Eduardo Reis
Comp2Comp: Open-Source Software with FDA-Cleared Artificial Intelligence Algorithms for Computed Tomography Image Analysis
Feb 10, 2026Abstract:Artificial intelligence allows automatic extraction of imaging biomarkers from already-acquired radiologic images. This paradigm of opportunistic imaging adds value to medical imaging without additional imaging costs or patient radiation exposure. However, many open-source image analysis solutions lack rigorous validation while commercial solutions lack transparency, leading to unexpected failures when deployed. Here, we report development and validation for two of the first fully open-sourced, FDA-510(k)-cleared deep learning pipelines to mitigate both challenges: Abdominal Aortic Quantification (AAQ) and Bone Mineral Density (BMD) estimation are both offered within the Comp2Comp package for opportunistic analysis of computed tomography scans. AAQ segments the abdominal aorta to assess aneurysm size; BMD segments vertebral bodies to estimate trabecular bone density and osteoporosis risk. AAQ-derived maximal aortic diameters were compared against radiologist ground-truth measurements on 258 patient scans enriched for abdominal aortic aneurysms from four external institutions. BMD binary classifications (low vs. normal bone density) were compared against concurrent DXA scan ground truths obtained on 371 patient scans from four external institutions. AAQ had an overall mean absolute error of 1.57 mm (95% CI 1.38-1.80 mm). BMD had a sensitivity of 81.0% (95% CI 74.0-86.8%) and specificity of 78.4% (95% CI 72.3-83.7%). Comp2Comp AAQ and BMD demonstrated sufficient accuracy for clinical use. Open-sourcing these algorithms improves transparency of typically opaque FDA clearance processes, allows hospitals to test the algorithms before cumbersome clinical pilots, and provides researchers with best-in-class methods.
MIRIAD: Augmenting LLMs with millions of medical query-response pairs
Jun 06, 2025


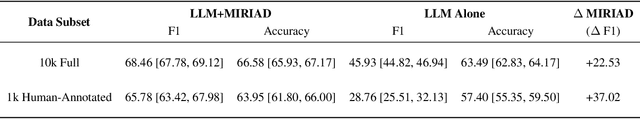
Abstract:LLMs are bound to transform healthcare with advanced decision support and flexible chat assistants. However, LLMs are prone to generate inaccurate medical content. To ground LLMs in high-quality medical knowledge, LLMs have been equipped with external knowledge via RAG, where unstructured medical knowledge is split into small text chunks that can be selectively retrieved and integrated into the LLMs context. Yet, existing RAG pipelines rely on raw, unstructured medical text, which can be noisy, uncurated and difficult for LLMs to effectively leverage. Systematic approaches to organize medical knowledge to best surface it to LLMs are generally lacking. To address these challenges, we introduce MIRIAD, a large-scale, curated corpus of 5,821,948 medical QA pairs, each rephrased from and grounded in a passage from peer-reviewed medical literature using a semi-automated pipeline combining LLM generation, filtering, grounding, and human annotation. Unlike prior medical corpora, which rely on unstructured text, MIRIAD encapsulates web-scale medical knowledge in an operationalized query-response format, which enables more targeted retrieval. Experiments on challenging medical QA benchmarks show that augmenting LLMs with MIRIAD improves accuracy up to 6.7% compared to unstructured RAG baselines with the same source corpus and with the same amount of retrieved text. Moreover, MIRIAD improved the ability of LLMs to detect medical hallucinations by 22.5 to 37% (increase in F1 score). We further introduce MIRIAD-Atlas, an interactive map of MIRIAD spanning 56 medical disciplines, enabling clinical users to visually explore, search, and refine medical knowledge. MIRIAD promises to unlock a wealth of down-stream applications, including medical information retrievers, enhanced RAG applications, and knowledge-grounded chat interfaces, which ultimately enables more reliable LLM applications in healthcare.
Automated Structured Radiology Report Generation
May 30, 2025Abstract:Automated radiology report generation from chest X-ray (CXR) images has the potential to improve clinical efficiency and reduce radiologists' workload. However, most datasets, including the publicly available MIMIC-CXR and CheXpert Plus, consist entirely of free-form reports, which are inherently variable and unstructured. This variability poses challenges for both generation and evaluation: existing models struggle to produce consistent, clinically meaningful reports, and standard evaluation metrics fail to capture the nuances of radiological interpretation. To address this, we introduce Structured Radiology Report Generation (SRRG), a new task that reformulates free-text radiology reports into a standardized format, ensuring clarity, consistency, and structured clinical reporting. We create a novel dataset by restructuring reports using large language models (LLMs) following strict structured reporting desiderata. Additionally, we introduce SRR-BERT, a fine-grained disease classification model trained on 55 labels, enabling more precise and clinically informed evaluation of structured reports. To assess report quality, we propose F1-SRR-BERT, a metric that leverages SRR-BERT's hierarchical disease taxonomy to bridge the gap between free-text variability and structured clinical reporting. We validate our dataset through a reader study conducted by five board-certified radiologists and extensive benchmarking experiments.
MedHELM: Holistic Evaluation of Large Language Models for Medical Tasks
May 26, 2025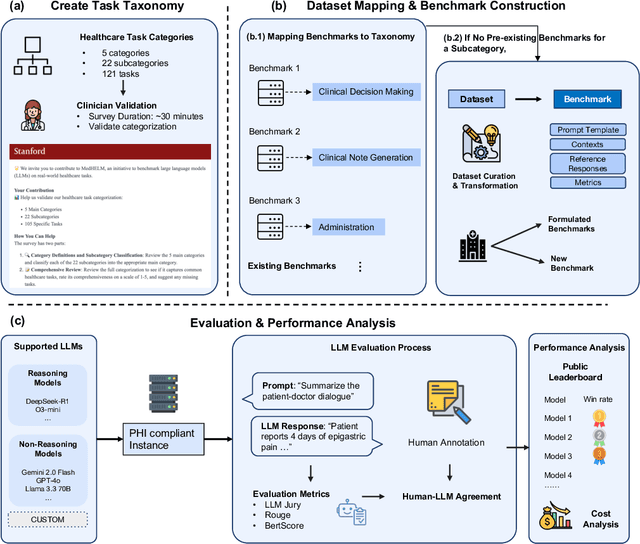
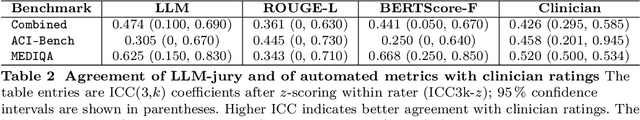
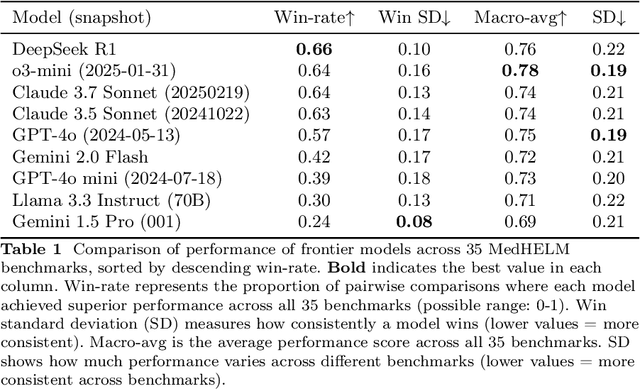
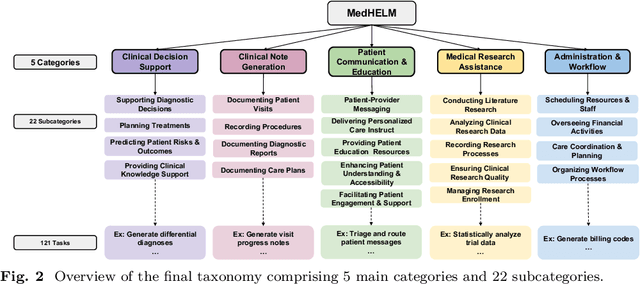

Abstract:While large language models (LLMs) achieve near-perfect scores on medical licensing exams, these evaluations inadequately reflect the complexity and diversity of real-world clinical practice. We introduce MedHELM, an extensible evaluation framework for assessing LLM performance for medical tasks with three key contributions. First, a clinician-validated taxonomy spanning 5 categories, 22 subcategories, and 121 tasks developed with 29 clinicians. Second, a comprehensive benchmark suite comprising 35 benchmarks (17 existing, 18 newly formulated) providing complete coverage of all categories and subcategories in the taxonomy. Third, a systematic comparison of LLMs with improved evaluation methods (using an LLM-jury) and a cost-performance analysis. Evaluation of 9 frontier LLMs, using the 35 benchmarks, revealed significant performance variation. Advanced reasoning models (DeepSeek R1: 66% win-rate; o3-mini: 64% win-rate) demonstrated superior performance, though Claude 3.5 Sonnet achieved comparable results at 40% lower estimated computational cost. On a normalized accuracy scale (0-1), most models performed strongly in Clinical Note Generation (0.73-0.85) and Patient Communication & Education (0.78-0.83), moderately in Medical Research Assistance (0.65-0.75), and generally lower in Clinical Decision Support (0.56-0.72) and Administration & Workflow (0.53-0.63). Our LLM-jury evaluation method achieved good agreement with clinician ratings (ICC = 0.47), surpassing both average clinician-clinician agreement (ICC = 0.43) and automated baselines including ROUGE-L (0.36) and BERTScore-F1 (0.44). Claude 3.5 Sonnet achieved comparable performance to top models at lower estimated cost. These findings highlight the importance of real-world, task-specific evaluation for medical use of LLMs and provides an open source framework to enable this.
Merlin: A Vision Language Foundation Model for 3D Computed Tomography
Jun 10, 2024



Abstract:Over 85 million computed tomography (CT) scans are performed annually in the US, of which approximately one quarter focus on the abdomen. Given the current radiologist shortage, there is a large impetus to use artificial intelligence to alleviate the burden of interpreting these complex imaging studies. Prior state-of-the-art approaches for automated medical image interpretation leverage vision language models (VLMs). However, current medical VLMs are generally limited to 2D images and short reports, and do not leverage electronic health record (EHR) data for supervision. We introduce Merlin - a 3D VLM that we train using paired CT scans (6+ million images from 15,331 CTs), EHR diagnosis codes (1.8+ million codes), and radiology reports (6+ million tokens). We evaluate Merlin on 6 task types and 752 individual tasks. The non-adapted (off-the-shelf) tasks include zero-shot findings classification (31 findings), phenotype classification (692 phenotypes), and zero-shot cross-modal retrieval (image to findings and image to impressions), while model adapted tasks include 5-year disease prediction (6 diseases), radiology report generation, and 3D semantic segmentation (20 organs). We perform internal validation on a test set of 5,137 CTs, and external validation on 7,000 clinical CTs and on two public CT datasets (VerSe, TotalSegmentator). Beyond these clinically-relevant evaluations, we assess the efficacy of various network architectures and training strategies to depict that Merlin has favorable performance to existing task-specific baselines. We derive data scaling laws to empirically assess training data needs for requisite downstream task performance. Furthermore, unlike conventional VLMs that require hundreds of GPUs for training, we perform all training on a single GPU.
Phasor-Driven Acceleration for FFT-based CNNs
Jun 01, 2024



Abstract:Recent research in deep learning (DL) has investigated the use of the Fast Fourier Transform (FFT) to accelerate the computations involved in Convolutional Neural Networks (CNNs) by replacing spatial convolution with element-wise multiplications on the spectral domain. These approaches mainly rely on the FFT to reduce the number of operations, which can be further decreased by adopting the Real-Valued FFT. In this paper, we propose using the phasor form, a polar representation of complex numbers, as a more efficient alternative to the traditional approach. The experimental results, evaluated on the CIFAR-10, demonstrate that our method achieves superior speed improvements of up to a factor of 1.376 (average of 1.316) during training and up to 1.390 (average of 1.321) during inference when compared to the traditional rectangular form employed in modern CNN architectures. Similarly, when evaluated on the CIFAR-100, our method achieves superior speed improvements of up to a factor of 1.375 (average of 1.299) during training and up to 1.387 (average of 1.300) during inference. Most importantly, given the modular aspect of our approach, the proposed method can be applied to any existing convolution-based DL model without design changes.
AgentClinic: a multimodal agent benchmark to evaluate AI in simulated clinical environments
May 13, 2024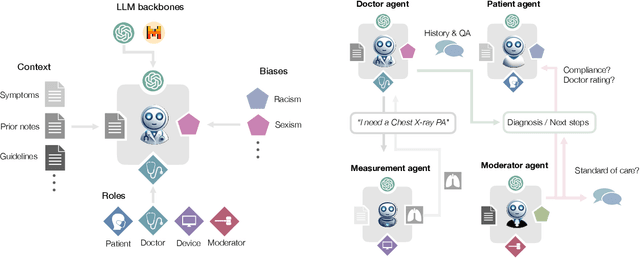
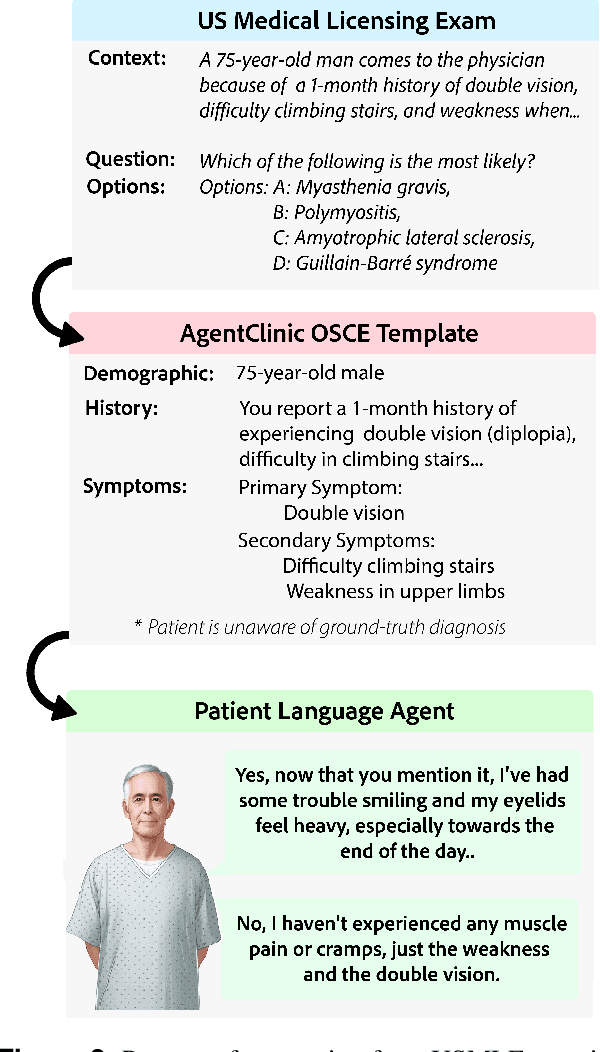
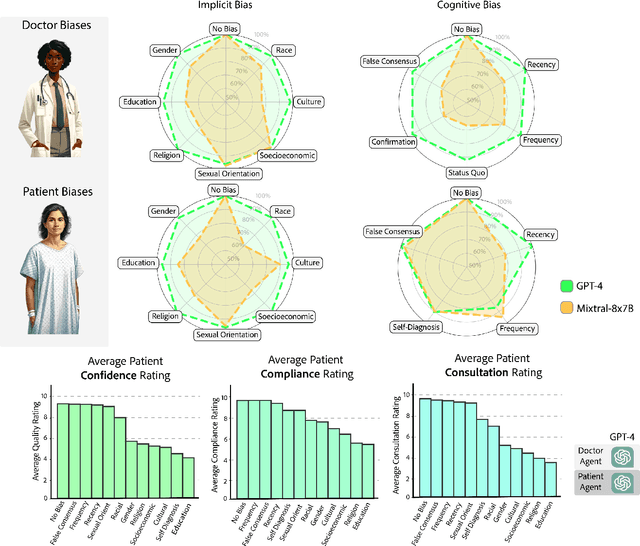

Abstract:Diagnosing and managing a patient is a complex, sequential decision making process that requires physicians to obtain information -- such as which tests to perform -- and to act upon it. Recent advances in artificial intelligence (AI) and large language models (LLMs) promise to profoundly impact clinical care. However, current evaluation schemes overrely on static medical question-answering benchmarks, falling short on interactive decision-making that is required in real-life clinical work. Here, we present AgentClinic: a multimodal benchmark to evaluate LLMs in their ability to operate as agents in simulated clinical environments. In our benchmark, the doctor agent must uncover the patient's diagnosis through dialogue and active data collection. We present two open benchmarks: a multimodal image and dialogue environment, AgentClinic-NEJM, and a dialogue-only environment, AgentClinic-MedQA. We embed cognitive and implicit biases both in patient and doctor agents to emulate realistic interactions between biased agents. We find that introducing bias leads to large reductions in diagnostic accuracy of the doctor agents, as well as reduced compliance, confidence, and follow-up consultation willingness in patient agents. Evaluating a suite of state-of-the-art LLMs, we find that several models that excel in benchmarks like MedQA are performing poorly in AgentClinic-MedQA. We find that the LLM used in the patient agent is an important factor for performance in the AgentClinic benchmark. We show that both having limited interactions as well as too many interaction reduces diagnostic accuracy in doctor agents. The code and data for this work is publicly available at https://AgentClinic.github.io.
Federated Learning on Heterogenous Data using Chest CT
Mar 23, 2023Abstract:Large data have accelerated advances in AI. While it is well known that population differences from genetics, sex, race, diet, and various environmental factors contribute significantly to disease, AI studies in medicine have largely focused on locoregional patient cohorts with less diverse data sources. Such limitation stems from barriers to large-scale data share in medicine and ethical concerns over data privacy. Federated learning (FL) is one potential pathway for AI development that enables learning across hospitals without data share. In this study, we show the results of various FL strategies on one of the largest and most diverse COVID-19 chest CT datasets: 21 participating hospitals across five continents that comprise >10,000 patients with >1 million images. We present three techniques: Fed Averaging (FedAvg), Incremental Institutional Learning (IIL), and Cyclical Incremental Institutional Learning (CIIL). We also propose an FL strategy that leverages synthetically generated data to overcome class imbalances and data size disparities across centers. We show that FL can achieve comparable performance to Centralized Data Sharing (CDS) while maintaining high performance across sites with small, underrepresented data. We investigate the strengths and weaknesses for all technical approaches on this heterogeneous dataset including the robustness to non-Independent and identically distributed (non-IID) diversity of data. We also describe the sources of data heterogeneity such as age, sex, and site locations in the context of FL and show how even among the correctly labeled populations, disparities can arise due to these biases.
Comp2Comp: Open-Source Body Composition Assessment on Computed Tomography
Feb 13, 2023Abstract:Computed tomography (CT) is routinely used in clinical practice to evaluate a wide variety of medical conditions. While CT scans provide diagnoses, they also offer the ability to extract quantitative body composition metrics to analyze tissue volume and quality. Extracting quantitative body composition measures manually from CT scans is a cumbersome and time-consuming task. Proprietary software has been developed recently to automate this process, but the closed-source nature impedes widespread use. There is a growing need for fully automated body composition software that is more accessible and easier to use, especially for clinicians and researchers who are not experts in medical image processing. To this end, we have built Comp2Comp, an open-source Python package for rapid and automated body composition analysis of CT scans. This package offers models, post-processing heuristics, body composition metrics, automated batching, and polychromatic visualizations. Comp2Comp currently computes body composition measures for bone, skeletal muscle, visceral adipose tissue, and subcutaneous adipose tissue on CT scans of the abdomen. We have created two pipelines for this purpose. The first pipeline computes vertebral measures, as well as muscle and adipose tissue measures, at the T12 - L5 vertebral levels from abdominal CT scans. The second pipeline computes muscle and adipose tissue measures on user-specified 2D axial slices. In this guide, we discuss the architecture of the Comp2Comp pipelines, provide usage instructions, and report internal and external validation results to measure the quality of segmentations and body composition measures. Comp2Comp can be found at https://github.com/StanfordMIMI/Comp2Comp.
DiffML: End-to-end Differentiable ML Pipelines
Jul 05, 2022



Abstract:In this paper, we present our vision of differentiable ML pipelines called DiffML to automate the construction of ML pipelines in an end-to-end fashion. The idea is that DiffML allows to jointly train not just the ML model itself but also the entire pipeline including data preprocessing steps, e.g., data cleaning, feature selection, etc. Our core idea is to formulate all pipeline steps in a differentiable way such that the entire pipeline can be trained using backpropagation. However, this is a non-trivial problem and opens up many new research questions. To show the feasibility of this direction, we demonstrate initial ideas and a general principle of how typical preprocessing steps such as data cleaning, feature selection and dataset selection can be formulated as differentiable programs and jointly learned with the ML model. Moreover, we discuss a research roadmap and core challenges that have to be systematically tackled to enable fully differentiable ML pipelines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge