Ann Zenobia Moore
Pitfalls of defacing whole-head MRI: re-identification risk with diffusion models and compromised research potential
Jan 31, 2025Abstract:Defacing is often applied to head magnetic resonance image (MRI) datasets prior to public release to address privacy concerns. The alteration of facial and nearby voxels has provoked discussions about the true capability of these techniques to ensure privacy as well as their impact on downstream tasks. With advancements in deep generative models, the extent to which defacing can protect privacy is uncertain. Additionally, while the altered voxels are known to contain valuable anatomical information, their potential to support research beyond the anatomical regions directly affected by defacing remains uncertain. To evaluate these considerations, we develop a refacing pipeline that recovers faces in defaced head MRIs using cascaded diffusion probabilistic models (DPMs). The DPMs are trained on images from 180 subjects and tested on images from 484 unseen subjects, 469 of whom are from a different dataset. To assess whether the altered voxels in defacing contain universally useful information, we also predict computed tomography (CT)-derived skeletal muscle radiodensity from facial voxels in both defaced and original MRIs. The results show that DPMs can generate high-fidelity faces that resemble the original faces from defaced images, with surface distances to the original faces significantly smaller than those of a population average face (p < 0.05). This performance also generalizes well to previously unseen datasets. For skeletal muscle radiodensity predictions, using defaced images results in significantly weaker Spearman's rank correlation coefficients compared to using original images (p < 10-4). For shin muscle, the correlation is statistically significant (p < 0.05) when using original images but not statistically significant (p > 0.05) when any defacing method is applied, suggesting that defacing might not only fail to protect privacy but also eliminate valuable information.
Deep conditional generative models for longitudinal single-slice abdominal computed tomography harmonization
Sep 17, 2023Abstract:Two-dimensional single-slice abdominal computed tomography (CT) provides a detailed tissue map with high resolution allowing quantitative characterization of relationships between health conditions and aging. However, longitudinal analysis of body composition changes using these scans is difficult due to positional variation between slices acquired in different years, which leading to different organs/tissues captured. To address this issue, we propose C-SliceGen, which takes an arbitrary axial slice in the abdominal region as a condition and generates a pre-defined vertebral level slice by estimating structural changes in the latent space. Our experiments on 2608 volumetric CT data from two in-house datasets and 50 subjects from the 2015 Multi-Atlas Abdomen Labeling Challenge dataset (BTCV) Challenge demonstrate that our model can generate high-quality images that are realistic and similar. We further evaluate our method's capability to harmonize longitudinal positional variation on 1033 subjects from the Baltimore Longitudinal Study of Aging (BLSA) dataset, which contains longitudinal single abdominal slices, and confirmed that our method can harmonize the slice positional variance in terms of visceral fat area. This approach provides a promising direction for mapping slices from different vertebral levels to a target slice and reducing positional variance for single-slice longitudinal analysis. The source code is available at: https://github.com/MASILab/C-SliceGen.
Single Slice Thigh CT Muscle Group Segmentation with Domain Adaptation and Self-Training
Nov 30, 2022Abstract:Objective: Thigh muscle group segmentation is important for assessment of muscle anatomy, metabolic disease and aging. Many efforts have been put into quantifying muscle tissues with magnetic resonance (MR) imaging including manual annotation of individual muscles. However, leveraging publicly available annotations in MR images to achieve muscle group segmentation on single slice computed tomography (CT) thigh images is challenging. Method: We propose an unsupervised domain adaptation pipeline with self-training to transfer labels from 3D MR to single CT slice. First, we transform the image appearance from MR to CT with CycleGAN and feed the synthesized CT images to a segmenter simultaneously. Single CT slices are divided into hard and easy cohorts based on the entropy of pseudo labels inferenced by the segmenter. After refining easy cohort pseudo labels based on anatomical assumption, self-training with easy and hard splits is applied to fine tune the segmenter. Results: On 152 withheld single CT thigh images, the proposed pipeline achieved a mean Dice of 0.888(0.041) across all muscle groups including sartorius, hamstrings, quadriceps femoris and gracilis. muscles Conclusion: To our best knowledge, this is the first pipeline to achieve thigh imaging domain adaptation from MR to CT. The proposed pipeline is effective and robust in extracting muscle groups on 2D single slice CT thigh images.The container is available for public use at https://github.com/MASILab/DA_CT_muscle_seg
Reducing Positional Variance in Cross-sectional Abdominal CT Slices with Deep Conditional Generative Models
Sep 28, 2022Abstract:2D low-dose single-slice abdominal computed tomography (CT) slice enables direct measurements of body composition, which are critical to quantitatively characterizing health relationships on aging. However, longitudinal analysis of body composition changes using 2D abdominal slices is challenging due to positional variance between longitudinal slices acquired in different years. To reduce the positional variance, we extend the conditional generative models to our C-SliceGen that takes an arbitrary axial slice in the abdominal region as the condition and generates a defined vertebral level slice by estimating the structural changes in the latent space. Experiments on 1170 subjects from an in-house dataset and 50 subjects from BTCV MICCAI Challenge 2015 show that our model can generate high quality images in terms of realism and similarity. External experiments on 20 subjects from the Baltimore Longitudinal Study of Aging (BLSA) dataset that contains longitudinal single abdominal slices validate that our method can harmonize the slice positional variance in terms of muscle and visceral fat area. Our approach provides a promising direction of mapping slices from different vertebral levels to a target slice to reduce positional variance for single slice longitudinal analysis. The source code is available at: https://github.com/MASILab/C-SliceGen.
* 11 pages, 4 figures
Longitudinal Variability Analysis on Low-dose Abdominal CT with Deep Learning-based Segmentation
Sep 28, 2022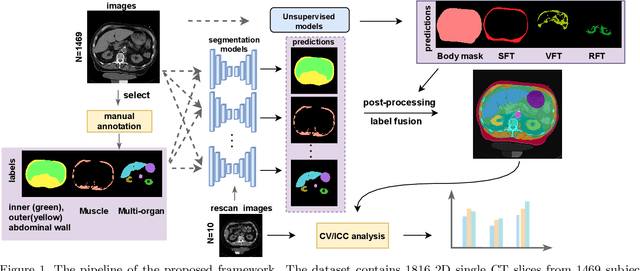
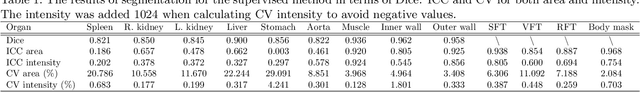
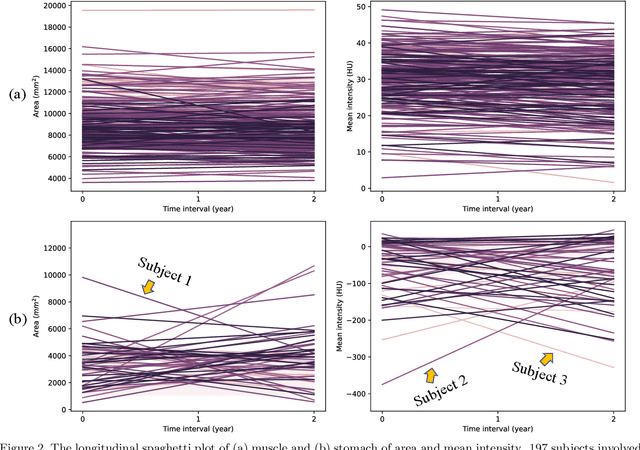
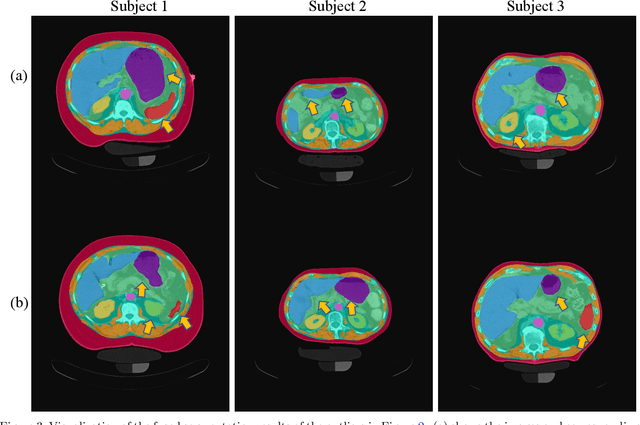
Abstract:Metabolic health is increasingly implicated as a risk factor across conditions from cardiology to neurology, and efficiency assessment of body composition is critical to quantitatively characterizing these relationships. 2D low dose single slice computed tomography (CT) provides a high resolution, quantitative tissue map, albeit with a limited field of view. Although numerous potential analyses have been proposed in quantifying image context, there has been no comprehensive study for low-dose single slice CT longitudinal variability with automated segmentation. We studied a total of 1816 slices from 1469 subjects of Baltimore Longitudinal Study on Aging (BLSA) abdominal dataset using supervised deep learning-based segmentation and unsupervised clustering method. 300 out of 1469 subjects that have two year gap in their first two scans were pick out to evaluate longitudinal variability with measurements including intraclass correlation coefficient (ICC) and coefficient of variation (CV) in terms of tissues/organs size and mean intensity. We showed that our segmentation methods are stable in longitudinal settings with Dice ranged from 0.821 to 0.962 for thirteen target abdominal tissues structures. We observed high variability in most organ with ICC<0.5, low variability in the area of muscle, abdominal wall, fat and body mask with average ICC>0.8. We found that the variability in organ is highly related to the cross-sectional position of the 2D slice. Our efforts pave quantitative exploration and quality control to reduce uncertainties in longitudinal analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge