Ziyue Xu
Standardized Methods and Recommendations for Green Federated Learning
Jan 30, 2026Abstract:Federated learning (FL) enables collaborative model training over privacy-sensitive, distributed data, but its environmental impact is difficult to compare across studies due to inconsistent measurement boundaries and heterogeneous reporting. We present a practical carbon-accounting methodology for FL CO2e tracking using NVIDIA NVFlare and CodeCarbon for explicit, phase-aware tasks (initialization, per-round training, evaluation, and idle/coordination). To capture non-compute effects, we additionally estimate communication emissions from transmitted model-update sizes under a network-configurable energy model. We validate the proposed approach on two representative workloads: CIFAR-10 image classification and retinal optic disk segmentation. In CIFAR-10, controlled client-efficiency scenarios show that system-level slowdowns and coordination effects can contribute meaningfully to carbon footprint under an otherwise fixed FL protocol, increasing total CO2e by 8.34x (medium) and 21.73x (low) relative to the high-efficiency baseline. In retinal segmentation, swapping GPU tiers (H100 vs.\ V100) yields a consistent 1.7x runtime gap (290 vs. 503 minutes) while producing non-uniform changes in total energy and CO2e across sites, underscoring the need for per-site and per-round reporting. Overall, our results support a standardized carbon accounting method that acts as a prerequisite for reproducible 'green' FL evaluation. Our code is available at https://github.com/Pediatric-Accelerated-Intelligence-Lab/carbon_footprint.
FeTTL: Federated Template and Task Learning for Multi-Institutional Medical Imaging
Jan 22, 2026Abstract:Federated learning enables collaborative model training across geographically distributed medical centers while preserving data privacy. However, domain shifts and heterogeneity in data often lead to a degradation in model performance. Medical imaging applications are particularly affected by variations in acquisition protocols, scanner types, and patient populations. To address these issues, we introduce Federated Template and Task Learning (FeTTL), a novel framework designed to harmonize multi-institutional medical imaging data in federated environments. FeTTL learns a global template together with a task model to align data distributions among clients. We evaluated FeTTL on two challenging and diverse multi-institutional medical imaging tasks: retinal fundus optical disc segmentation and histopathological metastasis classification. Experimental results show that FeTTL significantly outperforms the state-of-the-art federated learning baselines (p-values <0.002) for optical disc segmentation and classification of metastases from multi-institutional data. Our experiments further highlight the importance of jointly learning the template and the task. These findings suggest that FeTTL offers a principled and extensible solution for mitigating distribution shifts in federated learning, supporting robust model deployment in real-world, multi-institutional environments.
FedUMM: A General Framework for Federated Learning with Unified Multimodal Models
Jan 21, 2026Abstract:Unified multimodal models (UMMs) are emerging as strong foundation models that can do both generation and understanding tasks in a single architecture. However, they are typically trained in centralized settings where all training and downstream datasets are gathered in a central server, limiting the deployment in privacy-sensitive and geographically distributed scenarios. In this paper, we present FedUMM, a general federated learning framework for UMMs under non-IID multimodal data with low communication cost. Built on NVIDIA FLARE, FedUMM instantiates federation for a BLIP3o backbone via parameter-efficient fine-tuning: clients train lightweight LoRA adapters while freezing the foundation models, and the server aggregates only adapter updates. We evaluate on VQA v2 and the GenEval compositional generation benchmarks under Dirichlet-controlled heterogeneity with up to 16 clients. Results show slight degradation as client count and heterogeneity increase, while remaining competitive with centralized training. We further analyze computation--communication trade-offs and demonstrate that adapter-only federation reduces per-round communication by over an order of magnitude compared to full fine-tuning, enabling practical federated UMM training. This work provides empirical experience for future research on privacy-preserving federated unified multimodal models.
Cyst-X: AI-Powered Pancreatic Cancer Risk Prediction from Multicenter MRI in Centralized and Federated Learning
Jul 29, 2025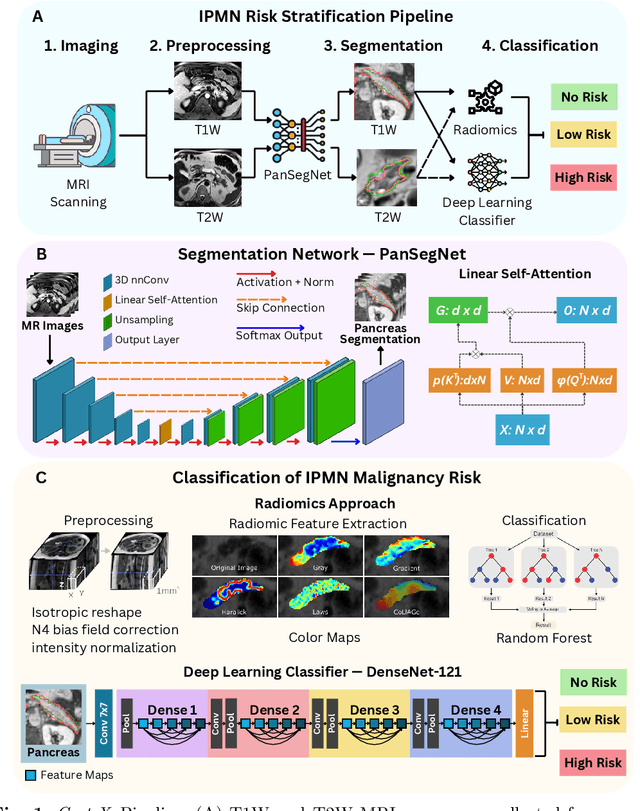

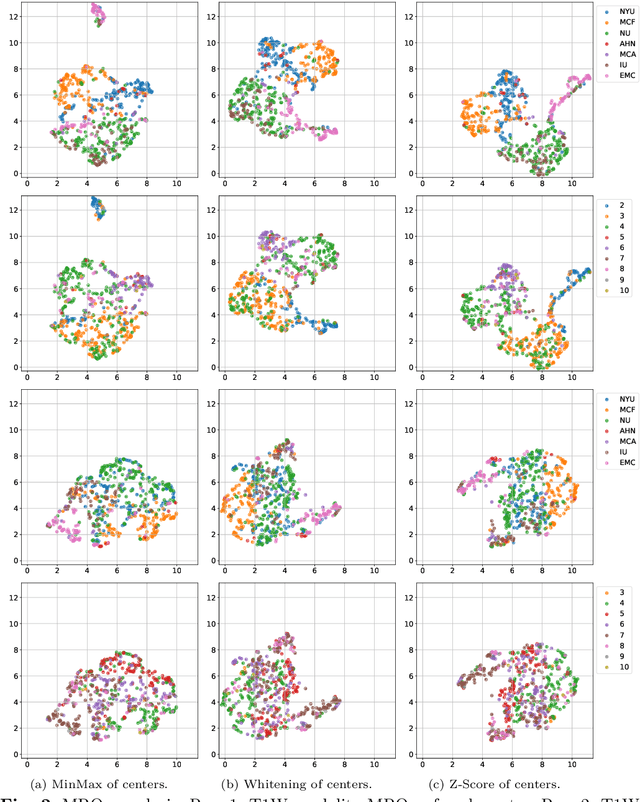
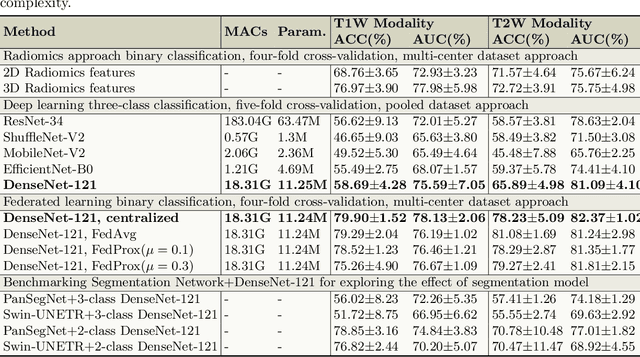
Abstract:Pancreatic cancer is projected to become the second-deadliest malignancy in Western countries by 2030, highlighting the urgent need for better early detection. Intraductal papillary mucinous neoplasms (IPMNs), key precursors to pancreatic cancer, are challenging to assess with current guidelines, often leading to unnecessary surgeries or missed malignancies. We present Cyst-X, an AI framework that predicts IPMN malignancy using multicenter MRI data, leveraging MRI's superior soft tissue contrast over CT. Trained on 723 T1- and 738 T2-weighted scans from 764 patients across seven institutions, our models (AUC=0.82) significantly outperform both Kyoto guidelines (AUC=0.75) and expert radiologists. The AI-derived imaging features align with known clinical markers and offer biologically meaningful insights. We also demonstrate strong performance in a federated learning setting, enabling collaborative training without sharing patient data. To promote privacy-preserving AI development and improve IPMN risk stratification, the Cyst-X dataset is released as the first large-scale, multi-center pancreatic cysts MRI dataset.
Text2CT: Towards 3D CT Volume Generation from Free-text Descriptions Using Diffusion Model
May 07, 2025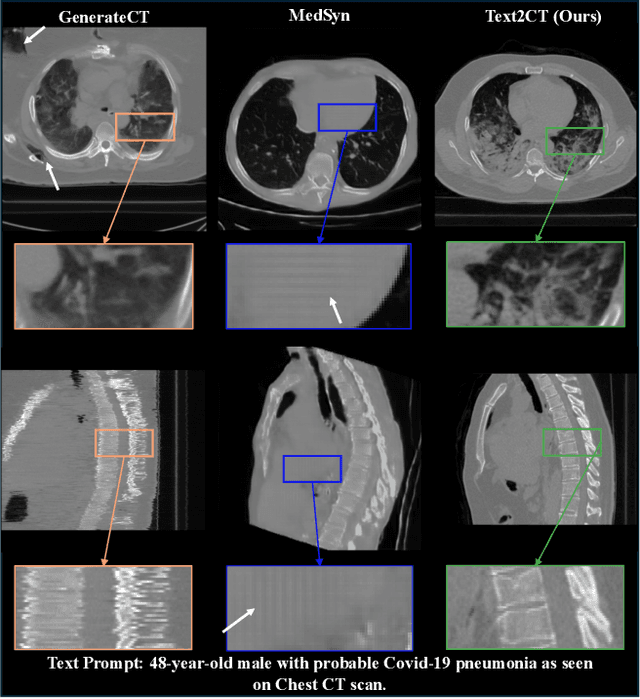
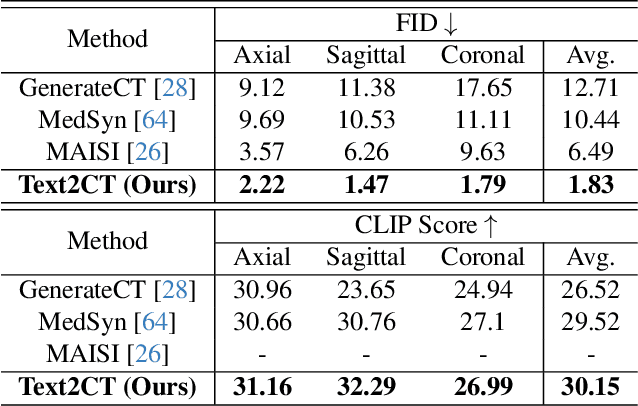
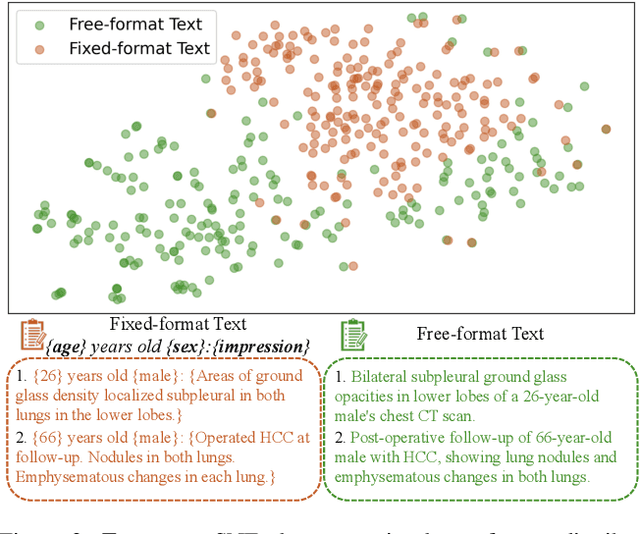
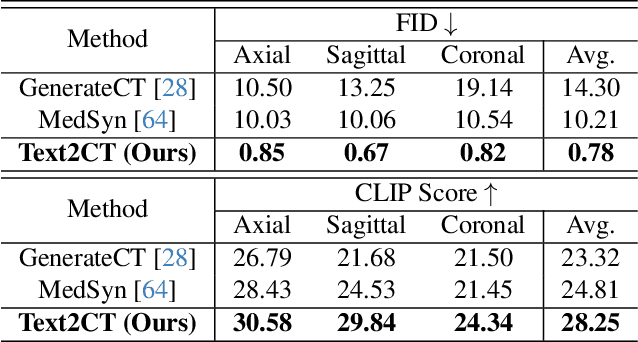
Abstract:Generating 3D CT volumes from descriptive free-text inputs presents a transformative opportunity in diagnostics and research. In this paper, we introduce Text2CT, a novel approach for synthesizing 3D CT volumes from textual descriptions using the diffusion model. Unlike previous methods that rely on fixed-format text input, Text2CT employs a novel prompt formulation that enables generation from diverse, free-text descriptions. The proposed framework encodes medical text into latent representations and decodes them into high-resolution 3D CT scans, effectively bridging the gap between semantic text inputs and detailed volumetric representations in a unified 3D framework. Our method demonstrates superior performance in preserving anatomical fidelity and capturing intricate structures as described in the input text. Extensive evaluations show that our approach achieves state-of-the-art results, offering promising potential applications in diagnostics, and data augmentation.
C-FedRAG: A Confidential Federated Retrieval-Augmented Generation System
Dec 17, 2024



Abstract:Organizations seeking to utilize Large Language Models (LLMs) for knowledge querying and analysis often encounter challenges in maintaining an LLM fine-tuned on targeted, up-to-date information that keeps answers relevant and grounded. Retrieval Augmented Generation (RAG) has quickly become a feasible solution for organizations looking to overcome the challenges of maintaining proprietary models and to help reduce LLM hallucinations in their query responses. However, RAG comes with its own issues regarding scaling data pipelines across tiered-access and disparate data sources. In many scenarios, it is necessary to query beyond a single data silo to provide richer and more relevant context for an LLM. Analyzing data sources within and across organizational trust boundaries is often limited by complex data-sharing policies that prohibit centralized data storage, therefore, inhibit the fast and effective setup and scaling of RAG solutions. In this paper, we introduce Confidential Computing (CC) techniques as a solution for secure Federated Retrieval Augmented Generation (FedRAG). Our proposed Confidential FedRAG system (C-FedRAG) enables secure connection and scaling of a RAG workflows across a decentralized network of data providers by ensuring context confidentiality. We also demonstrate how to implement a C-FedRAG system using the NVIDIA FLARE SDK and assess its performance using the MedRAG toolkit and MIRAGE benchmarking dataset.
VILA-M3: Enhancing Vision-Language Models with Medical Expert Knowledge
Nov 19, 2024



Abstract:Generalist vision language models (VLMs) have made significant strides in computer vision, but they fall short in specialized fields like healthcare, where expert knowledge is essential. In traditional computer vision tasks, creative or approximate answers may be acceptable, but in healthcare, precision is paramount.Current large multimodal models like Gemini and GPT-4o are insufficient for medical tasks due to their reliance on memorized internet knowledge rather than the nuanced expertise required in healthcare. VLMs are usually trained in three stages: vision pre-training, vision-language pre-training, and instruction fine-tuning (IFT). IFT has been typically applied using a mixture of generic and healthcare data. In contrast, we propose that for medical VLMs, a fourth stage of specialized IFT is necessary, which focuses on medical data and includes information from domain expert models. Domain expert models developed for medical use are crucial because they are specifically trained for certain clinical tasks, e.g. to detect tumors and classify abnormalities through segmentation and classification, which learn fine-grained features of medical data$-$features that are often too intricate for a VLM to capture effectively especially in radiology. This paper introduces a new framework, VILA-M3, for medical VLMs that utilizes domain knowledge via expert models. Through our experiments, we show an improved state-of-the-art (SOTA) performance with an average improvement of ~9% over the prior SOTA model Med-Gemini and ~6% over models trained on the specific tasks. Our approach emphasizes the importance of domain expertise in creating precise, reliable VLMs for medical applications.
IPMN Risk Assessment under Federated Learning Paradigm
Nov 08, 2024



Abstract:Accurate classification of Intraductal Papillary Mucinous Neoplasms (IPMN) is essential for identifying high-risk cases that require timely intervention. In this study, we develop a federated learning framework for multi-center IPMN classification utilizing a comprehensive pancreas MRI dataset. This dataset includes 653 T1-weighted and 656 T2-weighted MRI images, accompanied by corresponding IPMN risk scores from 7 leading medical institutions, making it the largest and most diverse dataset for IPMN classification to date. We assess the performance of DenseNet-121 in both centralized and federated settings for training on distributed data. Our results demonstrate that the federated learning approach achieves high classification accuracy comparable to centralized learning while ensuring data privacy across institutions. This work marks a significant advancement in collaborative IPMN classification, facilitating secure and high-accuracy model training across multiple centers.
Adaptive Aggregation Weights for Federated Segmentation of Pancreas MRI
Oct 29, 2024

Abstract:Federated learning (FL) enables collaborative model training across institutions without sharing sensitive data, making it an attractive solution for medical imaging tasks. However, traditional FL methods, such as Federated Averaging (FedAvg), face difficulties in generalizing across domains due to variations in imaging protocols and patient demographics across institutions. This challenge is particularly evident in pancreas MRI segmentation, where anatomical variability and imaging artifacts significantly impact performance. In this paper, we conduct a comprehensive evaluation of FL algorithms for pancreas MRI segmentation and introduce a novel approach that incorporates adaptive aggregation weights. By dynamically adjusting the contribution of each client during model aggregation, our method accounts for domain-specific differences and improves generalization across heterogeneous datasets. Experimental results demonstrate that our approach enhances segmentation accuracy and reduces the impact of domain shift compared to conventional FL methods while maintaining privacy-preserving capabilities. Significant performance improvements are observed across multiple hospitals (centers).
MAISI: Medical AI for Synthetic Imaging
Sep 13, 2024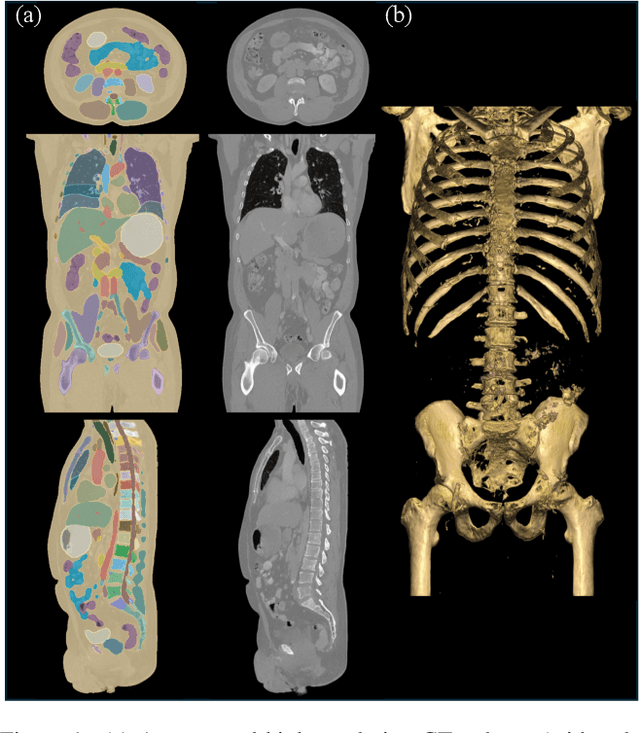
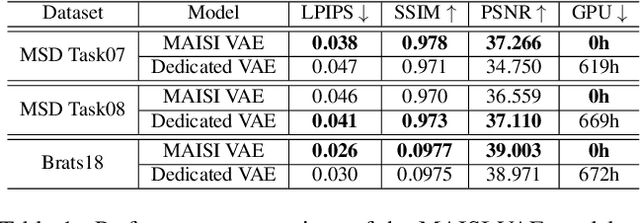
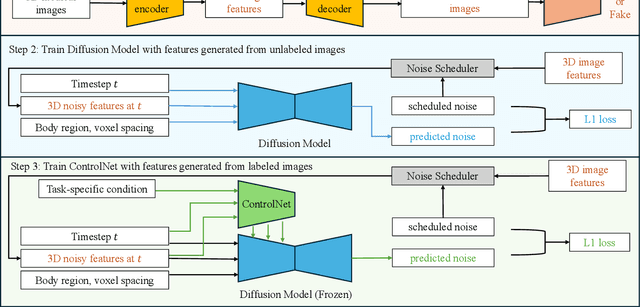
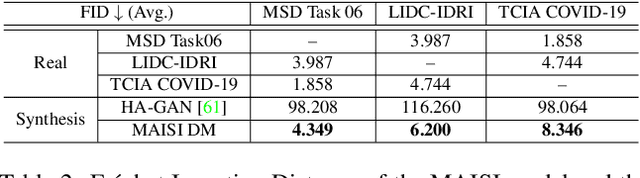
Abstract:Medical imaging analysis faces challenges such as data scarcity, high annotation costs, and privacy concerns. This paper introduces the Medical AI for Synthetic Imaging (MAISI), an innovative approach using the diffusion model to generate synthetic 3D computed tomography (CT) images to address those challenges. MAISI leverages the foundation volume compression network and the latent diffusion model to produce high-resolution CT images (up to a landmark volume dimension of 512 x 512 x 768 ) with flexible volume dimensions and voxel spacing. By incorporating ControlNet, MAISI can process organ segmentation, including 127 anatomical structures, as additional conditions and enables the generation of accurately annotated synthetic images that can be used for various downstream tasks. Our experiment results show that MAISI's capabilities in generating realistic, anatomically accurate images for diverse regions and conditions reveal its promising potential to mitigate challenges using synthetic data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge