A Short Review and Evaluation of SAM2's Performance in 3D CT Image Segmentation
Paper and Code
Aug 20, 2024
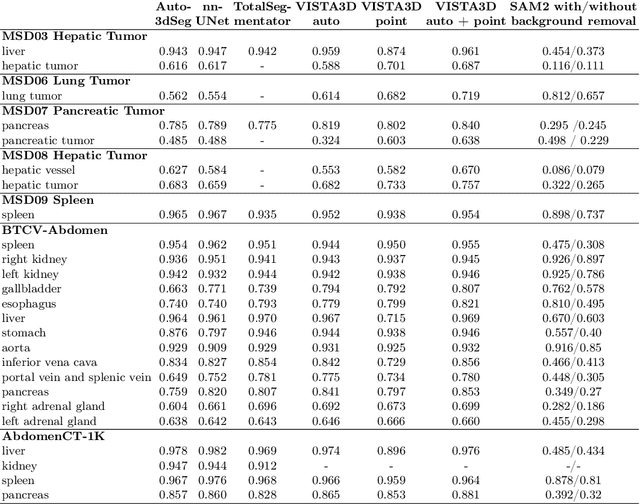
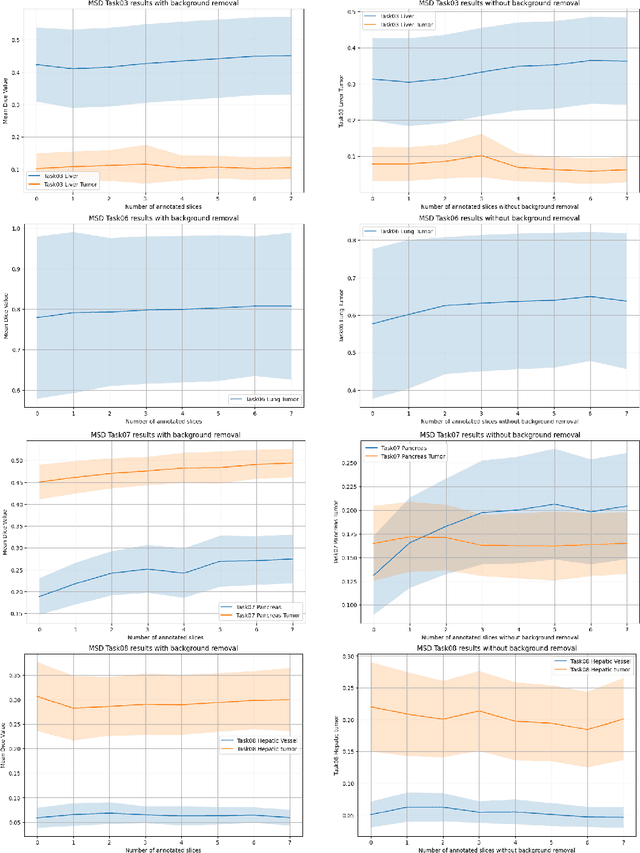
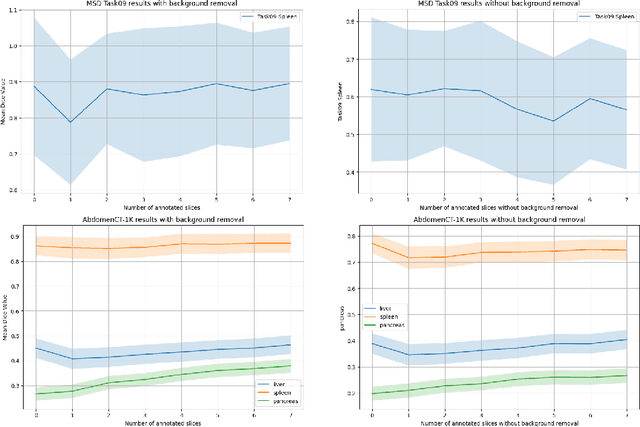
Since the release of Segment Anything 2 (SAM2), the medical imaging community has been actively evaluating its performance for 3D medical image segmentation. However, different studies have employed varying evaluation pipelines, resulting in conflicting outcomes that obscure a clear understanding of SAM2's capabilities and potential applications. We shortly review existing benchmarks and point out that the SAM2 paper clearly outlines a zero-shot evaluation pipeline, which simulates user clicks iteratively for up to eight iterations. We reproduced this interactive annotation simulation on 3D CT datasets and provided the results and code~\url{https://github.com/Project-MONAI/VISTA}. Our findings reveal that directly applying SAM2 on 3D medical imaging in a zero-shot manner is far from satisfactory. It is prone to generating false positives when foreground objects disappear, and annotating more slices cannot fully offset this tendency. For smaller single-connected objects like kidney and aorta, SAM2 performs reasonably well but for most organs it is still far behind state-of-the-art 3D annotation methods. More research and innovation are needed for 3D medical imaging community to use SAM2 correctly.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge