Sergios Gatidis
A Reasoning-Enabled Vision-Language Foundation Model for Chest X-ray Interpretation
Apr 01, 2026Abstract:Chest X-rays (CXRs) are among the most frequently performed imaging examinations worldwide, yet rising imaging volumes increase radiologist workload and the risk of diagnostic errors. Although artificial intelligence (AI) systems have shown promise for CXR interpretation, most generate only final predictions, without making explicit how visual evidence is translated into radiographic findings and diagnostic predictions. We present CheXOne, a reasoning-enabled vision-language model for CXR interpretation. CheXOne jointly generates diagnostic predictions and explicit, clinically grounded reasoning traces that connect visual evidence, radiographic findings, and these predictions. The model is trained on 14.7 million instruction and reasoning samples curated from 30 public datasets spanning 36 CXR interpretation tasks, using a two-stage framework that combines instruction tuning with reinforcement learning to improve reasoning quality. We evaluate CheXOne in zero-shot settings across visual question answering, report generation, visual grounding and reasoning assessment, covering 17 evaluation settings. CheXOne outperforms existing medical and general-domain foundation models and achieves strong performance on independent public benchmarks. A clinical reader study demonstrates that CheXOne-drafted reports are comparable to or better than resident-written reports in 55% of cases, while effectively addressing clinical indications and enhancing both report writing and CXR interpretation efficiency. Further analyses involving radiologists reveal that the generated reasoning traces show high clinical factuality and provide causal support for the final predictions, offering a plausible explanation for the performance gains. These results suggest that explicit reasoning can improve model performance, interpretability and clinical utility in AI-assisted CXR interpretation.
Sparse Autoencoders for Interpretable Medical Image Representation Learning
Mar 24, 2026Abstract:Vision foundation models (FMs) achieve state-of-the-art performance in medical imaging. However, they encode information in abstract latent representations that clinicians cannot interrogate or verify. The goal of this study is to investigate Sparse Autoencoders (SAEs) for replacing opaque FM image representations with human-interpretable, sparse features. We train SAEs on embeddings from BiomedParse (biomedical) and DINOv3 (general-purpose) using 909,873 CT and MRI 2D image slices from the TotalSegmentator dataset. We find that learned sparse features: (a) reconstruct original embeddings with high fidelity (R2 up to 0.941) and recover up to 87.8% of downstream performance using only 10 features (99.4% dimensionality reduction), (b) preserve semantic fidelity in image retrieval tasks, (c) correspond to specific concepts that can be expressed in language using large language model (LLM)-based auto-interpretation. (d) bridge clinical language and abstract latent representations in zero-shot language-driven image retrieval. Our work indicates SAEs are a promising pathway towards interpretable, concept-driven medical vision systems. Code repository: https://github.com/pwesp/sail.
A data- and compute-efficient chest X-ray foundation model beyond aggressive scaling
Feb 26, 2026Abstract:Foundation models for medical imaging are typically pretrained on increasingly large datasets, following a "scale-at-all-costs" paradigm. However, this strategy faces two critical challenges: large-scale medical datasets often contain substantial redundancy and severe class imbalance that bias representation learning toward over-represented patterns, and indiscriminate training regardless of heterogeneity in data quality incurs considerable computational inefficiency. Here we demonstrate that active, principled data curation during pretraining can serve as a viable, cost-effective alternative to brute-force dataset enlargement. We introduce CheXficient, a chest X-ray (CXR) foundation model that selectively prioritizes informative training samples. CheXficient is pretrained on only 22.7% of 1,235,004 paired CXR images and reports while consuming under 27.3% of the total compute budget, yet achieving comparable or superior performance to its full-data counterpart and other large-scale pretrained models. We assess CheXficient across 20 individual benchmarks spanning 5 task types, including non-adapted off-the-shelf evaluations (zero-shot findings classification and crossmodal retrieval) and adapted downstream tasks (disease prediction, semantic segmentation, and radiology report generation). Further analyses show that CheXficient systematically prioritizes under-represented training samples, improving generalizability on long-tailed or rare conditions. Overall, our work offers practical insights into the data and computation demands for efficient pretraining and downstream adaptation of medical vision-language foundation models.
Unpaired Image-to-Image Translation via a Self-Supervised Semantic Bridge
Feb 18, 2026Abstract:Adversarial diffusion and diffusion-inversion methods have advanced unpaired image-to-image translation, but each faces key limitations. Adversarial approaches require target-domain adversarial loss during training, which can limit generalization to unseen data, while diffusion-inversion methods often produce low-fidelity translations due to imperfect inversion into noise-latent representations. In this work, we propose the Self-Supervised Semantic Bridge (SSB), a versatile framework that integrates external semantic priors into diffusion bridge models to enable spatially faithful translation without cross-domain supervision. Our key idea is to leverage self-supervised visual encoders to learn representations that are invariant to appearance changes but capture geometric structure, forming a shared latent space that conditions the diffusion bridges. Extensive experiments show that SSB outperforms strong prior methods for challenging medical image synthesis in both in-domain and out-of-domain settings, and extends easily to high-quality text-guided editing.
Towards a Unified Theoretical Framework for Self-Supervised MRI Reconstruction
Jan 08, 2026Abstract:The demand for high-resolution, non-invasive imaging continues to drive innovation in magnetic resonance imaging (MRI), yet prolonged acquisition times hinder accessibility and real-time applications. While deep learning-based reconstruction methods have accelerated MRI, their predominant supervised paradigm depends on fully-sampled reference data that are challenging to acquire. Recently, self-supervised learning (SSL) approaches have emerged as promising alternatives, but most are empirically designed and fragmented. Therefore, we introduce UNITS (Unified Theory for Self-supervision), a general framework for self-supervised MRI reconstruction. UNITS unifies prior SSL strategies within a common formalism, enabling consistent interpretation and systematic benchmarking. We prove that SSL can achieve the same expected performance as supervised learning. Under this theoretical guarantee, we introduce sampling stochasticity and flexible data utilization, which improve network generalization under out-of-domain distributions and stabilize training. Together, these contributions establish UNITS as a theoretical foundation and a practical paradigm for interpretable, generalizable, and clinically applicable self-supervised MRI reconstruction.
Retrospective motion correction in MRI using disentangled embeddings
Nov 11, 2025Abstract:Physiological motion can affect the diagnostic quality of magnetic resonance imaging (MRI). While various retrospective motion correction methods exist, many struggle to generalize across different motion types and body regions. In particular, machine learning (ML)-based corrections are often tailored to specific applications and datasets. We hypothesize that motion artifacts, though diverse, share underlying patterns that can be disentangled and exploited. To address this, we propose a hierarchical vector-quantized (VQ) variational auto-encoder that learns a disentangled embedding of motion-to-clean image features. A codebook is deployed to capture finite collection of motion patterns at multiple resolutions, enabling coarse-to-fine correction. An auto-regressive model is trained to learn the prior distribution of motion-free images and is used at inference to guide the correction process. Unlike conventional approaches, our method does not require artifact-specific training and can generalize to unseen motion patterns. We demonstrate the approach on simulated whole-body motion artifacts and observe robust correction across varying motion severity. Our results suggest that the model effectively disentangled physical motion of the simulated motion-effective scans, therefore, improving the generalizability of the ML-based MRI motion correction. Our work of disentangling the motion features shed a light on its potential application across anatomical regions and motion types.
Adaptable Cardiovascular Disease Risk Prediction from Heterogeneous Data using Large Language Models
May 30, 2025Abstract:Cardiovascular disease (CVD) risk prediction models are essential for identifying high-risk individuals and guiding preventive actions. However, existing models struggle with the challenges of real-world clinical practice as they oversimplify patient profiles, rely on rigid input schemas, and are sensitive to distribution shifts. We developed AdaCVD, an adaptable CVD risk prediction framework built on large language models extensively fine-tuned on over half a million participants from the UK Biobank. In benchmark comparisons, AdaCVD surpasses established risk scores and standard machine learning approaches, achieving state-of-the-art performance. Crucially, for the first time, it addresses key clinical challenges across three dimensions: it flexibly incorporates comprehensive yet variable patient information; it seamlessly integrates both structured data and unstructured text; and it rapidly adapts to new patient populations using minimal additional data. In stratified analyses, it demonstrates robust performance across demographic, socioeconomic, and clinical subgroups, including underrepresented cohorts. AdaCVD offers a promising path toward more flexible, AI-driven clinical decision support tools suited to the realities of heterogeneous and dynamic healthcare environments.
Self-supervised feature learning for cardiac Cine MR image reconstruction
May 29, 2025Abstract:We propose a self-supervised feature learning assisted reconstruction (SSFL-Recon) framework for MRI reconstruction to address the limitation of existing supervised learning methods. Although recent deep learning-based methods have shown promising performance in MRI reconstruction, most require fully-sampled images for supervised learning, which is challenging in practice considering long acquisition times under respiratory or organ motion. Moreover, nearly all fully-sampled datasets are obtained from conventional reconstruction of mildly accelerated datasets, thus potentially biasing the achievable performance. The numerous undersampled datasets with different accelerations in clinical practice, hence, remain underutilized. To address these issues, we first train a self-supervised feature extractor on undersampled images to learn sampling-insensitive features. The pre-learned features are subsequently embedded in the self-supervised reconstruction network to assist in removing artifacts. Experiments were conducted retrospectively on an in-house 2D cardiac Cine dataset, including 91 cardiovascular patients and 38 healthy subjects. The results demonstrate that the proposed SSFL-Recon framework outperforms existing self-supervised MRI reconstruction methods and even exhibits comparable or better performance to supervised learning up to $16\times$ retrospective undersampling. The feature learning strategy can effectively extract global representations, which have proven beneficial in removing artifacts and increasing generalization ability during reconstruction.
MedHELM: Holistic Evaluation of Large Language Models for Medical Tasks
May 26, 2025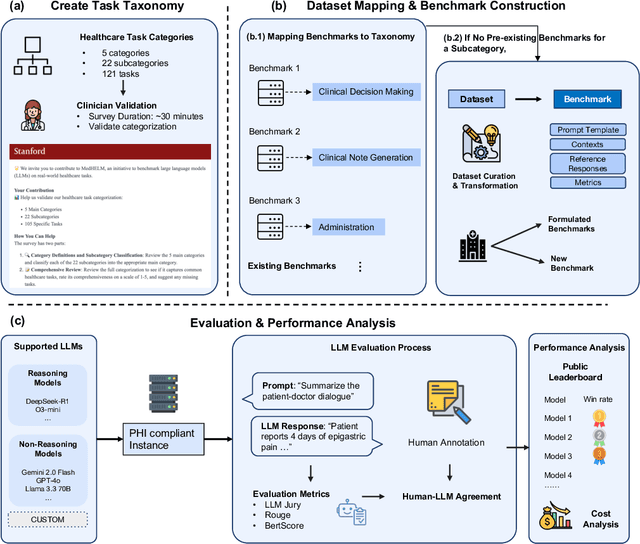
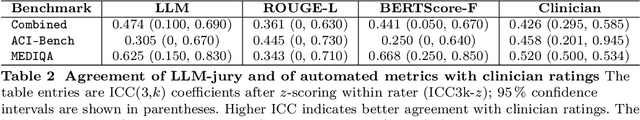
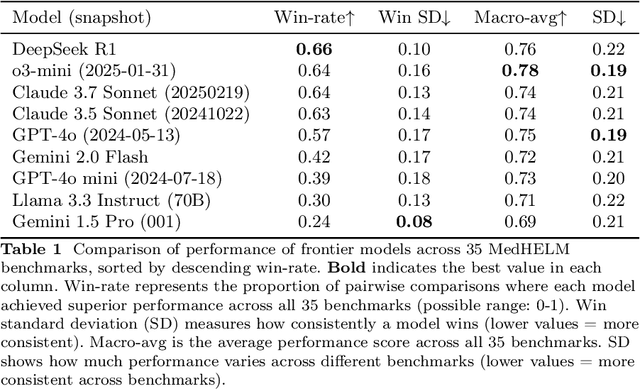
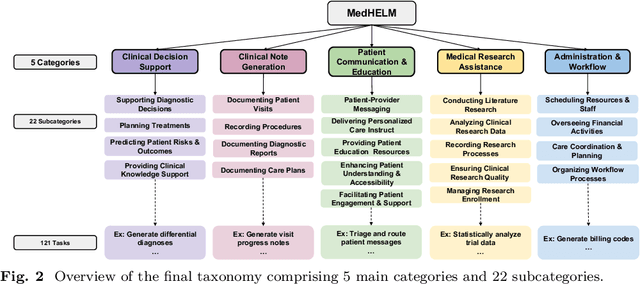

Abstract:While large language models (LLMs) achieve near-perfect scores on medical licensing exams, these evaluations inadequately reflect the complexity and diversity of real-world clinical practice. We introduce MedHELM, an extensible evaluation framework for assessing LLM performance for medical tasks with three key contributions. First, a clinician-validated taxonomy spanning 5 categories, 22 subcategories, and 121 tasks developed with 29 clinicians. Second, a comprehensive benchmark suite comprising 35 benchmarks (17 existing, 18 newly formulated) providing complete coverage of all categories and subcategories in the taxonomy. Third, a systematic comparison of LLMs with improved evaluation methods (using an LLM-jury) and a cost-performance analysis. Evaluation of 9 frontier LLMs, using the 35 benchmarks, revealed significant performance variation. Advanced reasoning models (DeepSeek R1: 66% win-rate; o3-mini: 64% win-rate) demonstrated superior performance, though Claude 3.5 Sonnet achieved comparable results at 40% lower estimated computational cost. On a normalized accuracy scale (0-1), most models performed strongly in Clinical Note Generation (0.73-0.85) and Patient Communication & Education (0.78-0.83), moderately in Medical Research Assistance (0.65-0.75), and generally lower in Clinical Decision Support (0.56-0.72) and Administration & Workflow (0.53-0.63). Our LLM-jury evaluation method achieved good agreement with clinician ratings (ICC = 0.47), surpassing both average clinician-clinician agreement (ICC = 0.43) and automated baselines including ROUGE-L (0.36) and BERTScore-F1 (0.44). Claude 3.5 Sonnet achieved comparable performance to top models at lower estimated cost. These findings highlight the importance of real-world, task-specific evaluation for medical use of LLMs and provides an open source framework to enable this.
Best Practices for Large Language Models in Radiology
Dec 02, 2024



Abstract:At the heart of radiological practice is the challenge of integrating complex imaging data with clinical information to produce actionable insights. Nuanced application of language is key for various activities, including managing requests, describing and interpreting imaging findings in the context of clinical data, and concisely documenting and communicating the outcomes. The emergence of large language models (LLMs) offers an opportunity to improve the management and interpretation of the vast data in radiology. Despite being primarily general-purpose, these advanced computational models demonstrate impressive capabilities in specialized language-related tasks, even without specific training. Unlocking the potential of LLMs for radiology requires basic understanding of their foundations and a strategic approach to navigate their idiosyncrasies. This review, drawing from practical radiology and machine learning expertise and recent literature, provides readers insight into the potential of LLMs in radiology. It examines best practices that have so far stood the test of time in the rapidly evolving landscape of LLMs. This includes practical advice for optimizing LLM characteristics for radiology practices along with limitations, effective prompting, and fine-tuning strategies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge