Bennett A. Landman
*: shared first/last authors
Large-Scale Deployment and Analytical Implications of Structured Quality Control in Diffusion Magnetic Resonance Imaging
May 20, 2026Abstract:Purpose: Diffusion MRI (dMRI) provides a diverse set of quantitative measures and derived datatypes to assess white matter microstructure and macrostructure. Coupled with the increasing size of imaging studies using dMRI, the number of downstream outputs requiring quality control (QC) will continue to grow. Previous work has shown that failure modes which are often not evident from aggregate metrics or summary statistics can be identified through structured visual inspection. This work aims to better understand common failure modes and the expected characteristics of valid dMRI processing outputs to ensure the validity and interpretability of quantitative findings. Approach: We deployed a structured QC framework to assess 18,328 dMRI scans across nine datasets, visually evaluating the outputs of seven processing pipelines representative of conventional dMRI analyses. Results: Downstream outputs that pass visual QC may still rely on failed upstream dependencies; such failures may only be visually detectable through systematic inspection of the full pipeline hierarchy. Additionally, appropriate QC granularity is algorithm-specific, as the spatial structure of each algorithm's outputs determines whether failures warrant selective or global exclusion. Conclusion: This work demonstrates the feasibility and analytical value of large-scale, structured QC for dMRI processing pipelines. Our results highlight the need for systematic QC spanning the full processing hierarchy to ensure the validity and interpretability of quantitative findings.
Unsupervised learning of acquisition variability in structural connectomes via hybrid latent space modeling
May 13, 2026Abstract:Acquisition differences across sites, scanners, and protocols in dMRI introduce variability that complicates structural connectome analysis. This motivates deep learning models that can represent high-dimensional connectomes in a low-dimensional space while explicitly separating acquisition-related effects from biological variation. Conventional dimensionality reduction methods model all variance as continuous, so acquisition effects often get absorbed into a continuous latent space. Recent hybrid latent-space models combine discrete and continuous components to address this, but typically require manual capacity tuning to ensure the discrete component captures the intended variability. We introduce an unsupervised framework that removes this manual tuning by architecturally annealing encoder outputs before decoding, allowing the model to adaptively balance discrete and continuous latent variables during training. To evaluate it, we curated a dataset of N=7,416 structural connectomes derived from dMRI, spanning ages 2 to 102 and 13 studies with 25 unique acquisition-parameter combinations. Of these, 5,900 are cognitively unimpaired, 877 have mild cognitive impairment (MCI), and 639 have Alzheimer's disease (AD). We compare against a standard VAE, PCA with k-means clustering, and hybrid models that anneal only through the loss function. Our architectural annealing produces stronger site learning (ARI=0.53, p<0.05) than these baselines. Results show that a hybrid continuous-discrete latent space, with architectural rather than loss-based annealing, provides a useful unsupervised mechanism for capturing acquisition variability in dMRI: by jointly modeling smooth and categorical structure, the Joint-VAE recovers clusters aligned with scanner and protocol differences.
Evaluation of neuroCombat and deep learning harmonization for multi-site magnetic resonance neuroimaging in youth with prenatal alcohol exposure
Mar 31, 2026Abstract:In cases of prevalent diseases and disorders, such as Prenatal Alcohol Exposure (PAE), multi-site data collection allows for increased study samples. However, multi-site studies introduce additional variability through heterogeneous collection materials, such as scanner and acquisition protocols, which confound with biologically relevant signals. Neuroscientists often utilize statistical methods on image-derived metrics, such as volume of regions of interest, after all image processing to minimize site-related variance. HACA3, a deep learning harmonization method, offers an opportunity to harmonize image signals prior to metric quantification; however, HACA3 has not yet been validated in a pediatric cohort. In this work, we investigate HACA3's ability to remove site-related variance and preserve biologically relevant signal compared to a statistical method, neuroCombat, and pair HACA3 processing with neuroCombat to evaluate the efficacy of multiple harmonization methods in a pediatric (age 7 to 21) population across three unique scanners with controls and cases of PAE with downstream MaCRUISE volume metrics. We find that HACA3 qualitatively improves inter-site contrast variations, but statistical methods reduce greater site-related variance within the MaCRUISE volume metrics following an ANCOVA test, and HACA3 relies on follow-up statistical methods to approach maximal biological preservation in this context.
Harmonization mitigates diffusion MRI scanner effects in infancy: insights from the HEALthy Brain and Childhood Development (HBCD) study
Mar 31, 2026Abstract:The HEALthy Brain and Childhood Development (HBCD) Study is an ongoing longitudinal initiative to understand population-level brain maturation; however, large-scale studies must overcome site-related variance and preserve biologically relevant signal. In addition to diffusion-weighted magnetic resonance imaging images, the HBCD dataset offers analysis-ready derivatives for scientists to conduct their analysis, including scalar diffusion tensor (DTI) metrics in a predetermined set of bundles. The purpose of this study is to characterize HBCD-specific site effects in diffusion MRI data, which have not been systematically reported. In this work, we investigate the sensitivity of HBCD bundle metrics to scanner model-related variance and address these variations with ComBat-GAM harmonization within the current HBCD data release 1.1 across six scanner models. Following ComBat-GAM, we observe zero statistically significant differences between the distributions from any scanner model following FDR correction and reduce Cohen's f effect sizes across all metrics. Our work underscores the importance of rigorous harmonization efforts in large-scale studies, and we encourage future investigations of HBCD data to control for these effects.
Insertion Network for Image Sequence Correspondence
Feb 13, 2026Abstract:We propose a novel method for establishing correspondence between two sequences of 2D images. One particular application of this technique is slice-level content navigation, where the goal is to localize specific 2D slices within a 3D volume or determine the anatomical coverage of a 3D scan based on its 2D slices. This serves as an important preprocessing step for various diagnostic tasks, as well as for automatic registration and segmentation pipelines. Our approach builds sequence correspondence by training a network to learn how to insert a slice from one sequence into the appropriate position in another. This is achieved by encoding contextual representations of each slice and modeling the insertion process using a slice-to-slice attention mechanism. We apply this method to localize manually labeled key slices in body CT scans and compare its performance to the current state-of-the-art alternative known as body part regression, which predicts anatomical position scores for individual slices. Unlike body part regression, which treats each slice independently, our method leverages contextual information from the entire sequence. Experimental results show that the insertion network reduces slice localization errors in supervised settings from 8.4 mm to 5.4 mm, demonstrating a substantial improvement in accuracy.
Personalized White Matter Bundle Segmentation for Early Childhood
Feb 04, 2026Abstract:White matter segmentation methods from diffusion magnetic resonance imaging range from streamline clustering-based approaches to bundle mask delineation, but none have proposed a pediatric-specific approach. We hypothesize that a deep learning model with a similar approach to TractSeg will improve similarity between an algorithm-generated mask and an expert-labeled ground truth. Given a cohort of 56 manually labelled white matter bundles, we take inspiration from TractSeg's 2D UNet architecture, and we modify inputs to match bundle definitions as determined by pediatric experts, evaluation to use k fold cross validation, the loss function to masked Dice loss. We evaluate Dice score, volume overlap, and volume overreach of 16 major regions of interest compared to the expert labeled dataset. To test whether our approach offers statistically significant improvements over TractSeg, we compare Dice voxels, volume overlap, and adjacency voxels with a Wilcoxon signed rank test followed by false discovery rate correction. We find statistical significance across all bundles for all metrics with one exception in volume overlap. After we run TractSeg and our model, we combine their output masks into a 60 label atlas to evaluate if TractSeg and our model combined can generate a robust, individualized atlas, and observe smoothed, continuous masks in cases that TractSeg did not produce an anatomically plausible output. With the improvement of white matter pathway segmentation masks, we can further understand neurodevelopment on a population level scale, and we can produce reliable estimates of individualized anatomy in pediatric white matter diseases and disorders.
AdaFuse: Adaptive Multimodal Fusion for Lung Cancer Risk Prediction via Reinforcement Learning
Jan 30, 2026Abstract:Multimodal fusion has emerged as a promising paradigm for disease diagnosis and prognosis, integrating complementary information from heterogeneous data sources such as medical images, clinical records, and radiology reports. However, existing fusion methods process all available modalities through the network, either treating them equally or learning to assign different contribution weights, leaving a fundamental question unaddressed: for a given patient, should certain modalities be used at all? We present AdaFuse, an adaptive multimodal fusion framework that leverages reinforcement learning (RL) to learn patient-specific modality selection and fusion strategies for lung cancer risk prediction. AdaFuse formulates multimodal fusion as a sequential decision process, where the policy network iteratively decides whether to incorporate an additional modality or proceed to prediction based on the information already acquired. This sequential formulation enables the model to condition each selection on previously observed modalities and terminate early when sufficient information is available, rather than committing to a fixed subset upfront. We evaluate AdaFuse on the National Lung Screening Trial (NLST) dataset. Experimental results demonstrate that AdaFuse achieves the highest AUC (0.762) compared to the best single-modality baseline (0.732), the best fixed fusion strategy (0.759), and adaptive baselines including DynMM (0.754) and MoE (0.742), while using fewer FLOPs than all triple-modality methods. Our work demonstrates the potential of reinforcement learning for personalized multimodal fusion in medical imaging, representing a shift from uniform fusion strategies toward adaptive diagnostic pipelines that learn when to consult additional modalities and when existing information suffices for accurate prediction.
Medical Imaging AI Competitions Lack Fairness
Dec 19, 2025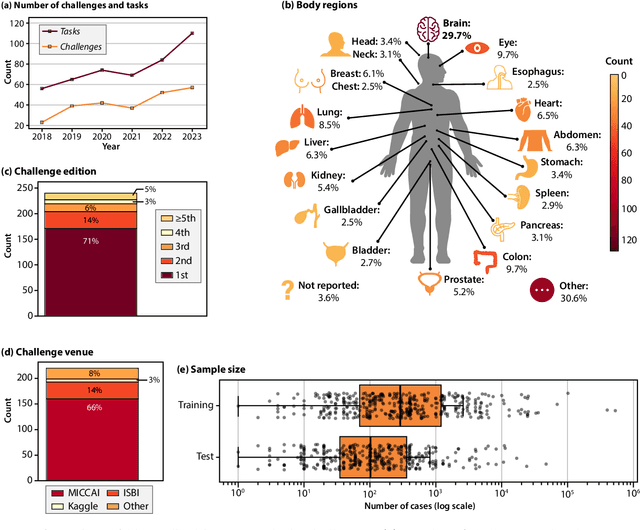
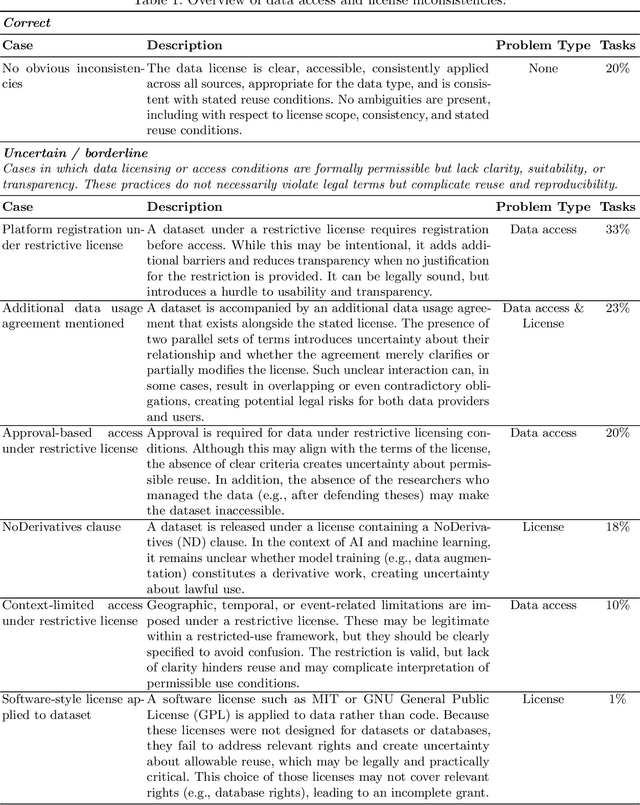
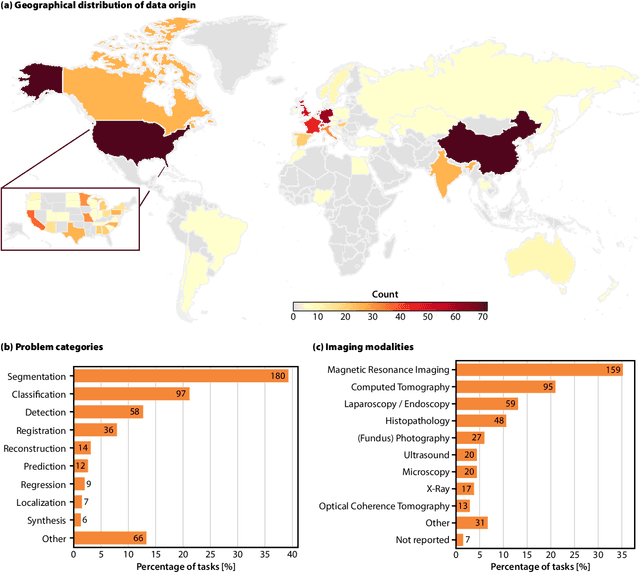
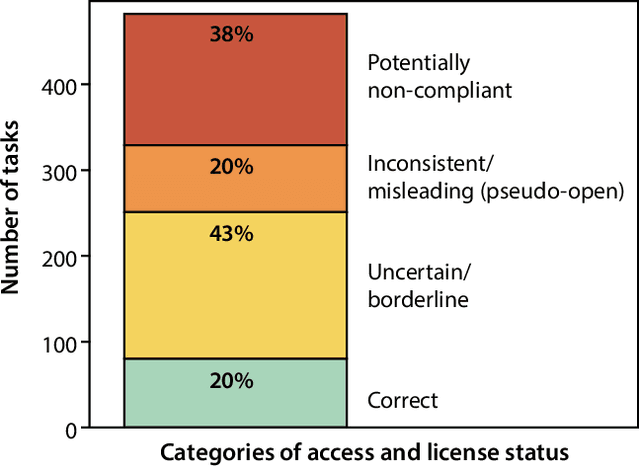
Abstract:Benchmarking competitions are central to the development of artificial intelligence (AI) in medical imaging, defining performance standards and shaping methodological progress. However, it remains unclear whether these benchmarks provide data that are sufficiently representative, accessible, and reusable to support clinically meaningful AI. In this work, we assess fairness along two complementary dimensions: (1) whether challenge datasets are representative of real-world clinical diversity, and (2) whether they are accessible and legally reusable in line with the FAIR principles. To address this question, we conducted a large-scale systematic study of 241 biomedical image analysis challenges comprising 458 tasks across 19 imaging modalities. Our findings show substantial biases in dataset composition, including geographic location, modality-, and problem type-related biases, indicating that current benchmarks do not adequately reflect real-world clinical diversity. Despite their widespread influence, challenge datasets were frequently constrained by restrictive or ambiguous access conditions, inconsistent or non-compliant licensing practices, and incomplete documentation, limiting reproducibility and long-term reuse. Together, these shortcomings expose foundational fairness limitations in our benchmarking ecosystem and highlight a disconnect between leaderboard success and clinical relevance.
MetaVoxel: Joint Diffusion Modeling of Imaging and Clinical Metadata
Dec 12, 2025Abstract:Modern deep learning methods have achieved impressive results across tasks from disease classification, estimating continuous biomarkers, to generating realistic medical images. Most of these approaches are trained to model conditional distributions defined by a specific predictive direction with a specific set of input variables. We introduce MetaVoxel, a generative joint diffusion modeling framework that models the joint distribution over imaging data and clinical metadata by learning a single diffusion process spanning all variables. By capturing the joint distribution, MetaVoxel unifies tasks that traditionally require separate conditional models and supports flexible zero-shot inference using arbitrary subsets of inputs without task-specific retraining. Using more than 10,000 T1-weighted MRI scans paired with clinical metadata from nine datasets, we show that a single MetaVoxel model can perform image generation, age estimation, and sex prediction, achieving performance comparable to established task-specific baselines. Additional experiments highlight its capabilities for flexible inference. Together, these findings demonstrate that joint multimodal diffusion offers a promising direction for unifying medical AI models and enabling broader clinical applicability.
Fully Differentiable dMRI Streamline Propagation in PyTorch
Nov 17, 2025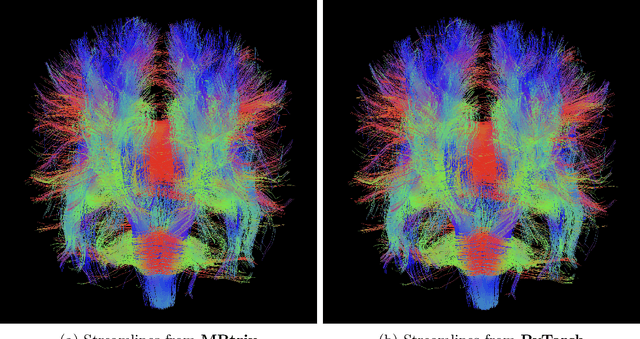
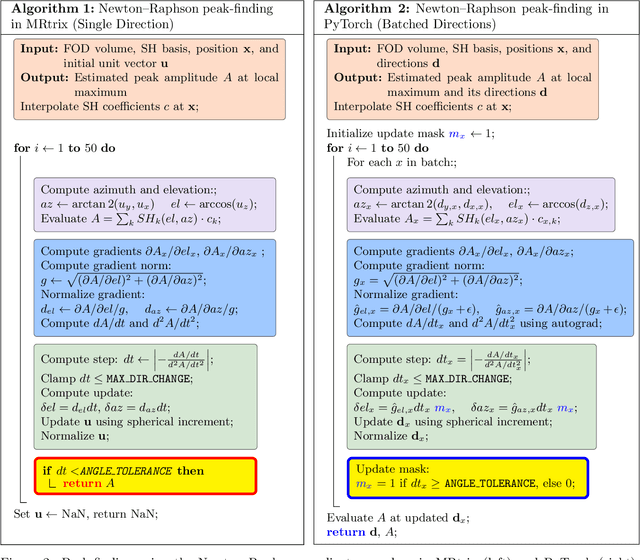
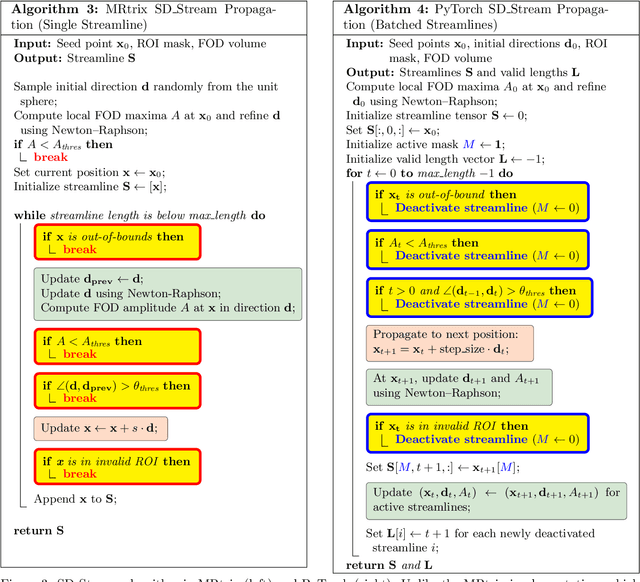
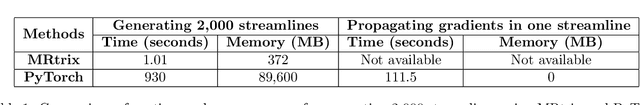


Abstract:Diffusion MRI (dMRI) provides a distinctive means to probe the microstructural architecture of living tissue, facilitating applications such as brain connectivity analysis, modeling across multiple conditions, and the estimation of macrostructural features. Tractography, which emerged in the final years of the 20th century and accelerated in the early 21st century, is a technique for visualizing white matter pathways in the brain using dMRI. Most diffusion tractography methods rely on procedural streamline propagators or global energy minimization methods. Although recent advancements in deep learning have enabled tasks that were previously challenging, existing tractography approaches are often non-differentiable, limiting their integration in end-to-end learning frameworks. While progress has been made in representing streamlines in differentiable frameworks, no existing method offers fully differentiable propagation. In this work, we propose a fully differentiable solution that retains numerical fidelity with a leading streamline algorithm. The key is that our PyTorch-engineered streamline propagator has no components that block gradient flow, making it fully differentiable. We show that our method matches standard propagators while remaining differentiable. By translating streamline propagation into a differentiable PyTorch framework, we enable deeper integration of tractography into deep learning workflows, laying the foundation for a new category of macrostructural reasoning that is not only computationally robust but also scientifically rigorous.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge