Mirza S. Khan
Longitudinal Multimodal Transformer Integrating Imaging and Latent Clinical Signatures From Routine EHRs for Pulmonary Nodule Classification
Apr 10, 2023



Abstract:The accuracy of predictive models for solitary pulmonary nodule (SPN) diagnosis can be greatly increased by incorporating repeat imaging and medical context, such as electronic health records (EHRs). However, clinically routine modalities such as imaging and diagnostic codes can be asynchronous and irregularly sampled over different time scales which are obstacles to longitudinal multimodal learning. In this work, we propose a transformer-based multimodal strategy to integrate repeat imaging with longitudinal clinical signatures from routinely collected EHRs for SPN classification. We perform unsupervised disentanglement of latent clinical signatures and leverage time-distance scaled self-attention to jointly learn from clinical signatures expressions and chest computed tomography (CT) scans. Our classifier is pretrained on 2,668 scans from a public dataset and 1,149 subjects with longitudinal chest CTs, billing codes, medications, and laboratory tests from EHRs of our home institution. Evaluation on 227 subjects with challenging SPNs revealed a significant AUC improvement over a longitudinal multimodal baseline (0.824 vs 0.752 AUC), as well as improvements over a single cross-section multimodal scenario (0.809 AUC) and a longitudinal imaging-only scenario (0.741 AUC). This work demonstrates significant advantages with a novel approach for co-learning longitudinal imaging and non-imaging phenotypes with transformers.
Body Composition Assessment with Limited Field-of-view Computed Tomography: A Semantic Image Extension Perspective
Jul 13, 2022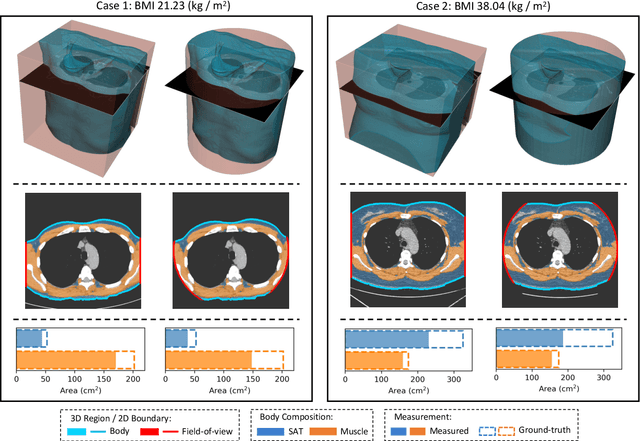
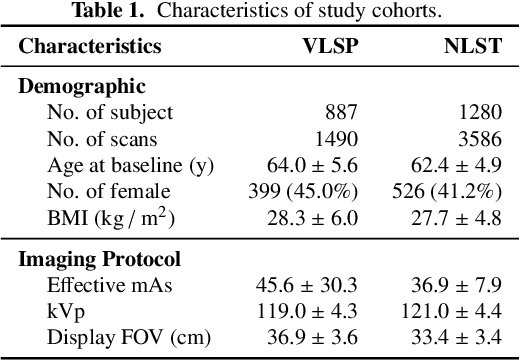
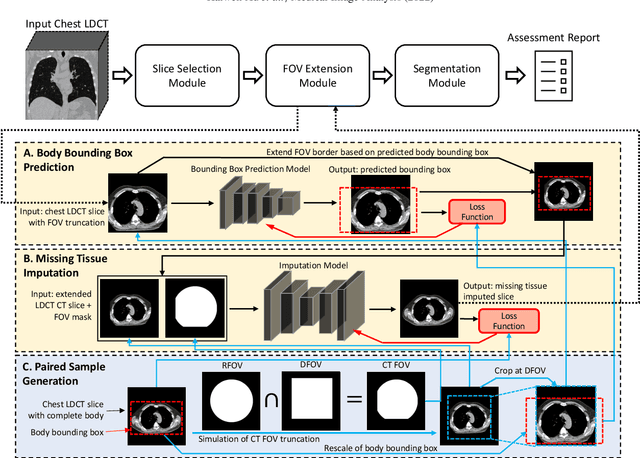
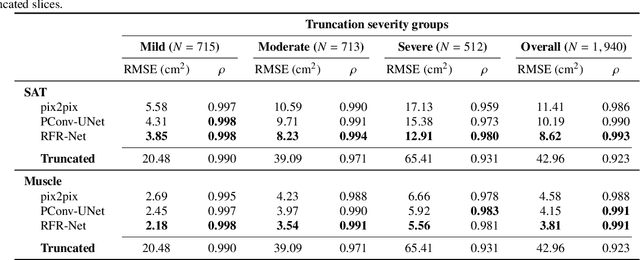
Abstract:Field-of-view (FOV) tissue truncation beyond the lungs is common in routine lung screening computed tomography (CT). This poses limitations for opportunistic CT- based body composition (BC) assessment as key anatomical structures are missing. Traditionally, extending the FOV of CT is considered as a CT reconstruction problem using limited data. However, this approach relies on the projection domain data which might not be available in application. In this work, we formulate the problem from the semantic image extension perspective which only requires image data as inputs. The proposed two-stage method identifies a new FOV border based on the estimated extent of the complete body and imputes missing tissues in the truncated region. The training samples are simulated using CT slices with complete body in FOV, making the model development self-supervised. We evaluate the validity of the proposed method in automatic BC assessment using lung screening CT with limited FOV. The proposed method effectively restores the missing tissues and reduces BC assessment error introduced by FOV tissue truncation. In the BC assessment for a large-scale lung screening CT dataset, this correction improves both the intra-subject consistency and the correlation with anthropometric approximations. The developed method is available at https://github.com/MASILab/S-EFOV.
Technical Report: Quality Assessment Tool for Machine Learning with Clinical CT
Jul 27, 2021



Abstract:Image Quality Assessment (IQA) is important for scientific inquiry, especially in medical imaging and machine learning. Potential data quality issues can be exacerbated when human-based workflows use limited views of the data that may obscure digital artifacts. In practice, multiple factors such as network issues, accelerated acquisitions, motion artifacts, and imaging protocol design can impede the interpretation of image collections. The medical image processing community has developed a wide variety of tools for the inspection and validation of imaging data. Yet, IQA of computed tomography (CT) remains an under-recognized challenge, and no user-friendly tool is commonly available to address these potential issues. Here, we create and illustrate a pipeline specifically designed to identify and resolve issues encountered with large-scale data mining of clinically acquired CT data. Using the widely studied National Lung Screening Trial (NLST), we have identified approximately 4% of image volumes with quality concerns out of 17,392 scans. To assess robustness, we applied the proposed pipeline to our internal datasets where we find our tool is generalizable to clinically acquired medical images. In conclusion, the tool has been useful and time-saving for research study of clinical data, and the code and tutorials are publicly available at https://github.com/MASILab/QA_tool.
Development and Characterization of a Chest CT Atlas
Dec 05, 2020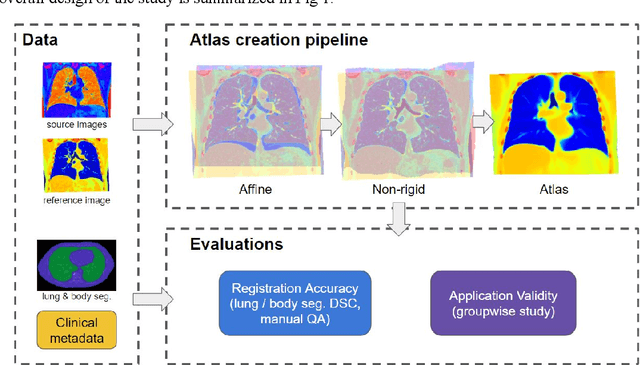
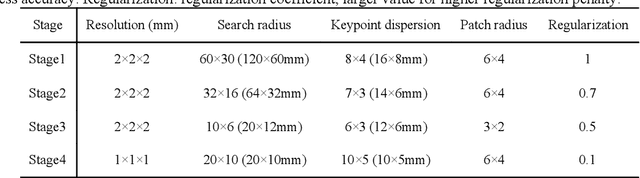

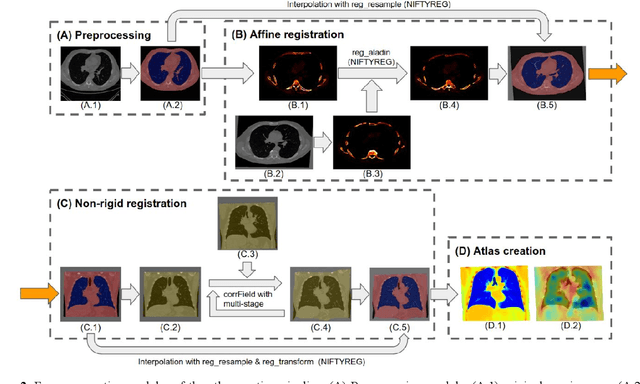
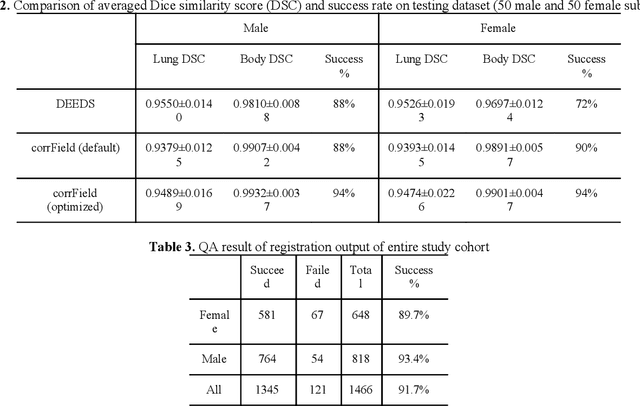
Abstract:A major goal of lung cancer screening is to identify individuals with particular phenotypes that are associated with high risk of cancer. Identifying relevant phenotypes is complicated by the variation in body position and body composition. In the brain, standardized coordinate systems (e.g., atlases) have enabled separate consideration of local features from gross/global structure. To date, no analogous standard atlas has been presented to enable spatial mapping and harmonization in chest computational tomography (CT). In this paper, we propose a thoracic atlas built upon a large low dose CT (LDCT) database of lung cancer screening program. The study cohort includes 466 male and 387 female subjects with no screening detected malignancy (age 46-79 years, mean 64.9 years). To provide spatial mapping, we optimize a multi-stage inter-subject non-rigid registration pipeline for the entire thoracic space. We evaluate the optimized pipeline relative to two baselines with alternative non-rigid registration module: the same software with default parameters and an alternative software. We achieve a significant improvement in terms of registration success rate based on manual QA. For the entire study cohort, the optimized pipeline achieves a registration success rate of 91.7%. The application validity of the developed atlas is evaluated in terms of discriminative capability for different anatomic phenotypes, including body mass index (BMI), chronic obstructive pulmonary disease (COPD), and coronary artery calcification (CAC).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge