Shunxing Bao
Lifespan Pancreas Morphology for Control vs Type 2 Diabetes using AI on Largescale Clinical Imaging
Aug 20, 2025Abstract:Purpose: Understanding how the pancreas changes is critical for detecting deviations in type 2 diabetes and other pancreatic disease. We measure pancreas size and shape using morphological measurements from ages 0 to 90. Our goals are to 1) identify reliable clinical imaging modalities for AI-based pancreas measurement, 2) establish normative morphological aging trends, and 3) detect potential deviations in type 2 diabetes. Approach: We analyzed a clinically acquired dataset of 2533 patients imaged with abdominal CT or MRI. We resampled the scans to 3mm isotropic resolution, segmented the pancreas using automated methods, and extracted 13 morphological pancreas features across the lifespan. First, we assessed CT and MRI measurements to determine which modalities provide consistent lifespan trends. Second, we characterized distributions of normative morphological patterns stratified by age group and sex. Third, we used GAMLSS regression to model pancreas morphology trends in 1350 patients matched for age, sex, and type 2 diabetes status to identify any deviations from normative aging associated with type 2 diabetes. Results: When adjusting for confounders, the aging trends for 10 of 13 morphological features were significantly different between patients with type 2 diabetes and non-diabetic controls (p < 0.05 after multiple comparisons corrections). Additionally, MRI appeared to yield different pancreas measurements than CT using our AI-based method. Conclusions: We provide lifespan trends demonstrating that the size and shape of the pancreas is altered in type 2 diabetes using 675 control patients and 675 diabetes patients. Moreover, our findings reinforce that the pancreas is smaller in type 2 diabetes. Additionally, we contribute a reference of lifespan pancreas morphology from a large cohort of non-diabetic control patients in a clinical setting.
Multipath cycleGAN for harmonization of paired and unpaired low-dose lung computed tomography reconstruction kernels
May 28, 2025Abstract:Reconstruction kernels in computed tomography (CT) affect spatial resolution and noise characteristics, introducing systematic variability in quantitative imaging measurements such as emphysema quantification. Choosing an appropriate kernel is therefore essential for consistent quantitative analysis. We propose a multipath cycleGAN model for CT kernel harmonization, trained on a mixture of paired and unpaired data from a low-dose lung cancer screening cohort. The model features domain-specific encoders and decoders with a shared latent space and uses discriminators tailored for each domain.We train the model on 42 kernel combinations using 100 scans each from seven representative kernels in the National Lung Screening Trial (NLST) dataset. To evaluate performance, 240 scans from each kernel are harmonized to a reference soft kernel, and emphysema is quantified before and after harmonization. A general linear model assesses the impact of age, sex, smoking status, and kernel on emphysema. We also evaluate harmonization from soft kernels to a reference hard kernel. To assess anatomical consistency, we compare segmentations of lung vessels, muscle, and subcutaneous adipose tissue generated by TotalSegmentator between harmonized and original images. Our model is benchmarked against traditional and switchable cycleGANs. For paired kernels, our approach reduces bias in emphysema scores, as seen in Bland-Altman plots (p<0.05). For unpaired kernels, harmonization eliminates confounding differences in emphysema (p>0.05). High Dice scores confirm preservation of muscle and fat anatomy, while lung vessel overlap remains reasonable. Overall, our shared latent space multipath cycleGAN enables robust harmonization across paired and unpaired CT kernels, improving emphysema quantification and preserving anatomical fidelity.
Rep3D: Re-parameterize Large 3D Kernels with Low-Rank Receptive Modeling for Medical Imaging
May 26, 2025Abstract:In contrast to vision transformers, which model long-range dependencies through global self-attention, large kernel convolutions provide a more efficient and scalable alternative, particularly in high-resolution 3D volumetric settings. However, naively increasing kernel size often leads to optimization instability and degradation in performance. Motivated by the spatial bias observed in effective receptive fields (ERFs), we hypothesize that different kernel elements converge at variable rates during training. To support this, we derive a theoretical connection between element-wise gradients and first-order optimization, showing that structurally re-parameterized convolution blocks inherently induce spatially varying learning rates. Building on this insight, we introduce Rep3D, a 3D convolutional framework that incorporates a learnable spatial prior into large kernel training. A lightweight two-stage modulation network generates a receptive-biased scaling mask, adaptively re-weighting kernel updates and enabling local-to-global convergence behavior. Rep3D adopts a plain encoder design with large depthwise convolutions, avoiding the architectural complexity of multi-branch compositions. We evaluate Rep3D on five challenging 3D segmentation benchmarks and demonstrate consistent improvements over state-of-the-art baselines, including transformer-based and fixed-prior re-parameterization methods. By unifying spatial inductive bias with optimization-aware learning, Rep3D offers an interpretable, and scalable solution for 3D medical image analysis. The source code is publicly available at https://github.com/leeh43/Rep3D.
Scale-up Unlearnable Examples Learning with High-Performance Computing
Jan 10, 2025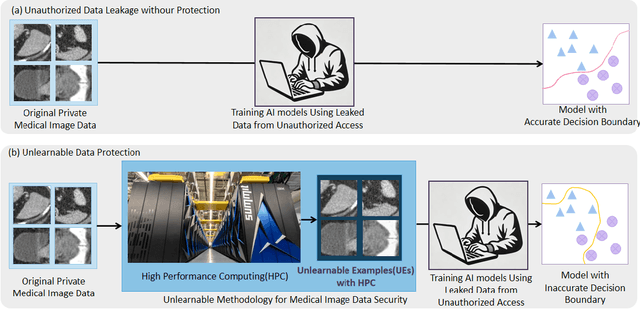
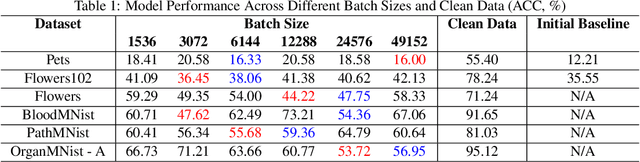
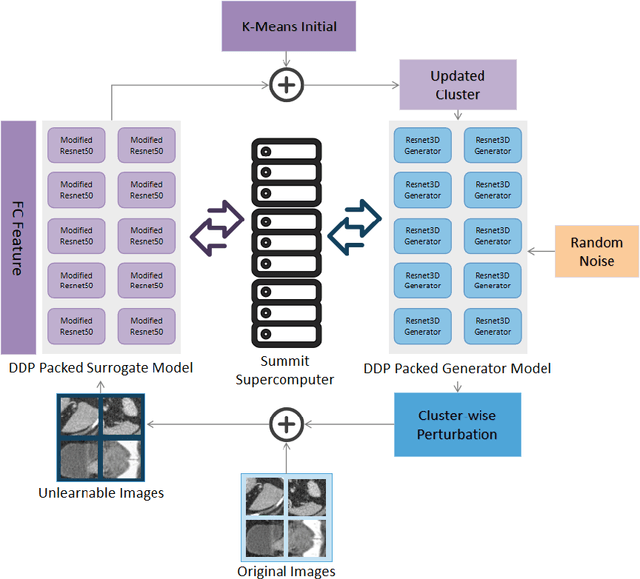
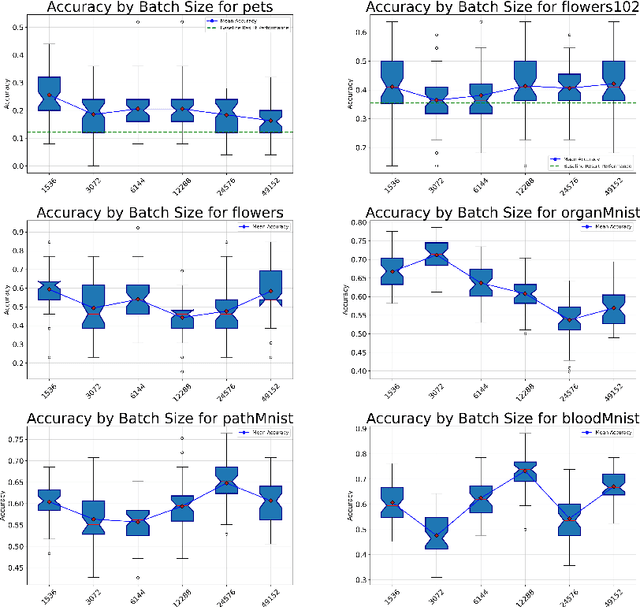
Abstract:Recent advancements in AI models are structured to retain user interactions, which could inadvertently include sensitive healthcare data. In the healthcare field, particularly when radiologists use AI-driven diagnostic tools hosted on online platforms, there is a risk that medical imaging data may be repurposed for future AI training without explicit consent, spotlighting critical privacy and intellectual property concerns around healthcare data usage. Addressing these privacy challenges, a novel approach known as Unlearnable Examples (UEs) has been introduced, aiming to make data unlearnable to deep learning models. A prominent method within this area, called Unlearnable Clustering (UC), has shown improved UE performance with larger batch sizes but was previously limited by computational resources. To push the boundaries of UE performance with theoretically unlimited resources, we scaled up UC learning across various datasets using Distributed Data Parallel (DDP) training on the Summit supercomputer. Our goal was to examine UE efficacy at high-performance computing (HPC) levels to prevent unauthorized learning and enhance data security, particularly exploring the impact of batch size on UE's unlearnability. Utilizing the robust computational capabilities of the Summit, extensive experiments were conducted on diverse datasets such as Pets, MedMNist, Flowers, and Flowers102. Our findings reveal that both overly large and overly small batch sizes can lead to performance instability and affect accuracy. However, the relationship between batch size and unlearnability varied across datasets, highlighting the necessity for tailored batch size strategies to achieve optimal data protection. Our results underscore the critical role of selecting appropriate batch sizes based on the specific characteristics of each dataset to prevent learning and ensure data security in deep learning applications.
Brain age identification from diffusion MRI synergistically predicts neurodegenerative disease
Oct 29, 2024



Abstract:Estimated brain age from magnetic resonance image (MRI) and its deviation from chronological age can provide early insights into potential neurodegenerative diseases, supporting early detection and implementation of prevention strategies. Diffusion MRI (dMRI), a widely used modality for brain age estimation, presents an opportunity to build an earlier biomarker for neurodegenerative disease prediction because it captures subtle microstructural changes that precede more perceptible macrostructural changes. However, the coexistence of macro- and micro-structural information in dMRI raises the question of whether current dMRI-based brain age estimation models are leveraging the intended microstructural information or if they inadvertently rely on the macrostructural information. To develop a microstructure-specific brain age, we propose a method for brain age identification from dMRI that minimizes the model's use of macrostructural information by non-rigidly registering all images to a standard template. Imaging data from 13,398 participants across 12 datasets were used for the training and evaluation. We compare our brain age models, trained with and without macrostructural information minimized, with an architecturally similar T1-weighted (T1w) MRI-based brain age model and two state-of-the-art T1w MRI-based brain age models that primarily use macrostructural information. We observe difference between our dMRI-based brain age and T1w MRI-based brain age across stages of neurodegeneration, with dMRI-based brain age being older than T1w MRI-based brain age in participants transitioning from cognitively normal (CN) to mild cognitive impairment (MCI), but younger in participants already diagnosed with Alzheimer's disease (AD). Approximately 4 years before MCI diagnosis, dMRI-based brain age yields better performance than T1w MRI-based brain ages in predicting transition from CN to MCI.
Multi-Modality Conditioned Variational U-Net for Field-of-View Extension in Brain Diffusion MRI
Sep 20, 2024
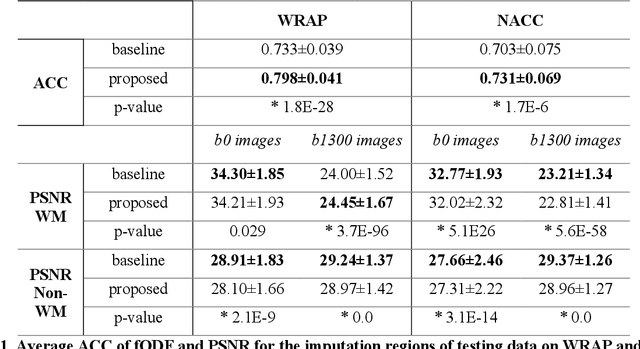

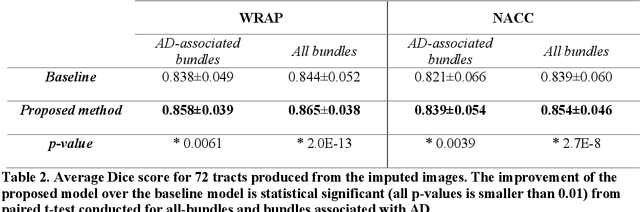
Abstract:An incomplete field-of-view (FOV) in diffusion magnetic resonance imaging (dMRI) can severely hinder the volumetric and bundle analyses of whole-brain white matter connectivity. Although existing works have investigated imputing the missing regions using deep generative models, it remains unclear how to specifically utilize additional information from paired multi-modality data and whether this can enhance the imputation quality and be useful for downstream tractography. To fill this gap, we propose a novel framework for imputing dMRI scans in the incomplete part of the FOV by integrating the learned diffusion features in the acquired part of the FOV to the complete brain anatomical structure. We hypothesize that by this design the proposed framework can enhance the imputation performance of the dMRI scans and therefore be useful for repairing whole-brain tractography in corrupted dMRI scans with incomplete FOV. We tested our framework on two cohorts from different sites with a total of 96 subjects and compared it with a baseline imputation method that treats the information from T1w and dMRI scans equally. The proposed framework achieved significant improvements in imputation performance, as demonstrated by angular correlation coefficient (p < 1E-5), and in downstream tractography accuracy, as demonstrated by Dice score (p < 0.01). Results suggest that the proposed framework improved imputation performance in dMRI scans by specifically utilizing additional information from paired multi-modality data, compared with the baseline method. The imputation achieved by the proposed framework enhances whole brain tractography, and therefore reduces the uncertainty when analyzing bundles associated with neurodegenerative.
Influence of Early through Late Fusion on Pancreas Segmentation from Imperfectly Registered Multimodal MRI
Sep 06, 2024Abstract:Multimodal fusion promises better pancreas segmentation. However, where to perform fusion in models is still an open question. It is unclear if there is a best location to fuse information when analyzing pairs of imperfectly aligned images. Two main alignment challenges in this pancreas segmentation study are 1) the pancreas is deformable and 2) breathing deforms the abdomen. Even after image registration, relevant deformations are often not corrected. We examine how early through late fusion impacts pancreas segmentation. We used 353 pairs of T2-weighted (T2w) and T1-weighted (T1w) abdominal MR images from 163 subjects with accompanying pancreas labels. We used image registration (deeds) to align the image pairs. We trained a collection of basic UNets with different fusion points, spanning from early to late, to assess how early through late fusion influenced segmentation performance on imperfectly aligned images. We assessed generalization of fusion points on nnUNet. The single-modality T2w baseline using a basic UNet model had a Dice score of 0.73, while the same baseline on the nnUNet model achieved 0.80. For the basic UNet, the best fusion approach occurred in the middle of the encoder (early/mid fusion), which led to a statistically significant improvement of 0.0125 on Dice score compared to the baseline. For the nnUNet, the best fusion approach was na\"ive image concatenation before the model (early fusion), which resulted in a statistically significant Dice score increase of 0.0021 compared to baseline. Fusion in specific blocks can improve performance, but the best blocks for fusion are model specific, and the gains are small. In imperfectly registered datasets, fusion is a nuanced problem, with the art of design remaining vital for uncovering potential insights. Future innovation is needed to better address fusion in cases of imperfect alignment of abdominal image pairs.
Persistence Image from 3D Medical Image: Superpixel and Optimized Gaussian Coefficient
Aug 15, 2024Abstract:Topological data analysis (TDA) uncovers crucial properties of objects in medical imaging. Methods based on persistent homology have demonstrated their advantages in capturing topological features that traditional deep learning methods cannot detect in both radiology and pathology. However, previous research primarily focused on 2D image analysis, neglecting the comprehensive 3D context. In this paper, we propose an innovative 3D TDA approach that incorporates the concept of superpixels to transform 3D medical image features into point cloud data. By Utilizing Optimized Gaussian Coefficient, the proposed 3D TDA method, for the first time, efficiently generate holistic Persistence Images for 3D volumetric data. Our 3D TDA method exhibits superior performance on the MedMNist3D dataset when compared to other traditional methods, showcasing its potential effectiveness in modeling 3D persistent homology-based topological analysis when it comes to classification tasks. The source code is publicly available at https://github.com/hrlblab/TopologicalDataAnalysis3D.
HATs: Hierarchical Adaptive Taxonomy Segmentation for Panoramic Pathology Image Analysis
Jun 30, 2024



Abstract:Panoramic image segmentation in computational pathology presents a remarkable challenge due to the morphologically complex and variably scaled anatomy. For instance, the intricate organization in kidney pathology spans multiple layers, from regions like the cortex and medulla to functional units such as glomeruli, tubules, and vessels, down to various cell types. In this paper, we propose a novel Hierarchical Adaptive Taxonomy Segmentation (HATs) method, which is designed to thoroughly segment panoramic views of kidney structures by leveraging detailed anatomical insights. Our approach entails (1) the innovative HATs technique which translates spatial relationships among 15 distinct object classes into a versatile "plug-and-play" loss function that spans across regions, functional units, and cells, (2) the incorporation of anatomical hierarchies and scale considerations into a unified simple matrix representation for all panoramic entities, (3) the adoption of the latest AI foundation model (EfficientSAM) as a feature extraction tool to boost the model's adaptability, yet eliminating the need for manual prompt generation in conventional segment anything model (SAM). Experimental findings demonstrate that the HATs method offers an efficient and effective strategy for integrating clinical insights and imaging precedents into a unified segmentation model across more than 15 categories. The official implementation is publicly available at https://github.com/hrlblab/HATs.
Enhancing Single-Slice Segmentation with 3D-to-2D Unpaired Scan Distillation
Jun 18, 2024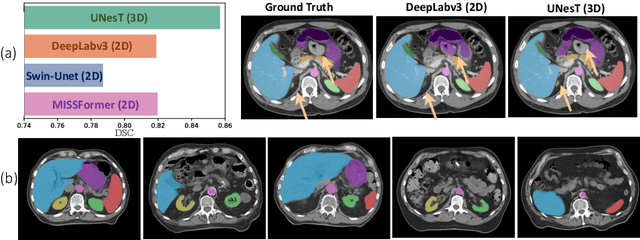
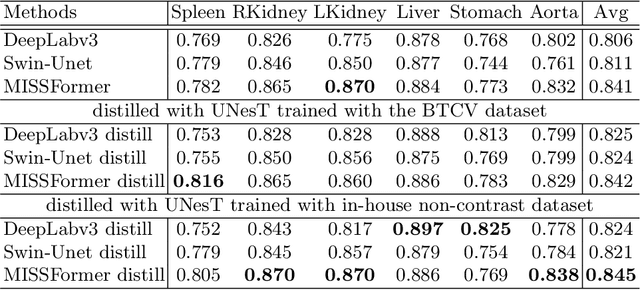
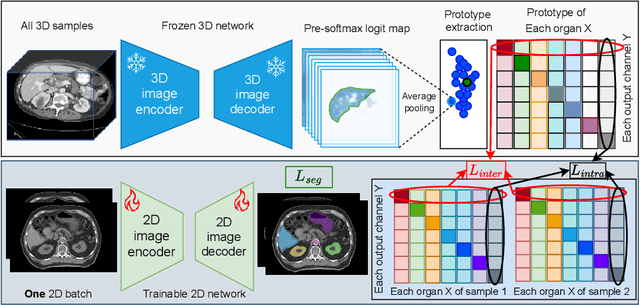
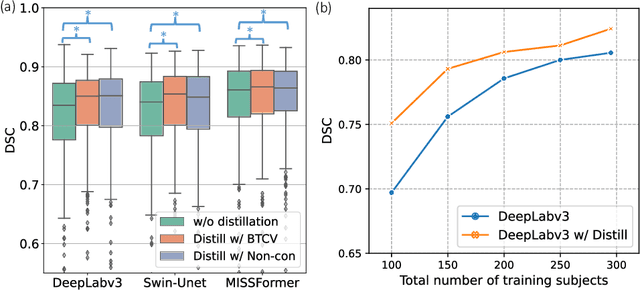
Abstract:2D single-slice abdominal computed tomography (CT) enables the assessment of body habitus and organ health with low radiation exposure. However, single-slice data necessitates the use of 2D networks for segmentation, but these networks often struggle to capture contextual information effectively. Consequently, even when trained on identical datasets, 3D networks typically achieve superior segmentation results. In this work, we propose a novel 3D-to-2D distillation framework, leveraging pre-trained 3D models to enhance 2D single-slice segmentation. Specifically, we extract the prediction distribution centroid from the 3D representations, to guide the 2D student by learning intra- and inter-class correlation. Unlike traditional knowledge distillation methods that require the same data input, our approach employs unpaired 3D CT scans with any contrast to guide the 2D student model. Experiments conducted on 707 subjects from the single-slice Baltimore Longitudinal Study of Aging (BLSA) dataset demonstrate that state-of-the-art 2D multi-organ segmentation methods can benefit from the 3D teacher model, achieving enhanced performance in single-slice multi-organ segmentation. Notably, our approach demonstrates considerable efficacy in low-data regimes, outperforming the model trained with all available training subjects even when utilizing only 200 training subjects. Thus, this work underscores the potential to alleviate manual annotation burdens.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge