Pengcheng Shi
for the ALFA study
Federated Distributional Reinforcement Learning with Distributional Critic Regularization
Mar 18, 2026Abstract:Federated reinforcement learning typically aggregates value functions or policies by parameter averaging, which emphasizes expected return and can obscure statistical multimodality and tail behavior that matter in safety-critical settings. We formalize federated distributional reinforcement learning (FedDistRL), where clients parametrize quantile value function critics and federate these networks only. We also propose TR-FedDistRL, which builds a per client, risk-aware Wasserstein barycenter over a temporal buffer. This local barycenter provides a reference region to constrain the parameter averaged critic, ensuring necessary distributional information is not averaged out during the federation process. The distributional trust region is implemented as a shrink-squash step around this reference. Under fixed-policy evaluation, the feasibility map is nonexpansive and the update is contractive in a probe-set Wasserstein metric under evaluation. Experiments on a bandit, multi-agent gridworld, and continuous highway environment show reduced mean-smearing, improved safety proxies (catastrophe/accident rate), and lower critic/policy drift versus mean-oriented and non-federated baselines.
Global Cross-Modal Geo-Localization: A Million-Scale Dataset and a Physical Consistency Learning Framework
Mar 09, 2026Abstract:Cross-modal Geo-localization (CMGL) matches ground-level text descriptions with geo-tagged aerial imagery, which is crucial for pedestrian navigation and emergency response. However, existing researches are constrained by narrow geographic coverage and simplistic scene diversity, failing to reflect the immense spatial heterogeneity of global architectural styles and topographic features. To bridge this gap and facilitate universal positioning, we introduce CORE, the first million-scale dataset dedicated to global CMGL. CORE comprises 1,034,786 cross-view images sampled from 225 distinct geographic regions across all continents, offering an unprecedented variety of perspectives in varying environmental conditions and urban layouts. We leverage the zero-shot reasoning of Large Vision-Language Models (LVLMs) to synthesize high-quality scene descriptions rich in discriminative cues. Furthermore, we propose a physical-law-aware network (PLANET) for cross-modal geo-localization. PLANET introduces a novel contrastive learning paradigm to guide textual representations in capturing the intrinsic physical signatures of satellite imagery. Extensive experiments across varied geographic regions demonstrate that PLANet significantly outperforms state-of-the-art methods, establishing a new benchmark for robust, global-scale geo-localization. The dataset and source code will be released at https://github.com/YtH0823/CORE.
U-VLM: Hierarchical Vision Language Modeling for Report Generation
Feb 28, 2026Abstract:Automated radiology report generation is key for reducing radiologist workload and improving diagnostic consistency, yet generating accurate reports for 3D medical imaging remains challenging. Existing vision-language models face two limitations: they do not leverage segmentation-pretrained encoders, and they inject visual features only at the input layer of language models, losing multi-scale information. We propose U-VLM, which enables hierarchical vision-language modeling in both training and architecture: (1) progressive training from segmentation to classification to report generation, and (2) multi-layer visual injection that routes U-Net encoder features to corresponding language model layers. Each training stage can leverage different datasets without unified annotations. U-VLM achieves state-of-the-art performance on CT-RATE (F1: 0.414 vs 0.258, BLEU-mean: 0.349 vs 0.305) and AbdomenAtlas 3.0 (F1: 0.624 vs 0.518 for segmentation-based detection) using only a 0.1B decoder trained from scratch, demonstrating that well-designed vision encoder pretraining outweighs the benefits of 7B+ pre-trained language models. Ablation studies show that progressive pretraining significantly improves F1, while multi-layer injection improves BLEU-mean. Code is available at https://github.com/yinghemedical/U-VLM.
Hierarchical Semantic Learning for Multi-Class Aorta Segmentation
Nov 18, 2025Abstract:The aorta, the body's largest artery, is prone to pathologies such as dissection, aneurysm, and atherosclerosis, which often require timely intervention. Minimally invasive repairs involving branch vessels necessitate detailed 3D anatomical analysis. Existing methods often overlook hierarchical anatomical relationships while struggling with severe class imbalance inherent in vascular structures. We address these challenges with a curriculum learning strategy that leverages a novel fractal softmax for hierarchical semantic learning. Inspired by human cognition, our approach progressively learns anatomical constraints by decomposing complex structures from simple to complex components. The curriculum learning framework naturally addresses class imbalance by first establishing robust feature representations for dominant classes before tackling rare but anatomically critical structures, significantly accelerating model convergence in multi-class scenarios. Our two-stage inference strategy achieves up to fivefold acceleration, enhancing clinical practicality. On the validation set at epoch 50, our hierarchical semantic loss improves the Dice score of nnU-Net ResEnc M by 11.65%. The proposed model demonstrates a 5.6% higher Dice score than baselines on the test set. Experimental results show significant improvements in segmentation accuracy and efficiency, making the framework suitable for real-time clinical applications. The implementation code for this challenge entry is publicly available at: https://github.com/PengchengShi1220/AortaSeg24. The code for fractal softmax will be available at https://github.com/PengchengShi1220/fractal-softmax.
Medal S: Spatio-Textual Prompt Model for Medical Segmentation
Nov 17, 2025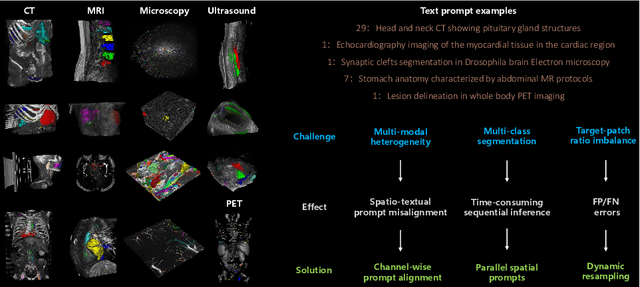
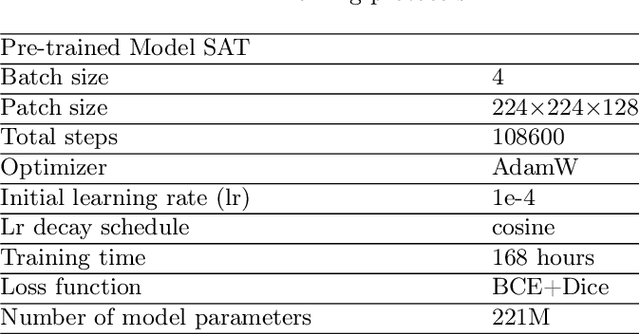
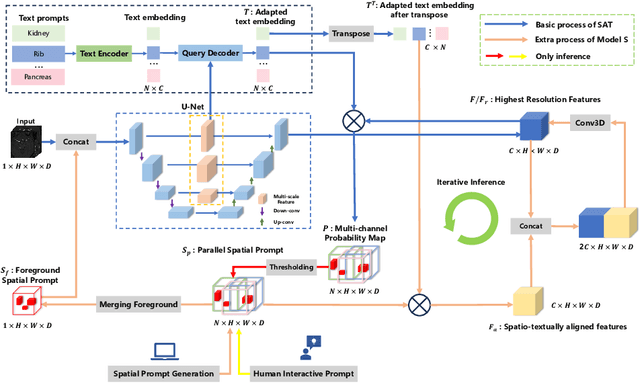
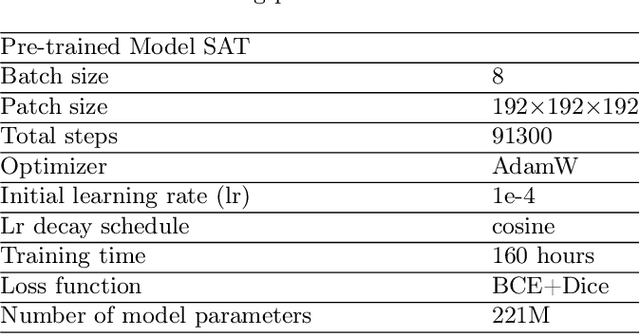
Abstract:We introduce Medal S, a medical segmentation foundation model that supports native-resolution spatial and textual prompts within an end-to-end trainable framework. Unlike text-only methods lacking spatial awareness, Medal S achieves channel-wise alignment between volumetric prompts and text embeddings, mitigating inaccuracies from resolution mismatches. By preserving full 3D context, it efficiently processes multiple native-resolution masks in parallel, enhancing multi-class segmentation performance. A lightweight 3D convolutional module enables precise voxel-space refinement guided by both prompt types, supporting up to 243 classes across CT, MRI, PET, ultrasound, and microscopy modalities in the BiomedSegFM dataset. Medal S offers two prompting modes: a text-only mode, where model predictions serve as spatial prompts for self-refinement without human input, and a hybrid mode, incorporating manual annotations for enhanced flexibility. For 24-class segmentation, parallel spatial prompting reduces inference time by more than 90% compared to sequential prompting. We propose dynamic resampling to address target-patch ratio imbalance, extending SAT and nnU-Net for data augmentation. Furthermore, we develop optimized text preprocessing, a two-stage inference strategy, and post-processing techniques to improve memory efficiency, precision, and inference speed. On the five-modality average on the validation set, Medal S outperforms SAT with a DSC of 75.44 (vs. 69.83), NSD of 77.34 (vs. 71.06), F1 of 38.24 (vs. 24.88), and DSC TP of 65.46 (vs. 46.97). Medal S achieves excellent performance by harmonizing spatial precision with semantic textual guidance, demonstrating superior efficiency and accuracy in multi-class medical segmentation tasks compared to sequential prompt-based approaches. Medal S will be publicly available at https://github.com/yinghemedical/Medal-S.
TurboReg: TurboClique for Robust and Efficient Point Cloud Registration
Jul 02, 2025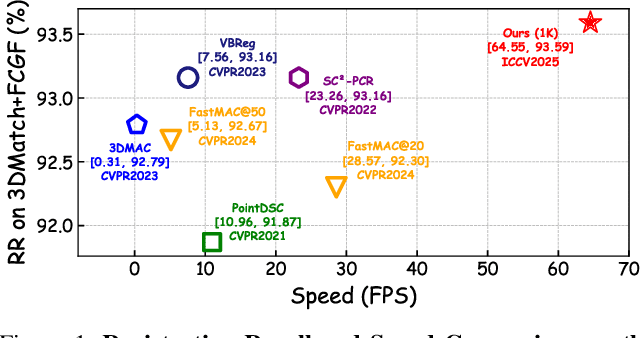
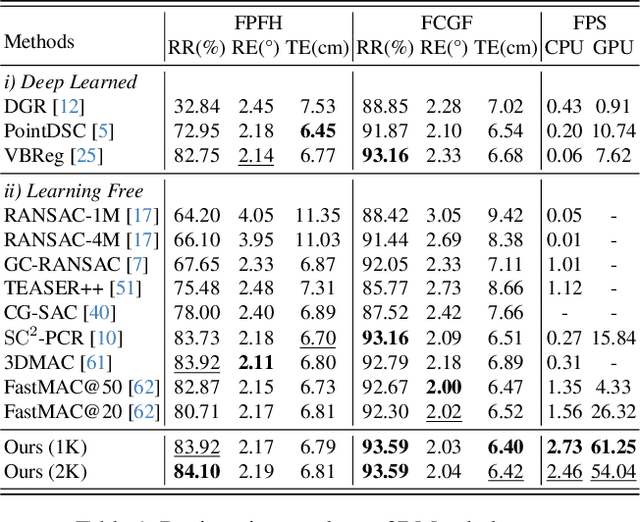


Abstract:Robust estimation is essential in correspondence-based Point Cloud Registration (PCR). Existing methods using maximal clique search in compatibility graphs achieve high recall but suffer from exponential time complexity, limiting their use in time-sensitive applications. To address this challenge, we propose a fast and robust estimator, TurboReg, built upon a novel lightweight clique, TurboClique, and a highly parallelizable Pivot-Guided Search (PGS) algorithm. First, we define the TurboClique as a 3-clique within a highly-constrained compatibility graph. The lightweight nature of the 3-clique allows for efficient parallel searching, and the highly-constrained compatibility graph ensures robust spatial consistency for stable transformation estimation. Next, PGS selects matching pairs with high SC$^2$ scores as pivots, effectively guiding the search toward TurboCliques with higher inlier ratios. Moreover, the PGS algorithm has linear time complexity and is significantly more efficient than the maximal clique search with exponential time complexity. Extensive experiments show that TurboReg achieves state-of-the-art performance across multiple real-world datasets, with substantial speed improvements. For example, on the 3DMatch+FCGF dataset, TurboReg (1K) operates $208.22\times$ faster than 3DMAC while also achieving higher recall. Our code is accessible at \href{https://github.com/Laka-3DV/TurboReg}{\texttt{TurboReg}}.
Building 3D In-Context Learning Universal Model in Neuroimaging
Mar 04, 2025Abstract:In-context learning (ICL), a type of universal model, demonstrates exceptional generalization across a wide range of tasks without retraining by leveraging task-specific guidance from context, making it particularly effective for the complex demands of neuroimaging. However, existing ICL models, which take 2D images as input, struggle to fully leverage the 3D anatomical structures in neuroimages, leading to a lack of global awareness and suboptimal performance. In this regard, we introduce Neuroverse3D, an ICL model capable of performing multiple neuroimaging tasks (e.g., segmentation, denoising, inpainting) in 3D. Neuroverse3D overcomes the large memory consumption due to 3D inputs through adaptive parallel-sequential context processing and a U-shape fusion strategy, allowing it to handle an unlimited number of context images. Additionally, we propose an optimized loss to balance multi-task training and enhance the focus on anatomical structures. Our study incorporates 43,674 3D scans from 19 neuroimaging datasets and evaluates Neuroverse3D on 14 diverse tasks using held-out test sets. The results demonstrate that Neuroverse3D significantly outperforms existing ICL models and closely matches the performance of task-specific models. The code and model weights are publicly released at: https://github.com/jiesihu/Neu3D.
Multi-Class Segmentation of Aortic Branches and Zones in Computed Tomography Angiography: The AortaSeg24 Challenge
Feb 07, 2025



Abstract:Multi-class segmentation of the aorta in computed tomography angiography (CTA) scans is essential for diagnosing and planning complex endovascular treatments for patients with aortic dissections. However, existing methods reduce aortic segmentation to a binary problem, limiting their ability to measure diameters across different branches and zones. Furthermore, no open-source dataset is currently available to support the development of multi-class aortic segmentation methods. To address this gap, we organized the AortaSeg24 MICCAI Challenge, introducing the first dataset of 100 CTA volumes annotated for 23 clinically relevant aortic branches and zones. This dataset was designed to facilitate both model development and validation. The challenge attracted 121 teams worldwide, with participants leveraging state-of-the-art frameworks such as nnU-Net and exploring novel techniques, including cascaded models, data augmentation strategies, and custom loss functions. We evaluated the submitted algorithms using the Dice Similarity Coefficient (DSC) and Normalized Surface Distance (NSD), highlighting the approaches adopted by the top five performing teams. This paper presents the challenge design, dataset details, evaluation metrics, and an in-depth analysis of the top-performing algorithms. The annotated dataset, evaluation code, and implementations of the leading methods are publicly available to support further research. All resources can be accessed at https://aortaseg24.grand-challenge.org.
Legal Evalutions and Challenges of Large Language Models
Nov 15, 2024



Abstract:In this paper, we review legal testing methods based on Large Language Models (LLMs), using the OPENAI o1 model as a case study to evaluate the performance of large models in applying legal provisions. We compare current state-of-the-art LLMs, including open-source, closed-source, and legal-specific models trained specifically for the legal domain. Systematic tests are conducted on English and Chinese legal cases, and the results are analyzed in depth. Through systematic testing of legal cases from common law systems and China, this paper explores the strengths and weaknesses of LLMs in understanding and applying legal texts, reasoning through legal issues, and predicting judgments. The experimental results highlight both the potential and limitations of LLMs in legal applications, particularly in terms of challenges related to the interpretation of legal language and the accuracy of legal reasoning. Finally, the paper provides a comprehensive analysis of the advantages and disadvantages of various types of models, offering valuable insights and references for the future application of AI in the legal field.
Touchstone Benchmark: Are We on the Right Way for Evaluating AI Algorithms for Medical Segmentation?
Nov 06, 2024



Abstract:How can we test AI performance? This question seems trivial, but it isn't. Standard benchmarks often have problems such as in-distribution and small-size test sets, oversimplified metrics, unfair comparisons, and short-term outcome pressure. As a consequence, good performance on standard benchmarks does not guarantee success in real-world scenarios. To address these problems, we present Touchstone, a large-scale collaborative segmentation benchmark of 9 types of abdominal organs. This benchmark is based on 5,195 training CT scans from 76 hospitals around the world and 5,903 testing CT scans from 11 additional hospitals. This diverse test set enhances the statistical significance of benchmark results and rigorously evaluates AI algorithms across various out-of-distribution scenarios. We invited 14 inventors of 19 AI algorithms to train their algorithms, while our team, as a third party, independently evaluated these algorithms on three test sets. In addition, we also evaluated pre-existing AI frameworks--which, differing from algorithms, are more flexible and can support different algorithms--including MONAI from NVIDIA, nnU-Net from DKFZ, and numerous other open-source frameworks. We are committed to expanding this benchmark to encourage more innovation of AI algorithms for the medical domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge