Aravind R. Krishnan
Personalized White Matter Bundle Segmentation for Early Childhood
Feb 04, 2026Abstract:White matter segmentation methods from diffusion magnetic resonance imaging range from streamline clustering-based approaches to bundle mask delineation, but none have proposed a pediatric-specific approach. We hypothesize that a deep learning model with a similar approach to TractSeg will improve similarity between an algorithm-generated mask and an expert-labeled ground truth. Given a cohort of 56 manually labelled white matter bundles, we take inspiration from TractSeg's 2D UNet architecture, and we modify inputs to match bundle definitions as determined by pediatric experts, evaluation to use k fold cross validation, the loss function to masked Dice loss. We evaluate Dice score, volume overlap, and volume overreach of 16 major regions of interest compared to the expert labeled dataset. To test whether our approach offers statistically significant improvements over TractSeg, we compare Dice voxels, volume overlap, and adjacency voxels with a Wilcoxon signed rank test followed by false discovery rate correction. We find statistical significance across all bundles for all metrics with one exception in volume overlap. After we run TractSeg and our model, we combine their output masks into a 60 label atlas to evaluate if TractSeg and our model combined can generate a robust, individualized atlas, and observe smoothed, continuous masks in cases that TractSeg did not produce an anatomically plausible output. With the improvement of white matter pathway segmentation masks, we can further understand neurodevelopment on a population level scale, and we can produce reliable estimates of individualized anatomy in pediatric white matter diseases and disorders.
Self-supervised learning of imaging and clinical signatures using a multimodal joint-embedding predictive architecture
Sep 18, 2025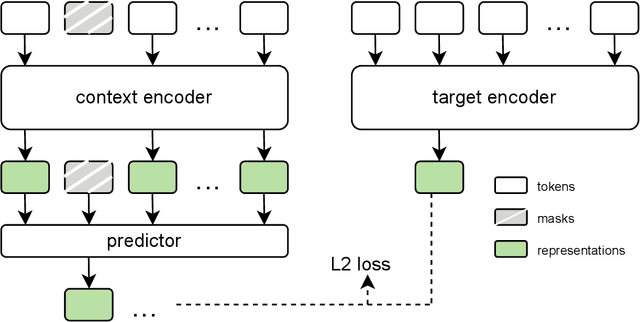
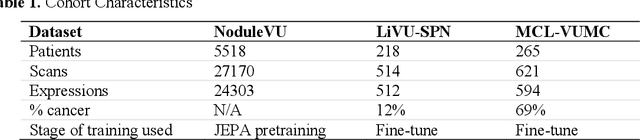

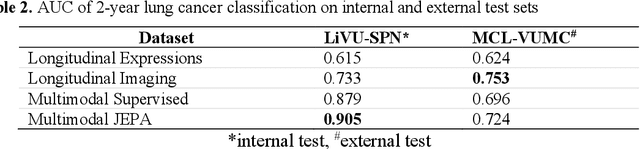
Abstract:The development of multimodal models for pulmonary nodule diagnosis is limited by the scarcity of labeled data and the tendency for these models to overfit on the training distribution. In this work, we leverage self-supervised learning from longitudinal and multimodal archives to address these challenges. We curate an unlabeled set of patients with CT scans and linked electronic health records from our home institution to power joint embedding predictive architecture (JEPA) pretraining. After supervised finetuning, we show that our approach outperforms an unregularized multimodal model and imaging-only model in an internal cohort (ours: 0.91, multimodal: 0.88, imaging-only: 0.73 AUC), but underperforms in an external cohort (ours: 0.72, imaging-only: 0.75 AUC). We develop a synthetic environment that characterizes the context in which JEPA may underperform. This work innovates an approach that leverages unlabeled multimodal medical archives to improve predictive models and demonstrates its advantages and limitations in pulmonary nodule diagnosis.
Lifespan Pancreas Morphology for Control vs Type 2 Diabetes using AI on Largescale Clinical Imaging
Aug 20, 2025Abstract:Purpose: Understanding how the pancreas changes is critical for detecting deviations in type 2 diabetes and other pancreatic disease. We measure pancreas size and shape using morphological measurements from ages 0 to 90. Our goals are to 1) identify reliable clinical imaging modalities for AI-based pancreas measurement, 2) establish normative morphological aging trends, and 3) detect potential deviations in type 2 diabetes. Approach: We analyzed a clinically acquired dataset of 2533 patients imaged with abdominal CT or MRI. We resampled the scans to 3mm isotropic resolution, segmented the pancreas using automated methods, and extracted 13 morphological pancreas features across the lifespan. First, we assessed CT and MRI measurements to determine which modalities provide consistent lifespan trends. Second, we characterized distributions of normative morphological patterns stratified by age group and sex. Third, we used GAMLSS regression to model pancreas morphology trends in 1350 patients matched for age, sex, and type 2 diabetes status to identify any deviations from normative aging associated with type 2 diabetes. Results: When adjusting for confounders, the aging trends for 10 of 13 morphological features were significantly different between patients with type 2 diabetes and non-diabetic controls (p < 0.05 after multiple comparisons corrections). Additionally, MRI appeared to yield different pancreas measurements than CT using our AI-based method. Conclusions: We provide lifespan trends demonstrating that the size and shape of the pancreas is altered in type 2 diabetes using 675 control patients and 675 diabetes patients. Moreover, our findings reinforce that the pancreas is smaller in type 2 diabetes. Additionally, we contribute a reference of lifespan pancreas morphology from a large cohort of non-diabetic control patients in a clinical setting.
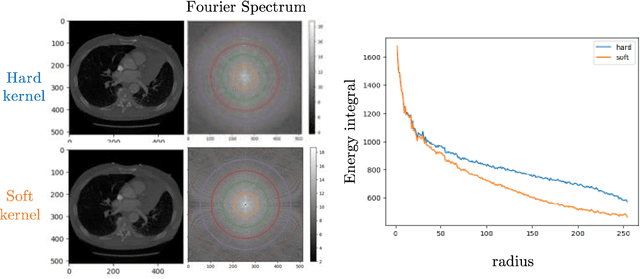
Multipath cycleGAN for harmonization of paired and unpaired low-dose lung computed tomography reconstruction kernels
May 28, 2025Abstract:Reconstruction kernels in computed tomography (CT) affect spatial resolution and noise characteristics, introducing systematic variability in quantitative imaging measurements such as emphysema quantification. Choosing an appropriate kernel is therefore essential for consistent quantitative analysis. We propose a multipath cycleGAN model for CT kernel harmonization, trained on a mixture of paired and unpaired data from a low-dose lung cancer screening cohort. The model features domain-specific encoders and decoders with a shared latent space and uses discriminators tailored for each domain.We train the model on 42 kernel combinations using 100 scans each from seven representative kernels in the National Lung Screening Trial (NLST) dataset. To evaluate performance, 240 scans from each kernel are harmonized to a reference soft kernel, and emphysema is quantified before and after harmonization. A general linear model assesses the impact of age, sex, smoking status, and kernel on emphysema. We also evaluate harmonization from soft kernels to a reference hard kernel. To assess anatomical consistency, we compare segmentations of lung vessels, muscle, and subcutaneous adipose tissue generated by TotalSegmentator between harmonized and original images. Our model is benchmarked against traditional and switchable cycleGANs. For paired kernels, our approach reduces bias in emphysema scores, as seen in Bland-Altman plots (p<0.05). For unpaired kernels, harmonization eliminates confounding differences in emphysema (p>0.05). High Dice scores confirm preservation of muscle and fat anatomy, while lung vessel overlap remains reasonable. Overall, our shared latent space multipath cycleGAN enables robust harmonization across paired and unpaired CT kernels, improving emphysema quantification and preserving anatomical fidelity.
Investigating the impact of kernel harmonization and deformable registration on inspiratory and expiratory chest CT images for people with COPD
Feb 07, 2025



Abstract:Paired inspiratory-expiratory CT scans enable the quantification of gas trapping due to small airway disease and emphysema by analyzing lung tissue motion in COPD patients. Deformable image registration of these scans assesses regional lung volumetric changes. However, variations in reconstruction kernels between paired scans introduce errors in quantitative analysis. This work proposes a two-stage pipeline to harmonize reconstruction kernels and perform deformable image registration using data acquired from the COPDGene study. We use a cycle generative adversarial network (GAN) to harmonize inspiratory scans reconstructed with a hard kernel (BONE) to match expiratory scans reconstructed with a soft kernel (STANDARD). We then deformably register the expiratory scans to inspiratory scans. We validate harmonization by measuring emphysema using a publicly available segmentation algorithm before and after harmonization. Results show harmonization significantly reduces emphysema measurement inconsistencies, decreasing median emphysema scores from 10.479% to 3.039%, with a reference median score of 1.305% from the STANDARD kernel as the target. Registration accuracy is evaluated via Dice overlap between emphysema regions on inspiratory, expiratory, and deformed images. The Dice coefficient between inspiratory emphysema masks and deformably registered emphysema masks increases significantly across registration stages (p<0.001). Additionally, we demonstrate that deformable registration is robust to kernel variations.
Brain age identification from diffusion MRI synergistically predicts neurodegenerative disease
Oct 29, 2024



Abstract:Estimated brain age from magnetic resonance image (MRI) and its deviation from chronological age can provide early insights into potential neurodegenerative diseases, supporting early detection and implementation of prevention strategies. Diffusion MRI (dMRI), a widely used modality for brain age estimation, presents an opportunity to build an earlier biomarker for neurodegenerative disease prediction because it captures subtle microstructural changes that precede more perceptible macrostructural changes. However, the coexistence of macro- and micro-structural information in dMRI raises the question of whether current dMRI-based brain age estimation models are leveraging the intended microstructural information or if they inadvertently rely on the macrostructural information. To develop a microstructure-specific brain age, we propose a method for brain age identification from dMRI that minimizes the model's use of macrostructural information by non-rigidly registering all images to a standard template. Imaging data from 13,398 participants across 12 datasets were used for the training and evaluation. We compare our brain age models, trained with and without macrostructural information minimized, with an architecturally similar T1-weighted (T1w) MRI-based brain age model and two state-of-the-art T1w MRI-based brain age models that primarily use macrostructural information. We observe difference between our dMRI-based brain age and T1w MRI-based brain age across stages of neurodegeneration, with dMRI-based brain age being older than T1w MRI-based brain age in participants transitioning from cognitively normal (CN) to mild cognitive impairment (MCI), but younger in participants already diagnosed with Alzheimer's disease (AD). Approximately 4 years before MCI diagnosis, dMRI-based brain age yields better performance than T1w MRI-based brain ages in predicting transition from CN to MCI.
Sensitivity of quantitative diffusion MRI tractography and microstructure to anisotropic spatial sampling
Sep 26, 2024Abstract:Purpose: Diffusion weighted MRI (dMRI) and its models of neural structure provide insight into human brain organization and variations in white matter. A recent study by McMaster, et al. showed that complex graph measures of the connectome, the graphical representation of a tractogram, vary with spatial sampling changes, but biases introduced by anisotropic voxels in the process have not been well characterized. This study uses microstructural measures (fractional anisotropy and mean diffusivity) and white matter bundle properties (bundle volume, length, and surface area) to further understand the effect of anisotropic voxels on microstructure and tractography. Methods: The statistical significance of the selected measures derived from dMRI data were assessed by comparing three white matter bundles at different spatial resolutions with 44 subjects from the Human Connectome Project Young Adult dataset scan/rescan data using the Wilcoxon Signed Rank test. The original isotropic resolution (1.25 mm isotropic) was explored with six anisotropic resolutions with 0.25 mm incremental steps in the z dimension. Then, all generated resolutions were upsampled to 1.25 mm isotropic and 1 mm isotropic. Results: There were statistically significant differences between at least one microstructural and one bundle measure at every resolution (p less than or equal to 0.05, corrected for multiple comparisons). Cohen's d coefficient evaluated the effect size of anisotropic voxels on microstructure and tractography. Conclusion: Fractional anisotropy and mean diffusivity cannot be recovered with basic up sampling from low quality data with gold standard data. However, the bundle measures from tractogram become more repeatable when voxels are resampled to 1 mm isotropic.
Harmonized connectome resampling for variance in voxel sizes
Aug 02, 2024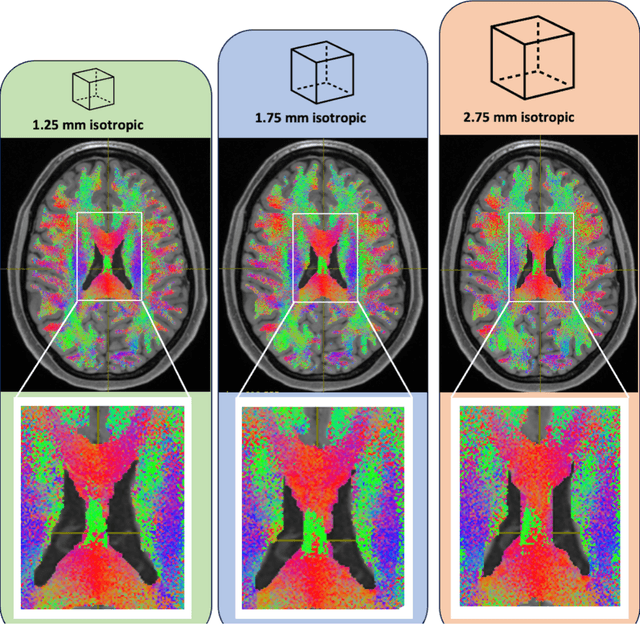
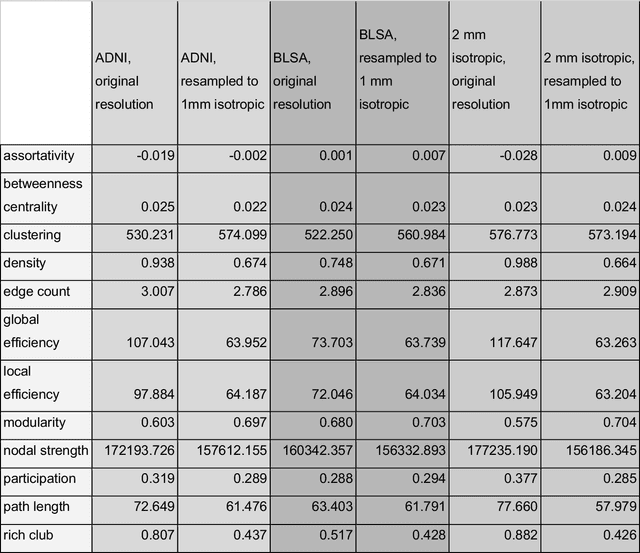
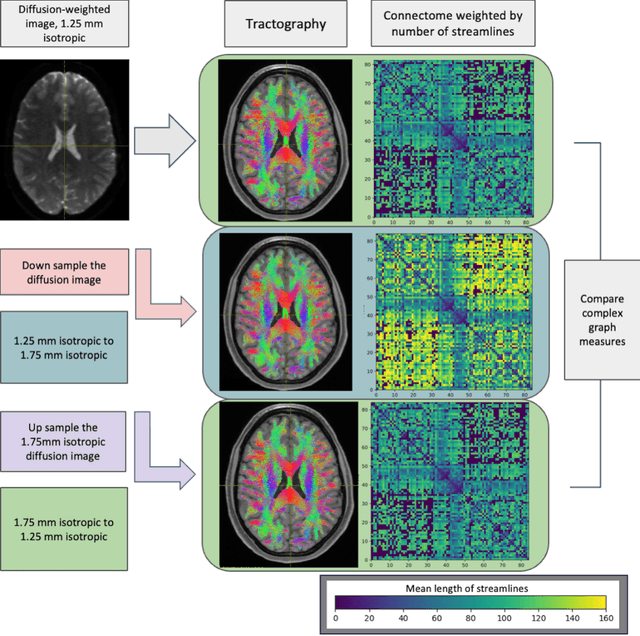
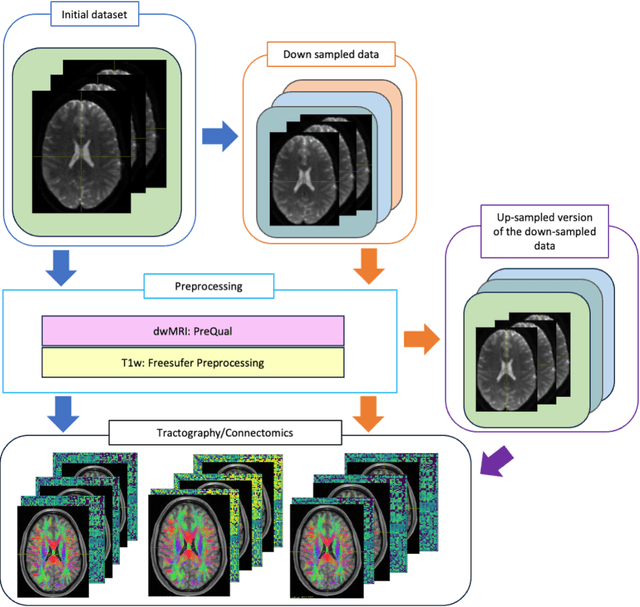
Abstract:To date, there has been no comprehensive study characterizing the effect of diffusion-weighted magnetic resonance imaging voxel resolution on the resulting connectome for high resolution subject data. Similarity in results improved with higher resolution, even after initial down-sampling. To ensure robust tractography and connectomes, resample data to 1 mm isotropic resolution.
Evaluation of Mean Shift, ComBat, and CycleGAN for Harmonizing Brain Connectivity Matrices Across Sites
Jan 24, 2024



Abstract:Connectivity matrices derived from diffusion MRI (dMRI) provide an interpretable and generalizable way of understanding the human brain connectome. However, dMRI suffers from inter-site and between-scanner variation, which impedes analysis across datasets to improve robustness and reproducibility of results. To evaluate different harmonization approaches on connectivity matrices, we compared graph measures derived from these matrices before and after applying three harmonization techniques: mean shift, ComBat, and CycleGAN. The sample comprises 168 age-matched, sex-matched normal subjects from two studies: the Vanderbilt Memory and Aging Project (VMAP) and the Biomarkers of Cognitive Decline Among Normal Individuals (BIOCARD). First, we plotted the graph measures and used coefficient of variation (CoV) and the Mann-Whitney U test to evaluate different methods' effectiveness in removing site effects on the matrices and the derived graph measures. ComBat effectively eliminated site effects for global efficiency and modularity and outperformed the other two methods. However, all methods exhibited poor performance when harmonizing average betweenness centrality. Second, we tested whether our harmonization methods preserved correlations between age and graph measures. All methods except for CycleGAN in one direction improved correlations between age and global efficiency and between age and modularity from insignificant to significant with p-values less than 0.05.
Inter-vendor harmonization of Computed Tomography (CT) reconstruction kernels using unpaired image translation
Sep 22, 2023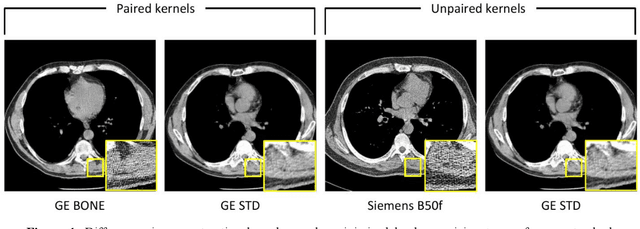
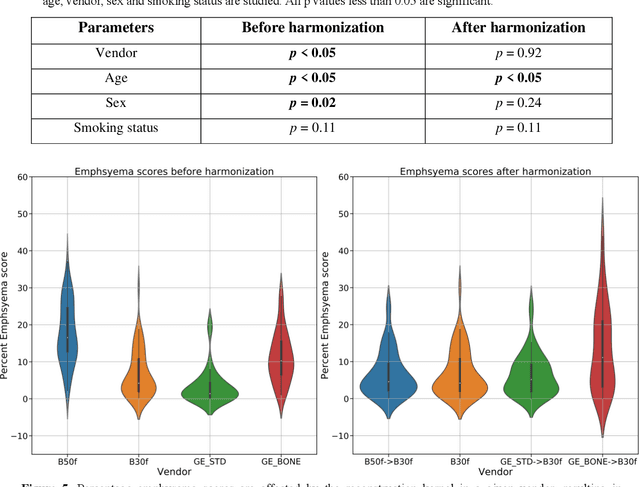
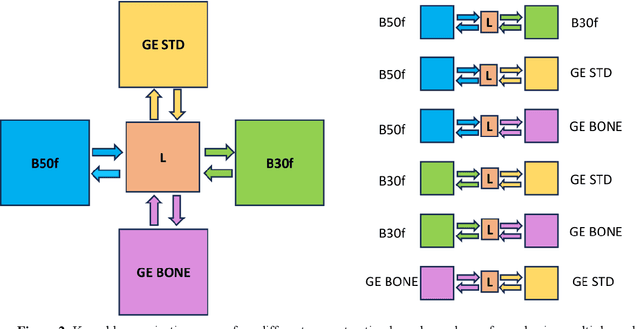
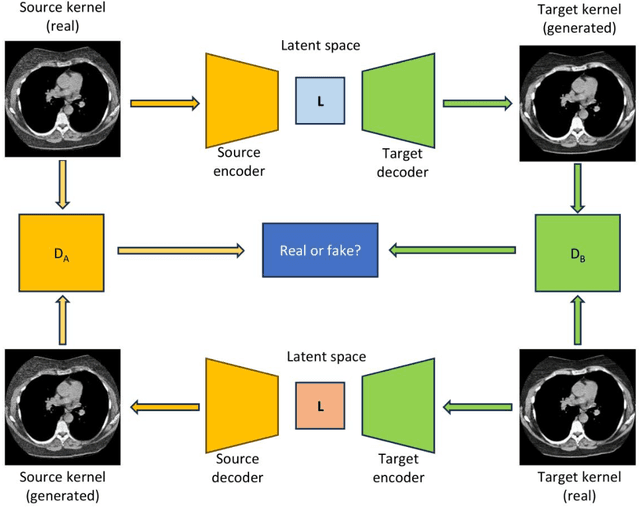
Abstract:The reconstruction kernel in computed tomography (CT) generation determines the texture of the image. Consistency in reconstruction kernels is important as the underlying CT texture can impact measurements during quantitative image analysis. Harmonization (i.e., kernel conversion) minimizes differences in measurements due to inconsistent reconstruction kernels. Existing methods investigate harmonization of CT scans in single or multiple manufacturers. However, these methods require paired scans of hard and soft reconstruction kernels that are spatially and anatomically aligned. Additionally, a large number of models need to be trained across different kernel pairs within manufacturers. In this study, we adopt an unpaired image translation approach to investigate harmonization between and across reconstruction kernels from different manufacturers by constructing a multipath cycle generative adversarial network (GAN). We use hard and soft reconstruction kernels from the Siemens and GE vendors from the National Lung Screening Trial dataset. We use 50 scans from each reconstruction kernel and train a multipath cycle GAN. To evaluate the effect of harmonization on the reconstruction kernels, we harmonize 50 scans each from Siemens hard kernel, GE soft kernel and GE hard kernel to a reference Siemens soft kernel (B30f) and evaluate percent emphysema. We fit a linear model by considering the age, smoking status, sex and vendor and perform an analysis of variance (ANOVA) on the emphysema scores. Our approach minimizes differences in emphysema measurement and highlights the impact of age, sex, smoking status and vendor on emphysema quantification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge