuring it aligns with realistic physical contact conditions. Trained through contrastive learning on limited expert demonstrations, the CPM provides a tactile-informed feasibility score to steer the sampling process toward refined actions that satisfy physical contact constraints. Furthermore, to facilitate TouchGuide training with high-quality and cost-effective data, we introduce TacUMI, a data collection system. TacUMI achieves a favorable trade-off between precision and affordability; by leveraging rigid fingertips, it obtains direct tactile feedback, thereby enabling the collection of reliable tactile data. Extensive experiments on five challenging contact-rich tasks, such as shoe lacing and chip handover, show that TouchGuide consistently and significantly outperforms state-of-the-art visuo-tactile policies.
Cross-Modal Attention Network with Dual Graph Learning in Multimodal Recommendation
Jan 16, 2026Abstract:Multimedia recommendation systems leverage user-item interactions and multimodal information to capture user preferences, enabling more accurate and personalized recommendations. Despite notable advancements, existing approaches still face two critical limitations: first, shallow modality fusion often relies on simple concatenation, failing to exploit rich synergic intra- and inter-modal relationships; second, asymmetric feature treatment-where users are only characterized by interaction IDs while items benefit from rich multimodal content-hinders the learning of a shared semantic space. To address these issues, we propose a Cross-modal Recursive Attention Network with dual graph Embedding (CRANE). To tackle shallow fusion, we design a core Recursive Cross-Modal Attention (RCA) mechanism that iteratively refines modality features based on cross-correlations in a joint latent space, effectively capturing high-order intra- and inter-modal dependencies. For symmetric multimodal learning, we explicitly construct users' multimodal profiles by aggregating features of their interacted items. Furthermore, CRANE integrates a symmetric dual-graph framework-comprising a heterogeneous user-item interaction graph and a homogeneous item-item semantic graph-unified by a self-supervised contrastive learning objective to fuse behavioral and semantic signals. Despite these complex modeling capabilities, CRANE maintains high computational efficiency. Theoretical and empirical analyses confirm its scalability and high practical efficiency, achieving faster convergence on small datasets and superior performance ceilings on large-scale ones. Comprehensive experiments on four public real-world datasets validate an average 5% improvement in key metrics over state-of-the-art baselines.
Knots: A Large-Scale Multi-Agent Enhanced Expert-Annotated Dataset and LLM Prompt Optimization for NOTAM Semantic Parsing
Nov 16, 2025Abstract:Notice to Air Missions (NOTAMs) serve as a critical channel for disseminating key flight safety information, yet their complex linguistic structures and implicit reasoning pose significant challenges for automated parsing. Existing research mainly focuses on surface-level tasks such as classification and named entity recognition, lacking deep semantic understanding. To address this gap, we propose NOTAM semantic parsing, a task emphasizing semantic inference and the integration of aviation domain knowledge to produce structured, inference-rich outputs. To support this task, we construct Knots (Knowledge and NOTAM Semantics), a high-quality dataset of 12,347 expert-annotated NOTAMs covering 194 Flight Information Regions, enhanced through a multi-agent collaborative framework for comprehensive field discovery. We systematically evaluate a wide range of prompt-engineering strategies and model-adaptation techniques, achieving substantial improvements in aviation text understanding and processing. Our experimental results demonstrate the effectiveness of the proposed approach and offer valuable insights for automated NOTAM analysis systems. Our code is available at: https://github.com/Estrellajer/Knots.
NOTAM-Evolve: A Knowledge-Guided Self-Evolving Optimization Framework with LLMs for NOTAM Interpretation
Nov 11, 2025Abstract:Accurate interpretation of Notices to Airmen (NOTAMs) is critical for aviation safety, yet their condensed and cryptic language poses significant challenges to both manual and automated processing. Existing automated systems are typically limited to shallow parsing, failing to extract the actionable intelligence needed for operational decisions. We formalize the complete interpretation task as deep parsing, a dual-reasoning challenge requiring both dynamic knowledge grounding (linking the NOTAM to evolving real-world aeronautical data) and schema-based inference (applying static domain rules to deduce operational status). To tackle this challenge, we propose NOTAM-Evolve, a self-evolving framework that enables a large language model (LLM) to autonomously master complex NOTAM interpretation. Leveraging a knowledge graph-enhanced retrieval module for data grounding, the framework introduces a closed-loop learning process where the LLM progressively improves from its own outputs, minimizing the need for extensive human-annotated reasoning traces. In conjunction with this framework, we introduce a new benchmark dataset of 10,000 expert-annotated NOTAMs. Our experiments demonstrate that NOTAM-Evolve achieves a 30.4% absolute accuracy improvement over the base LLM, establishing a new state of the art on the task of structured NOTAM interpretation.
RANA: Robust Active Learning for Noisy Network Alignment
Jul 30, 2025Abstract:Network alignment has attracted widespread attention in various fields. However, most existing works mainly focus on the problem of label sparsity, while overlooking the issue of noise in network alignment, which can substantially undermine model performance. Such noise mainly includes structural noise from noisy edges and labeling noise caused by human-induced and process-driven errors. To address these problems, we propose RANA, a Robust Active learning framework for noisy Network Alignment. RANA effectively tackles both structure noise and label noise while addressing the sparsity of anchor link annotations, which can improve the robustness of network alignment models. Specifically, RANA introduces the proposed Noise-aware Selection Module and the Label Denoising Module to address structural noise and labeling noise, respectively. In the first module, we design a noise-aware maximization objective to select node pairs, incorporating a cleanliness score to address structural noise. In the second module, we propose a novel multi-source fusion denoising strategy that leverages model and twin node pairs labeling to provide more accurate labels for node pairs. Empirical results on three real-world datasets demonstrate that RANA outperforms state-of-the-art active learning-based methods in alignment accuracy. Our code is available at https://github.com/YXNan0110/RANA.
Text2CT: Towards 3D CT Volume Generation from Free-text Descriptions Using Diffusion Model
May 07, 2025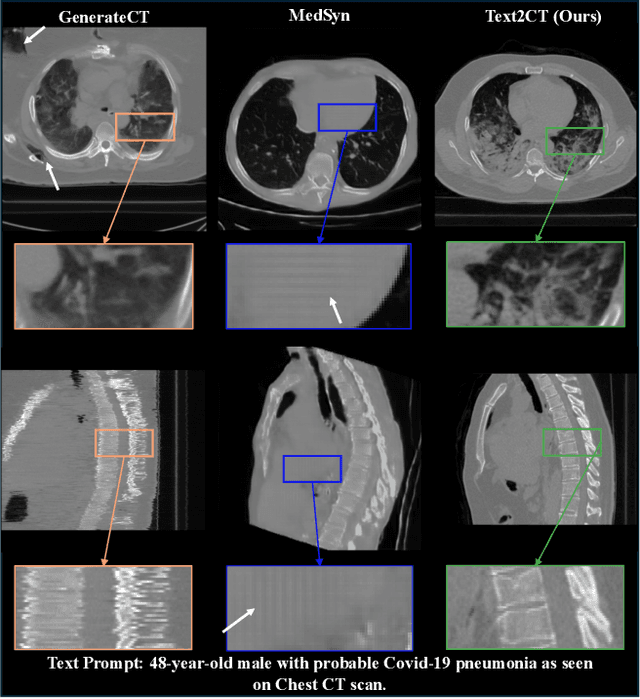
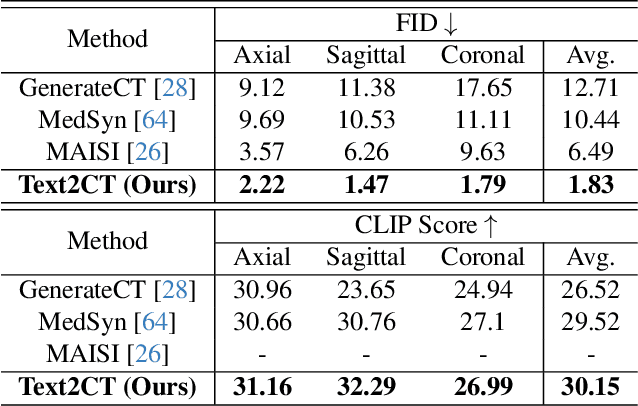
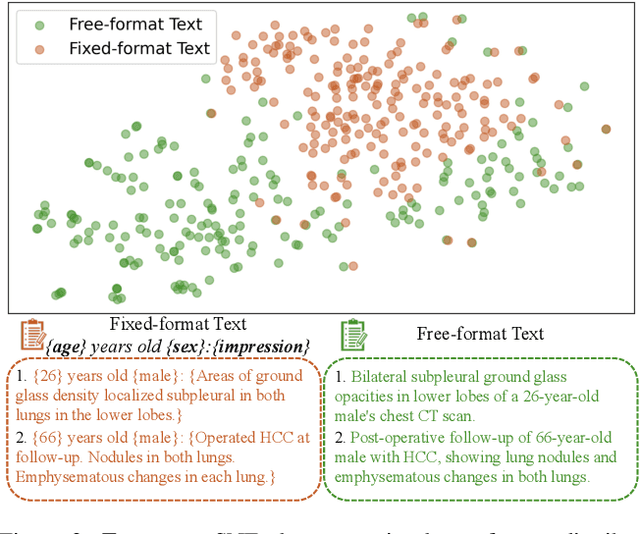
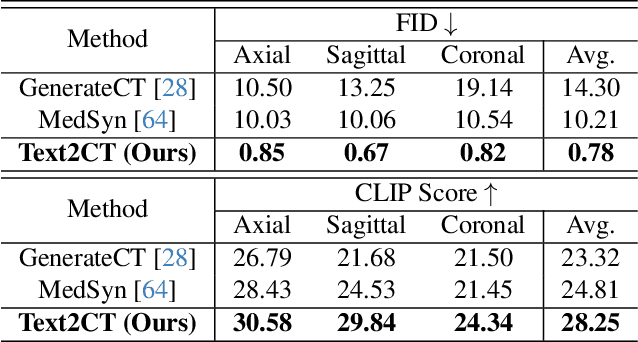
Abstract:Generating 3D CT volumes from descriptive free-text inputs presents a transformative opportunity in diagnostics and research. In this paper, we introduce Text2CT, a novel approach for synthesizing 3D CT volumes from textual descriptions using the diffusion model. Unlike previous methods that rely on fixed-format text input, Text2CT employs a novel prompt formulation that enables generation from diverse, free-text descriptions. The proposed framework encodes medical text into latent representations and decodes them into high-resolution 3D CT scans, effectively bridging the gap between semantic text inputs and detailed volumetric representations in a unified 3D framework. Our method demonstrates superior performance in preserving anatomical fidelity and capturing intricate structures as described in the input text. Extensive evaluations show that our approach achieves state-of-the-art results, offering promising potential applications in diagnostics, and data augmentation.
Pushing DSP-Free Coherent Interconnect to the Last Inch by Optically Analog Signal Processing
Mar 14, 2025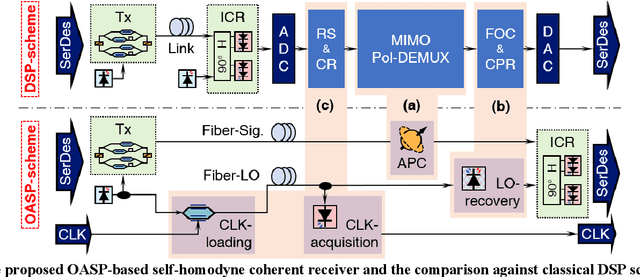
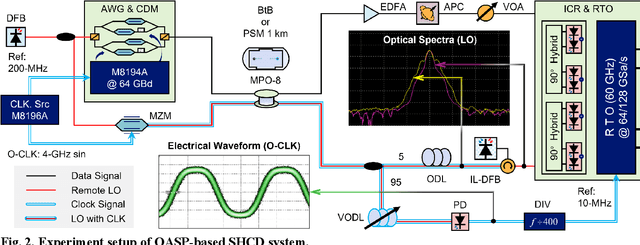
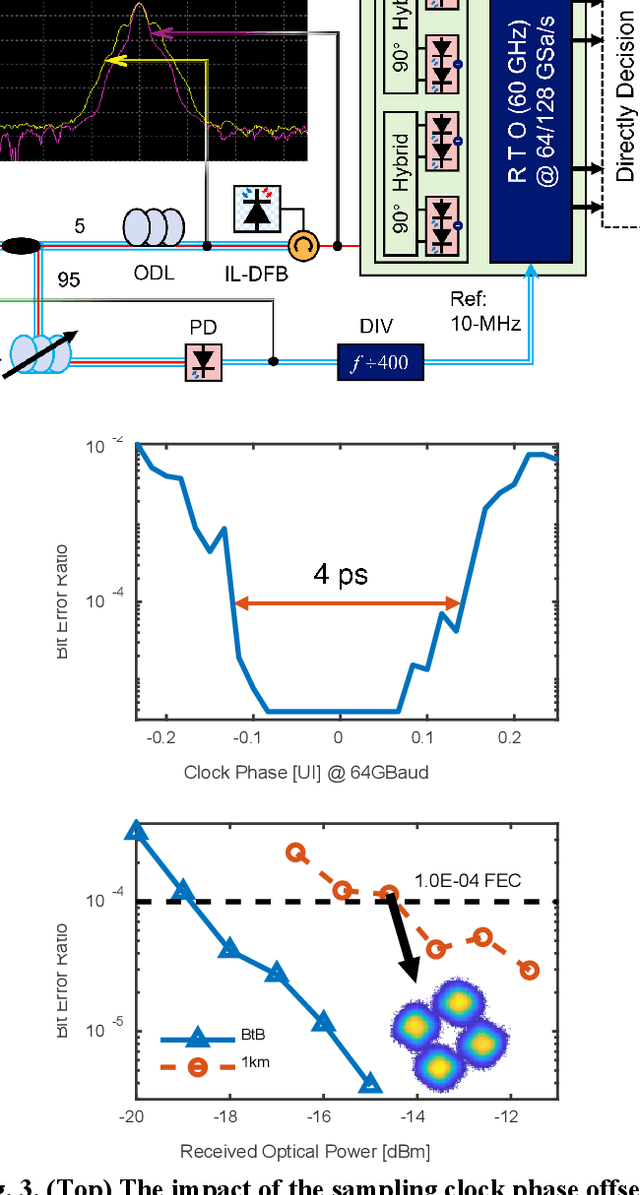
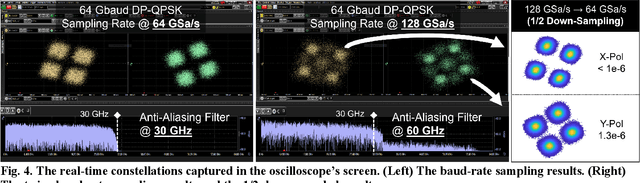
Abstract:To support the boosting interconnect capacity of the AI-related data centers, novel techniques enabled high-speed and low-cost optics are continuously emerging. When the baud rate approaches 200 GBaud per lane, the bottle-neck of traditional intensity modulation direct detection (IM-DD) architectures becomes increasingly evident. The simplified coherent solutions are widely discussed and considered as one of the most promising candidates. In this paper, a novel coherent architecture based on self-homodyne coherent detection and optically analog signal processing (OASP) is demonstrated. Proved by experiment, the first DSP-free baud-rate sampled 64-GBaud QPSK/16-QAM receptions are achieved, with BERs of 1e-6 and 2e-2, respectively. Even with 1-km fiber link propagation, the BER for QPSK reception remains at 3.6e-6. When an ultra-simple 1-sps SISO filter is utilized, the performance degradation of the proposed scheme is less than 1 dB compared to legacy DSP-based coherent reception. The proposed results pave the way for the ultra-high-speed coherent optical interconnections, offering high power and cost efficiency.
Interactive Tumor Progression Modeling via Sketch-Based Image Editing
Mar 10, 2025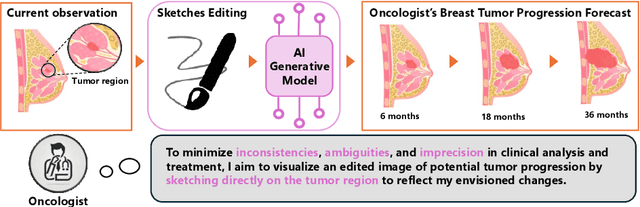
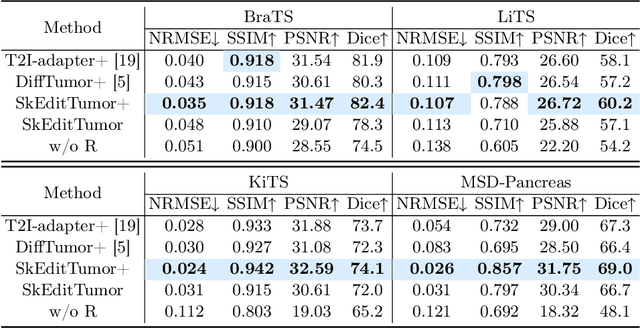
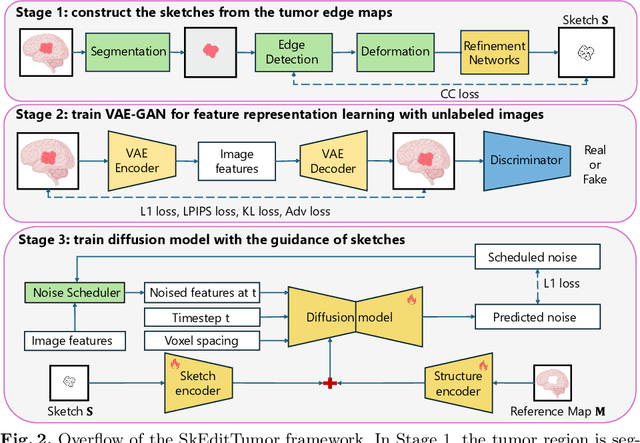
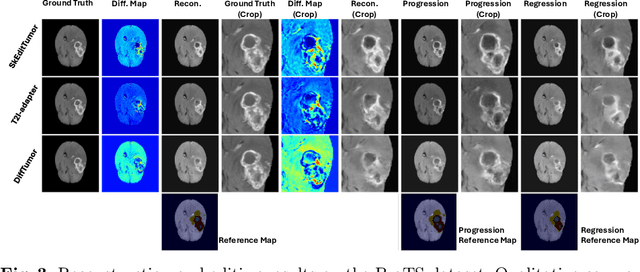
Abstract:Accurately visualizing and editing tumor progression in medical imaging is crucial for diagnosis, treatment planning, and clinical communication. To address the challenges of subjectivity and limited precision in existing methods, we propose SkEditTumor, a sketch-based diffusion model for controllable tumor progression editing. By leveraging sketches as structural priors, our method enables precise modifications of tumor regions while maintaining structural integrity and visual realism. We evaluate SkEditTumor on four public datasets - BraTS, LiTS, KiTS, and MSD-Pancreas - covering diverse organs and imaging modalities. Experimental results demonstrate that our method outperforms state-of-the-art baselines, achieving superior image fidelity and segmentation accuracy. Our contributions include a novel integration of sketches with diffusion models for medical image editing, fine-grained control over tumor progression visualization, and extensive validation across multiple datasets, setting a new benchmark in the field.
Text-Driven Tumor Synthesis
Dec 24, 2024



Abstract:Tumor synthesis can generate examples that AI often misses or over-detects, improving AI performance by training on these challenging cases. However, existing synthesis methods, which are typically unconditional -- generating images from random variables -- or conditioned only by tumor shapes, lack controllability over specific tumor characteristics such as texture, heterogeneity, boundaries, and pathology type. As a result, the generated tumors may be overly similar or duplicates of existing training data, failing to effectively address AI's weaknesses. We propose a new text-driven tumor synthesis approach, termed TextoMorph, that provides textual control over tumor characteristics. This is particularly beneficial for examples that confuse the AI the most, such as early tumor detection (increasing Sensitivity by +8.5%), tumor segmentation for precise radiotherapy (increasing DSC by +6.3%), and classification between benign and malignant tumors (improving Sensitivity by +8.2%). By incorporating text mined from radiology reports into the synthesis process, we increase the variability and controllability of the synthetic tumors to target AI's failure cases more precisely. Moreover, TextoMorph uses contrastive learning across different texts and CT scans, significantly reducing dependence on scarce image-report pairs (only 141 pairs used in this study) by leveraging a large corpus of 34,035 radiology reports. Finally, we have developed rigorous tests to evaluate synthetic tumors, including Text-Driven Visual Turing Test and Radiomics Pattern Analysis, showing that our synthetic tumors is realistic and diverse in texture, heterogeneity, boundaries, and pathology.
VILA-M3: Enhancing Vision-Language Models with Medical Expert Knowledge
Nov 19, 2024



Abstract:Generalist vision language models (VLMs) have made significant strides in computer vision, but they fall short in specialized fields like healthcare, where expert knowledge is essential. In traditional computer vision tasks, creative or approximate answers may be acceptable, but in healthcare, precision is paramount.Current large multimodal models like Gemini and GPT-4o are insufficient for medical tasks due to their reliance on memorized internet knowledge rather than the nuanced expertise required in healthcare. VLMs are usually trained in three stages: vision pre-training, vision-language pre-training, and instruction fine-tuning (IFT). IFT has been typically applied using a mixture of generic and healthcare data. In contrast, we propose that for medical VLMs, a fourth stage of specialized IFT is necessary, which focuses on medical data and includes information from domain expert models. Domain expert models developed for medical use are crucial because they are specifically trained for certain clinical tasks, e.g. to detect tumors and classify abnormalities through segmentation and classification, which learn fine-grained features of medical data$-$features that are often too intricate for a VLM to capture effectively especially in radiology. This paper introduces a new framework, VILA-M3, for medical VLMs that utilizes domain knowledge via expert models. Through our experiments, we show an improved state-of-the-art (SOTA) performance with an average improvement of ~9% over the prior SOTA model Med-Gemini and ~6% over models trained on the specific tasks. Our approach emphasizes the importance of domain expertise in creating precise, reliable VLMs for medical applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge