Jean-Benoit Delbrouck
Improving the Performance of Radiology Report De-identification with Large-Scale Training and Benchmarking Against Cloud Vendor Methods
Nov 06, 2025


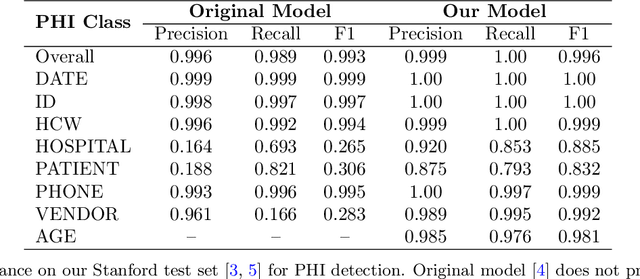
Abstract:Objective: To enhance automated de-identification of radiology reports by scaling transformer-based models through extensive training datasets and benchmarking performance against commercial cloud vendor systems for protected health information (PHI) detection. Materials and Methods: In this retrospective study, we built upon a state-of-the-art, transformer-based, PHI de-identification pipeline by fine-tuning on two large annotated radiology corpora from Stanford University, encompassing chest X-ray, chest CT, abdomen/pelvis CT, and brain MR reports and introducing an additional PHI category (AGE) into the architecture. Model performance was evaluated on test sets from Stanford and the University of Pennsylvania (Penn) for token-level PHI detection. We further assessed (1) the stability of synthetic PHI generation using a "hide-in-plain-sight" method and (2) performance against commercial systems. Precision, recall, and F1 scores were computed across all PHI categories. Results: Our model achieved overall F1 scores of 0.973 on the Penn dataset and 0.996 on the Stanford dataset, outperforming or maintaining the previous state-of-the-art model performance. Synthetic PHI evaluation showed consistent detectability (overall F1: 0.959 [0.958-0.960]) across 50 independently de-identified Penn datasets. Our model outperformed all vendor systems on synthetic Penn reports (overall F1: 0.960 vs. 0.632-0.754). Discussion: Large-scale, multimodal training improved cross-institutional generalization and robustness. Synthetic PHI generation preserved data utility while ensuring privacy. Conclusion: A transformer-based de-identification model trained on diverse radiology datasets outperforms prior academic and commercial systems in PHI detection and establishes a new benchmark for secure clinical text processing.
SMMILE: An Expert-Driven Benchmark for Multimodal Medical In-Context Learning
Jun 26, 2025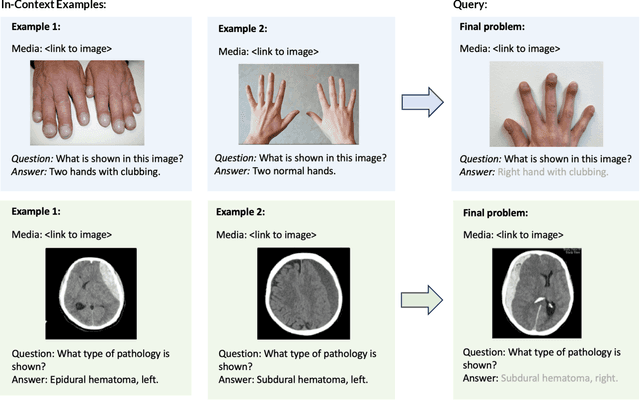



Abstract:Multimodal in-context learning (ICL) remains underexplored despite significant potential for domains such as medicine. Clinicians routinely encounter diverse, specialized tasks requiring adaptation from limited examples, such as drawing insights from a few relevant prior cases or considering a constrained set of differential diagnoses. While multimodal large language models (MLLMs) have shown advances in medical visual question answering (VQA), their ability to learn multimodal tasks from context is largely unknown. We introduce SMMILE, the first expert-driven multimodal ICL benchmark for medical tasks. Eleven medical experts curated problems, each including a multimodal query and multimodal in-context examples as task demonstrations. SMMILE encompasses 111 problems (517 question-image-answer triplets) covering 6 medical specialties and 13 imaging modalities. We further introduce SMMILE++, an augmented variant with 1038 permuted problems. A comprehensive evaluation of 15 MLLMs demonstrates that most models exhibit moderate to poor multimodal ICL ability in medical tasks. In open-ended evaluations, ICL contributes only 8% average improvement over zero-shot on SMMILE and 9.4% on SMMILE++. We observe a susceptibility for irrelevant in-context examples: even a single noisy or irrelevant example can degrade performance by up to 9.5%. Moreover, example ordering exhibits a recency bias, i.e., placing the most relevant example last can lead to substantial performance improvements by up to 71%. Our findings highlight critical limitations and biases in current MLLMs when learning multimodal medical tasks from context.
Automated Structured Radiology Report Generation
May 30, 2025Abstract:Automated radiology report generation from chest X-ray (CXR) images has the potential to improve clinical efficiency and reduce radiologists' workload. However, most datasets, including the publicly available MIMIC-CXR and CheXpert Plus, consist entirely of free-form reports, which are inherently variable and unstructured. This variability poses challenges for both generation and evaluation: existing models struggle to produce consistent, clinically meaningful reports, and standard evaluation metrics fail to capture the nuances of radiological interpretation. To address this, we introduce Structured Radiology Report Generation (SRRG), a new task that reformulates free-text radiology reports into a standardized format, ensuring clarity, consistency, and structured clinical reporting. We create a novel dataset by restructuring reports using large language models (LLMs) following strict structured reporting desiderata. Additionally, we introduce SRR-BERT, a fine-grained disease classification model trained on 55 labels, enabling more precise and clinically informed evaluation of structured reports. To assess report quality, we propose F1-SRR-BERT, a metric that leverages SRR-BERT's hierarchical disease taxonomy to bridge the gap between free-text variability and structured clinical reporting. We validate our dataset through a reader study conducted by five board-certified radiologists and extensive benchmarking experiments.
RaVL: Discovering and Mitigating Spurious Correlations in Fine-Tuned Vision-Language Models
Nov 06, 2024



Abstract:Fine-tuned vision-language models (VLMs) often capture spurious correlations between image features and textual attributes, resulting in degraded zero-shot performance at test time. Existing approaches for addressing spurious correlations (i) primarily operate at the global image-level rather than intervening directly on fine-grained image features and (ii) are predominantly designed for unimodal settings. In this work, we present RaVL, which takes a fine-grained perspective on VLM robustness by discovering and mitigating spurious correlations using local image features rather than operating at the global image level. Given a fine-tuned VLM, RaVL first discovers spurious correlations by leveraging a region-level clustering approach to identify precise image features contributing to zero-shot classification errors. Then, RaVL mitigates the identified spurious correlation with a novel region-aware loss function that enables the VLM to focus on relevant regions and ignore spurious relationships during fine-tuning. We evaluate RaVL on 654 VLMs with various model architectures, data domains, and learned spurious correlations. Our results show that RaVL accurately discovers (191% improvement over the closest baseline) and mitigates (8.2% improvement on worst-group image classification accuracy) spurious correlations. Qualitative evaluations on general-domain and medical-domain VLMs confirm our findings.
Overview of the First Shared Task on Clinical Text Generation: RRG24 and "Discharge Me!"
Sep 25, 2024



Abstract:Recent developments in natural language generation have tremendous implications for healthcare. For instance, state-of-the-art systems could automate the generation of sections in clinical reports to alleviate physician workload and streamline hospital documentation. To explore these applications, we present a shared task consisting of two subtasks: (1) Radiology Report Generation (RRG24) and (2) Discharge Summary Generation ("Discharge Me!"). RRG24 involves generating the 'Findings' and 'Impression' sections of radiology reports given chest X-rays. "Discharge Me!" involves generating the 'Brief Hospital Course' and 'Discharge Instructions' sections of discharge summaries for patients admitted through the emergency department. "Discharge Me!" submissions were subsequently reviewed by a team of clinicians. Both tasks emphasize the goal of reducing clinician burnout and repetitive workloads by generating documentation. We received 201 submissions from across 8 teams for RRG24, and 211 submissions from across 16 teams for "Discharge Me!".
* ACL Proceedings. BioNLP workshop
Merlin: A Vision Language Foundation Model for 3D Computed Tomography
Jun 10, 2024



Abstract:Over 85 million computed tomography (CT) scans are performed annually in the US, of which approximately one quarter focus on the abdomen. Given the current radiologist shortage, there is a large impetus to use artificial intelligence to alleviate the burden of interpreting these complex imaging studies. Prior state-of-the-art approaches for automated medical image interpretation leverage vision language models (VLMs). However, current medical VLMs are generally limited to 2D images and short reports, and do not leverage electronic health record (EHR) data for supervision. We introduce Merlin - a 3D VLM that we train using paired CT scans (6+ million images from 15,331 CTs), EHR diagnosis codes (1.8+ million codes), and radiology reports (6+ million tokens). We evaluate Merlin on 6 task types and 752 individual tasks. The non-adapted (off-the-shelf) tasks include zero-shot findings classification (31 findings), phenotype classification (692 phenotypes), and zero-shot cross-modal retrieval (image to findings and image to impressions), while model adapted tasks include 5-year disease prediction (6 diseases), radiology report generation, and 3D semantic segmentation (20 organs). We perform internal validation on a test set of 5,137 CTs, and external validation on 7,000 clinical CTs and on two public CT datasets (VerSe, TotalSegmentator). Beyond these clinically-relevant evaluations, we assess the efficacy of various network architectures and training strategies to depict that Merlin has favorable performance to existing task-specific baselines. We derive data scaling laws to empirically assess training data needs for requisite downstream task performance. Furthermore, unlike conventional VLMs that require hundreds of GPUs for training, we perform all training on a single GPU.
CheXpert Plus: Augmenting a Large Chest X-ray Dataset with Text Radiology Reports, Patient Demographics and Additional Image Formats
Jun 03, 2024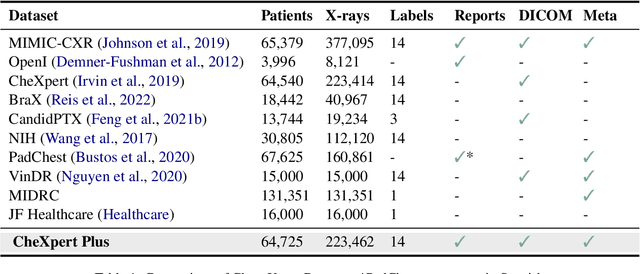

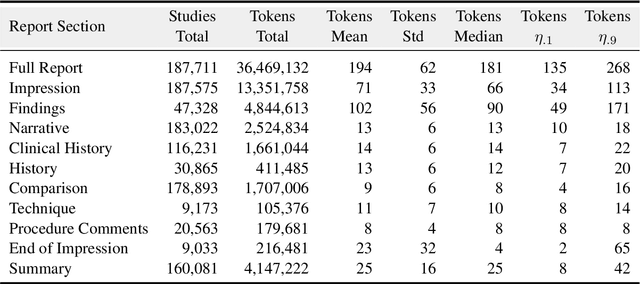
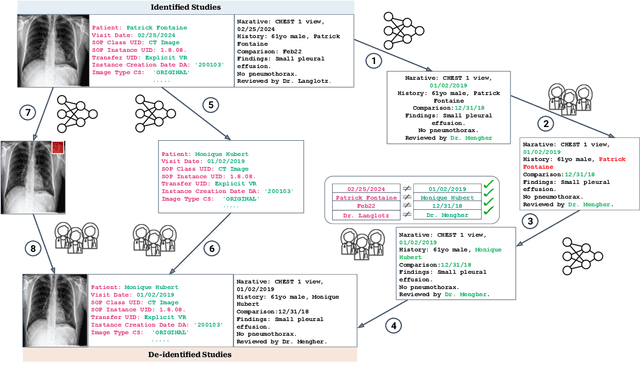
Abstract:Since the release of the original CheXpert paper five years ago, CheXpert has become one of the most widely used and cited clinical AI datasets. The emergence of vision language models has sparked an increase in demands for sharing reports linked to CheXpert images, along with a growing interest among AI fairness researchers in obtaining demographic data. To address this, CheXpert Plus serves as a new collection of radiology data sources, made publicly available to enhance the scaling, performance, robustness, and fairness of models for all subsequent machine learning tasks in the field of radiology. CheXpert Plus is the largest text dataset publicly released in radiology, with a total of 36 million text tokens, including 13 million impression tokens. To the best of our knowledge, it represents the largest text de-identification effort in radiology, with almost 1 million PHI spans anonymized. It is only the second time that a large-scale English paired dataset has been released in radiology, thereby enabling, for the first time, cross-institution training at scale. All reports are paired with high-quality images in DICOM format, along with numerous image and patient metadata covering various clinical and socio-economic groups, as well as many pathology labels and RadGraph annotations. We hope this dataset will boost research for AI models that can further assist radiologists and help improve medical care. Data is available at the following URL: https://stanfordaimi.azurewebsites.net/datasets/5158c524-d3ab-4e02-96e9-6ee9efc110a1 Models are available at the following URL: https://github.com/Stanford-AIMI/chexpert-plus
GREEN: Generative Radiology Report Evaluation and Error Notation
May 06, 2024
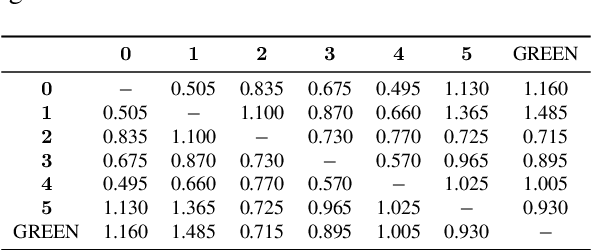


Abstract:Evaluating radiology reports is a challenging problem as factual correctness is extremely important due to the need for accurate medical communication about medical images. Existing automatic evaluation metrics either suffer from failing to consider factual correctness (e.g., BLEU and ROUGE) or are limited in their interpretability (e.g., F1CheXpert and F1RadGraph). In this paper, we introduce GREEN (Generative Radiology Report Evaluation and Error Notation), a radiology report generation metric that leverages the natural language understanding of language models to identify and explain clinically significant errors in candidate reports, both quantitatively and qualitatively. Compared to current metrics, GREEN offers: 1) a score aligned with expert preferences, 2) human interpretable explanations of clinically significant errors, enabling feedback loops with end-users, and 3) a lightweight open-source method that reaches the performance of commercial counterparts. We validate our GREEN metric by comparing it to GPT-4, as well as to error counts of 6 experts and preferences of 2 experts. Our method demonstrates not only higher correlation with expert error counts, but simultaneously higher alignment with expert preferences when compared to previous approaches."
CheXagent: Towards a Foundation Model for Chest X-Ray Interpretation
Jan 22, 2024



Abstract:Chest X-rays (CXRs) are the most frequently performed imaging test in clinical practice. Recent advances in the development of vision-language foundation models (FMs) give rise to the possibility of performing automated CXR interpretation, which can assist physicians with clinical decision-making and improve patient outcomes. However, developing FMs that can accurately interpret CXRs is challenging due to the (1) limited availability of large-scale vision-language datasets in the medical image domain, (2) lack of vision and language encoders that can capture the complexities of medical data, and (3) absence of evaluation frameworks for benchmarking the abilities of FMs on CXR interpretation. In this work, we address these challenges by first introducing \emph{CheXinstruct} - a large-scale instruction-tuning dataset curated from 28 publicly-available datasets. We then present \emph{CheXagent} - an instruction-tuned FM capable of analyzing and summarizing CXRs. To build CheXagent, we design a clinical large language model (LLM) for parsing radiology reports, a vision encoder for representing CXR images, and a network to bridge the vision and language modalities. Finally, we introduce \emph{CheXbench} - a novel benchmark designed to systematically evaluate FMs across 8 clinically-relevant CXR interpretation tasks. Extensive quantitative evaluations and qualitative reviews with five expert radiologists demonstrate that CheXagent outperforms previously-developed general- and medical-domain FMs on CheXbench tasks. Furthermore, in an effort to improve model transparency, we perform a fairness evaluation across factors of sex, race and age to highlight potential performance disparities. Our project is at \url{https://stanford-aimi.github.io/chexagent.html}.
Clinical Text Summarization: Adapting Large Language Models Can Outperform Human Experts
Sep 14, 2023Abstract:Sifting through vast textual data and summarizing key information imposes a substantial burden on how clinicians allocate their time. Although large language models (LLMs) have shown immense promise in natural language processing (NLP) tasks, their efficacy across diverse clinical summarization tasks has not yet been rigorously examined. In this work, we employ domain adaptation methods on eight LLMs, spanning six datasets and four distinct summarization tasks: radiology reports, patient questions, progress notes, and doctor-patient dialogue. Our thorough quantitative assessment reveals trade-offs between models and adaptation methods in addition to instances where recent advances in LLMs may not lead to improved results. Further, in a clinical reader study with six physicians, we depict that summaries from the best adapted LLM are preferable to human summaries in terms of completeness and correctness. Our ensuing qualitative analysis delineates mutual challenges faced by both LLMs and human experts. Lastly, we correlate traditional quantitative NLP metrics with reader study scores to enhance our understanding of how these metrics align with physician preferences. Our research marks the first evidence of LLMs outperforming human experts in clinical text summarization across multiple tasks. This implies that integrating LLMs into clinical workflows could alleviate documentation burden, empowering clinicians to focus more on personalized patient care and other irreplaceable human aspects of medicine.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge