Jeya Maria Jose Valanarasu
Student Member, IEEE
Evaluating and Improving the Effectiveness of Synthetic Chest X-Rays for Medical Image Analysis
Nov 27, 2024Abstract:Purpose: To explore best-practice approaches for generating synthetic chest X-ray images and augmenting medical imaging datasets to optimize the performance of deep learning models in downstream tasks like classification and segmentation. Materials and Methods: We utilized a latent diffusion model to condition the generation of synthetic chest X-rays on text prompts and/or segmentation masks. We explored methods like using a proxy model and using radiologist feedback to improve the quality of synthetic data. These synthetic images were then generated from relevant disease information or geometrically transformed segmentation masks and added to ground truth training set images from the CheXpert, CANDID-PTX, SIIM, and RSNA Pneumonia datasets to measure improvements in classification and segmentation model performance on the test sets. F1 and Dice scores were used to evaluate classification and segmentation respectively. One-tailed t-tests with Bonferroni correction assessed the statistical significance of performance improvements with synthetic data. Results: Across all experiments, the synthetic data we generated resulted in a maximum mean classification F1 score improvement of 0.150453 (CI: 0.099108-0.201798; P=0.0031) compared to using only real data. For segmentation, the maximum Dice score improvement was 0.14575 (CI: 0.108267-0.183233; P=0.0064). Conclusion: Best practices for generating synthetic chest X-ray images for downstream tasks include conditioning on single-disease labels or geometrically transformed segmentation masks, as well as potentially using proxy modeling for fine-tuning such generations.
Time-to-Event Pretraining for 3D Medical Imaging
Nov 14, 2024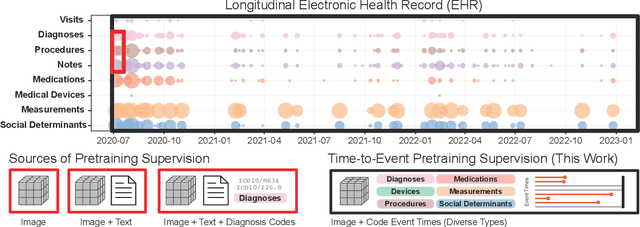
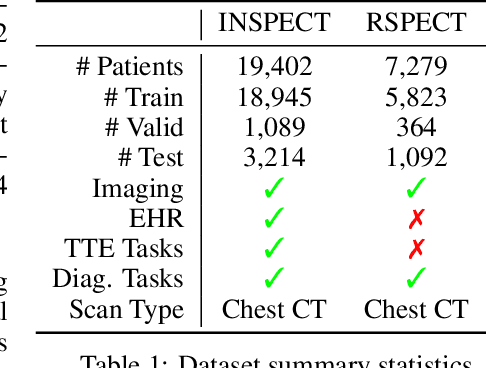
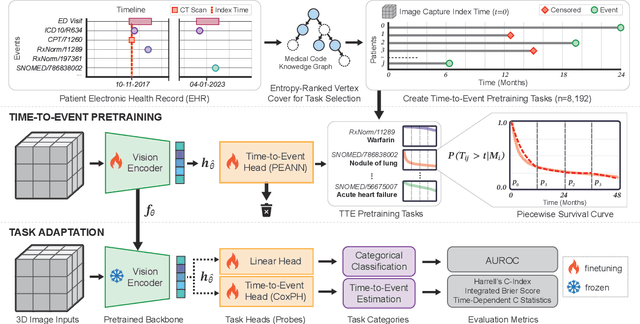
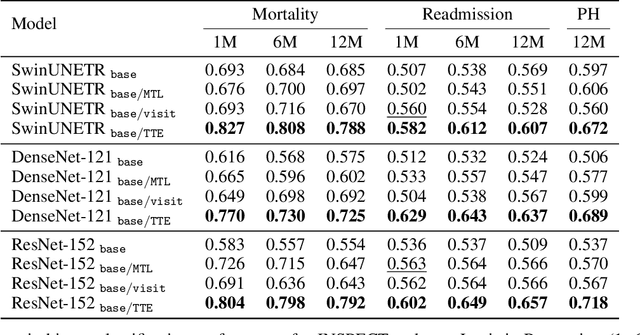
Abstract:With the rise of medical foundation models and the growing availability of imaging data, scalable pretraining techniques offer a promising way to identify imaging biomarkers predictive of future disease risk. While current self-supervised methods for 3D medical imaging models capture local structural features like organ morphology, they fail to link pixel biomarkers with long-term health outcomes due to a missing context problem. Current approaches lack the temporal context necessary to identify biomarkers correlated with disease progression, as they rely on supervision derived only from images and concurrent text descriptions. To address this, we introduce time-to-event pretraining, a pretraining framework for 3D medical imaging models that leverages large-scale temporal supervision from paired, longitudinal electronic health records (EHRs). Using a dataset of 18,945 CT scans (4.2 million 2D images) and time-to-event distributions across thousands of EHR-derived tasks, our method improves outcome prediction, achieving an average AUROC increase of 23.7% and a 29.4% gain in Harrell's C-index across 8 benchmark tasks. Importantly, these gains are achieved without sacrificing diagnostic classification performance. This study lays the foundation for integrating longitudinal EHR and 3D imaging data to advance clinical risk prediction.
Merlin: A Vision Language Foundation Model for 3D Computed Tomography
Jun 10, 2024



Abstract:Over 85 million computed tomography (CT) scans are performed annually in the US, of which approximately one quarter focus on the abdomen. Given the current radiologist shortage, there is a large impetus to use artificial intelligence to alleviate the burden of interpreting these complex imaging studies. Prior state-of-the-art approaches for automated medical image interpretation leverage vision language models (VLMs). However, current medical VLMs are generally limited to 2D images and short reports, and do not leverage electronic health record (EHR) data for supervision. We introduce Merlin - a 3D VLM that we train using paired CT scans (6+ million images from 15,331 CTs), EHR diagnosis codes (1.8+ million codes), and radiology reports (6+ million tokens). We evaluate Merlin on 6 task types and 752 individual tasks. The non-adapted (off-the-shelf) tasks include zero-shot findings classification (31 findings), phenotype classification (692 phenotypes), and zero-shot cross-modal retrieval (image to findings and image to impressions), while model adapted tasks include 5-year disease prediction (6 diseases), radiology report generation, and 3D semantic segmentation (20 organs). We perform internal validation on a test set of 5,137 CTs, and external validation on 7,000 clinical CTs and on two public CT datasets (VerSe, TotalSegmentator). Beyond these clinically-relevant evaluations, we assess the efficacy of various network architectures and training strategies to depict that Merlin has favorable performance to existing task-specific baselines. We derive data scaling laws to empirically assess training data needs for requisite downstream task performance. Furthermore, unlike conventional VLMs that require hundreds of GPUs for training, we perform all training on a single GPU.
Auto-Generating Weak Labels for Real & Synthetic Data to Improve Label-Scarce Medical Image Segmentation
Apr 25, 2024



Abstract:The high cost of creating pixel-by-pixel gold-standard labels, limited expert availability, and presence of diverse tasks make it challenging to generate segmentation labels to train deep learning models for medical imaging tasks. In this work, we present a new approach to overcome the hurdle of costly medical image labeling by leveraging foundation models like Segment Anything Model (SAM) and its medical alternate MedSAM. Our pipeline has the ability to generate weak labels for any unlabeled medical image and subsequently use it to augment label-scarce datasets. We perform this by leveraging a model trained on a few gold-standard labels and using it to intelligently prompt MedSAM for weak label generation. This automation eliminates the manual prompting step in MedSAM, creating a streamlined process for generating labels for both real and synthetic images, regardless of quantity. We conduct experiments on label-scarce settings for multiple tasks pertaining to modalities ranging from ultrasound, dermatology, and X-rays to demonstrate the usefulness of our pipeline. The code is available at https://github.com/stanfordmlgroup/Auto-Generate-WLs/.
Unlocking Robust Segmentation Across All Age Groups via Continual Learning
Apr 19, 2024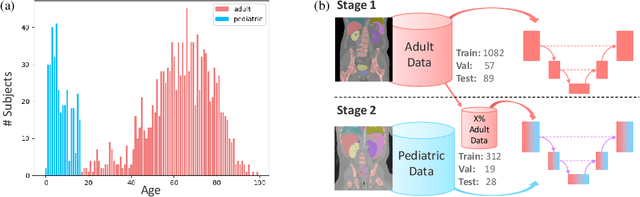

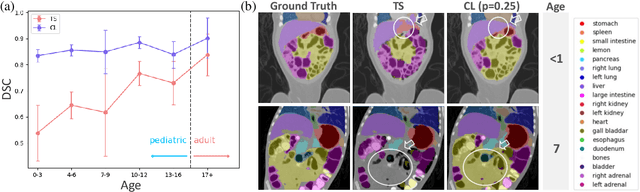
Abstract:Most deep learning models in medical imaging are trained on adult data with unclear performance on pediatric images. In this work, we aim to address this challenge in the context of automated anatomy segmentation in whole-body Computed Tomography (CT). We evaluate the performance of CT organ segmentation algorithms trained on adult data when applied to pediatric CT volumes and identify substantial age-dependent underperformance. We subsequently propose and evaluate strategies, including data augmentation and continual learning approaches, to achieve good segmentation accuracy across all age groups. Our best-performing model, trained using continual learning, achieves high segmentation accuracy on both adult and pediatric data (Dice scores of 0.90 and 0.84 respectively).
MaxFusion: Plug&Play Multi-Modal Generation in Text-to-Image Diffusion Models
Apr 15, 2024



Abstract:Large diffusion-based Text-to-Image (T2I) models have shown impressive generative powers for text-to-image generation as well as spatially conditioned image generation. For most applications, we can train the model end-toend with paired data to obtain photorealistic generation quality. However, to add an additional task, one often needs to retrain the model from scratch using paired data across all modalities to retain good generation performance. In this paper, we tackle this issue and propose a novel strategy to scale a generative model across new tasks with minimal compute. During our experiments, we discovered that the variance maps of intermediate feature maps of diffusion models capture the intensity of conditioning. Utilizing this prior information, we propose MaxFusion, an efficient strategy to scale up text-to-image generation models to accommodate new modality conditions. Specifically, we combine aligned features of multiple models, hence bringing a compositional effect. Our fusion strategy can be integrated into off-the-shelf models to enhance their generative prowess.
Diffscaler: Enhancing the Generative Prowess of Diffusion Transformers
Apr 15, 2024Abstract:Recently, diffusion transformers have gained wide attention with its excellent performance in text-to-image and text-to-vidoe models, emphasizing the need for transformers as backbone for diffusion models. Transformer-based models have shown better generalization capability compared to CNN-based models for general vision tasks. However, much less has been explored in the existing literature regarding the capabilities of transformer-based diffusion backbones and expanding their generative prowess to other datasets. This paper focuses on enabling a single pre-trained diffusion transformer model to scale across multiple datasets swiftly, allowing for the completion of diverse generative tasks using just one model. To this end, we propose DiffScaler, an efficient scaling strategy for diffusion models where we train a minimal amount of parameters to adapt to different tasks. In particular, we learn task-specific transformations at each layer by incorporating the ability to utilize the learned subspaces of the pre-trained model, as well as the ability to learn additional task-specific subspaces, which may be absent in the pre-training dataset. As these parameters are independent, a single diffusion model with these task-specific parameters can be used to perform multiple tasks simultaneously. Moreover, we find that transformer-based diffusion models significantly outperform CNN-based diffusion models methods while performing fine-tuning over smaller datasets. We perform experiments on four unconditional image generation datasets. We show that using our proposed method, a single pre-trained model can scale up to perform these conditional and unconditional tasks, respectively, with minimal parameter tuning while performing as close as fine-tuning an entire diffusion model for that particular task.
CheXagent: Towards a Foundation Model for Chest X-Ray Interpretation
Jan 22, 2024



Abstract:Chest X-rays (CXRs) are the most frequently performed imaging test in clinical practice. Recent advances in the development of vision-language foundation models (FMs) give rise to the possibility of performing automated CXR interpretation, which can assist physicians with clinical decision-making and improve patient outcomes. However, developing FMs that can accurately interpret CXRs is challenging due to the (1) limited availability of large-scale vision-language datasets in the medical image domain, (2) lack of vision and language encoders that can capture the complexities of medical data, and (3) absence of evaluation frameworks for benchmarking the abilities of FMs on CXR interpretation. In this work, we address these challenges by first introducing \emph{CheXinstruct} - a large-scale instruction-tuning dataset curated from 28 publicly-available datasets. We then present \emph{CheXagent} - an instruction-tuned FM capable of analyzing and summarizing CXRs. To build CheXagent, we design a clinical large language model (LLM) for parsing radiology reports, a vision encoder for representing CXR images, and a network to bridge the vision and language modalities. Finally, we introduce \emph{CheXbench} - a novel benchmark designed to systematically evaluate FMs across 8 clinically-relevant CXR interpretation tasks. Extensive quantitative evaluations and qualitative reviews with five expert radiologists demonstrate that CheXagent outperforms previously-developed general- and medical-domain FMs on CheXbench tasks. Furthermore, in an effort to improve model transparency, we perform a fairness evaluation across factors of sex, race and age to highlight potential performance disparities. Our project is at \url{https://stanford-aimi.github.io/chexagent.html}.
Disruptive Autoencoders: Leveraging Low-level features for 3D Medical Image Pre-training
Jul 31, 2023Abstract:Harnessing the power of pre-training on large-scale datasets like ImageNet forms a fundamental building block for the progress of representation learning-driven solutions in computer vision. Medical images are inherently different from natural images as they are acquired in the form of many modalities (CT, MR, PET, Ultrasound etc.) and contain granulated information like tissue, lesion, organs etc. These characteristics of medical images require special attention towards learning features representative of local context. In this work, we focus on designing an effective pre-training framework for 3D radiology images. First, we propose a new masking strategy called local masking where the masking is performed across channel embeddings instead of tokens to improve the learning of local feature representations. We combine this with classical low-level perturbations like adding noise and downsampling to further enable low-level representation learning. To this end, we introduce Disruptive Autoencoders, a pre-training framework that attempts to reconstruct the original image from disruptions created by a combination of local masking and low-level perturbations. Additionally, we also devise a cross-modal contrastive loss (CMCL) to accommodate the pre-training of multiple modalities in a single framework. We curate a large-scale dataset to enable pre-training of 3D medical radiology images (MRI and CT). The proposed pre-training framework is tested across multiple downstream tasks and achieves state-of-the-art performance. Notably, our proposed method tops the public test leaderboard of BTCV multi-organ segmentation challenge.
Ambiguous Medical Image Segmentation using Diffusion Models
Apr 10, 2023



Abstract:Collective insights from a group of experts have always proven to outperform an individual's best diagnostic for clinical tasks. For the task of medical image segmentation, existing research on AI-based alternatives focuses more on developing models that can imitate the best individual rather than harnessing the power of expert groups. In this paper, we introduce a single diffusion model-based approach that produces multiple plausible outputs by learning a distribution over group insights. Our proposed model generates a distribution of segmentation masks by leveraging the inherent stochastic sampling process of diffusion using only minimal additional learning. We demonstrate on three different medical image modalities- CT, ultrasound, and MRI that our model is capable of producing several possible variants while capturing the frequencies of their occurrences. Comprehensive results show that our proposed approach outperforms existing state-of-the-art ambiguous segmentation networks in terms of accuracy while preserving naturally occurring variation. We also propose a new metric to evaluate the diversity as well as the accuracy of segmentation predictions that aligns with the interest of clinical practice of collective insights.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge