Gaurav Rudravaram
Personalized White Matter Bundle Segmentation for Early Childhood
Feb 04, 2026Abstract:White matter segmentation methods from diffusion magnetic resonance imaging range from streamline clustering-based approaches to bundle mask delineation, but none have proposed a pediatric-specific approach. We hypothesize that a deep learning model with a similar approach to TractSeg will improve similarity between an algorithm-generated mask and an expert-labeled ground truth. Given a cohort of 56 manually labelled white matter bundles, we take inspiration from TractSeg's 2D UNet architecture, and we modify inputs to match bundle definitions as determined by pediatric experts, evaluation to use k fold cross validation, the loss function to masked Dice loss. We evaluate Dice score, volume overlap, and volume overreach of 16 major regions of interest compared to the expert labeled dataset. To test whether our approach offers statistically significant improvements over TractSeg, we compare Dice voxels, volume overlap, and adjacency voxels with a Wilcoxon signed rank test followed by false discovery rate correction. We find statistical significance across all bundles for all metrics with one exception in volume overlap. After we run TractSeg and our model, we combine their output masks into a 60 label atlas to evaluate if TractSeg and our model combined can generate a robust, individualized atlas, and observe smoothed, continuous masks in cases that TractSeg did not produce an anatomically plausible output. With the improvement of white matter pathway segmentation masks, we can further understand neurodevelopment on a population level scale, and we can produce reliable estimates of individualized anatomy in pediatric white matter diseases and disorders.
Fully Differentiable dMRI Streamline Propagation in PyTorch
Nov 17, 2025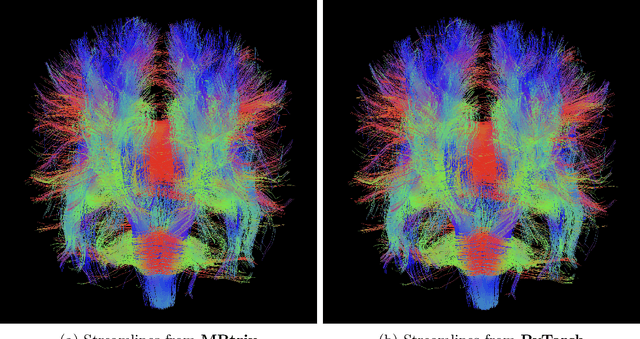
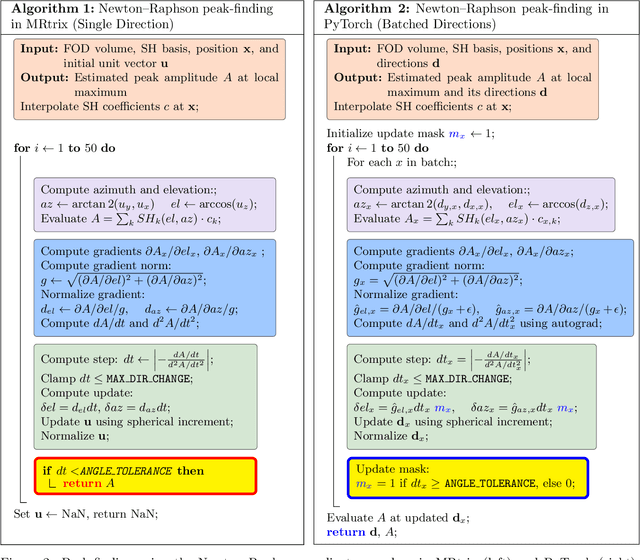
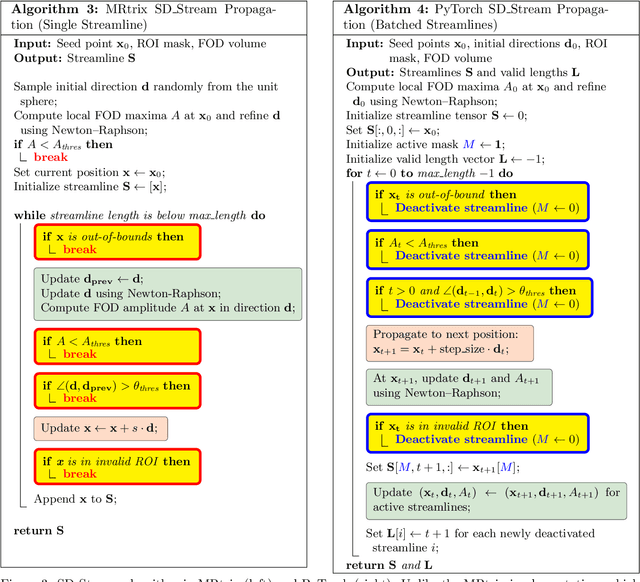
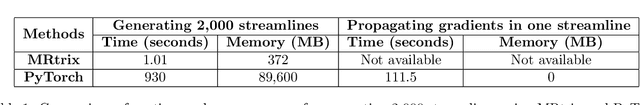


Abstract:Diffusion MRI (dMRI) provides a distinctive means to probe the microstructural architecture of living tissue, facilitating applications such as brain connectivity analysis, modeling across multiple conditions, and the estimation of macrostructural features. Tractography, which emerged in the final years of the 20th century and accelerated in the early 21st century, is a technique for visualizing white matter pathways in the brain using dMRI. Most diffusion tractography methods rely on procedural streamline propagators or global energy minimization methods. Although recent advancements in deep learning have enabled tasks that were previously challenging, existing tractography approaches are often non-differentiable, limiting their integration in end-to-end learning frameworks. While progress has been made in representing streamlines in differentiable frameworks, no existing method offers fully differentiable propagation. In this work, we propose a fully differentiable solution that retains numerical fidelity with a leading streamline algorithm. The key is that our PyTorch-engineered streamline propagator has no components that block gradient flow, making it fully differentiable. We show that our method matches standard propagators while remaining differentiable. By translating streamline propagation into a differentiable PyTorch framework, we enable deeper integration of tractography into deep learning workflows, laying the foundation for a new category of macrostructural reasoning that is not only computationally robust but also scientifically rigorous.
Phenotype discovery of traumatic brain injury segmentations from heterogeneous multi-site data
Nov 05, 2025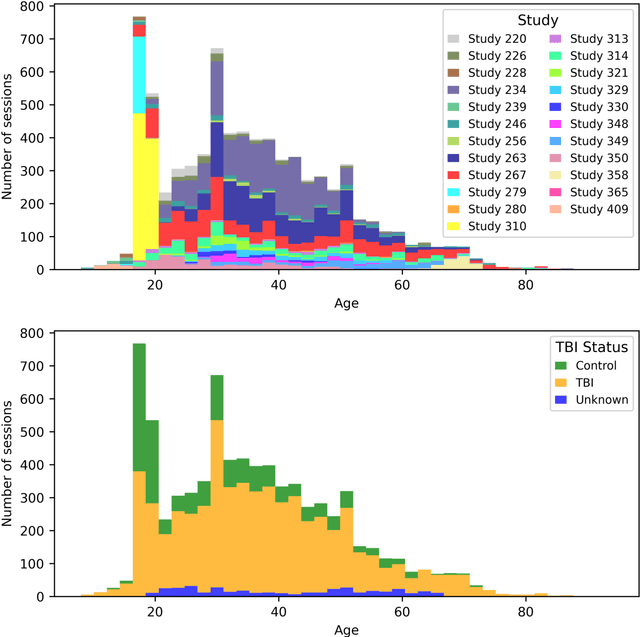
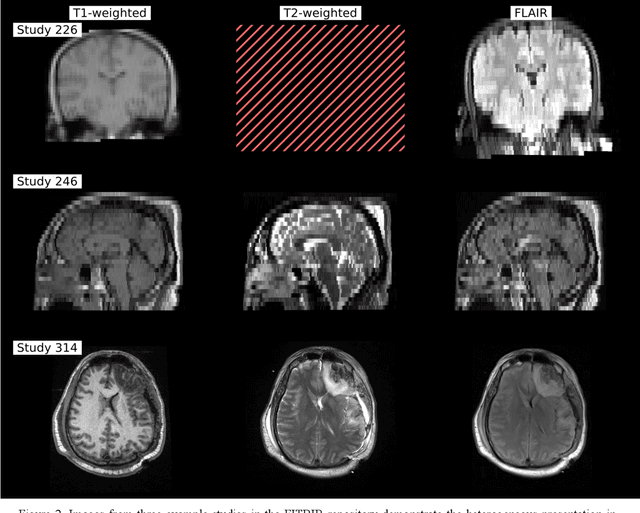
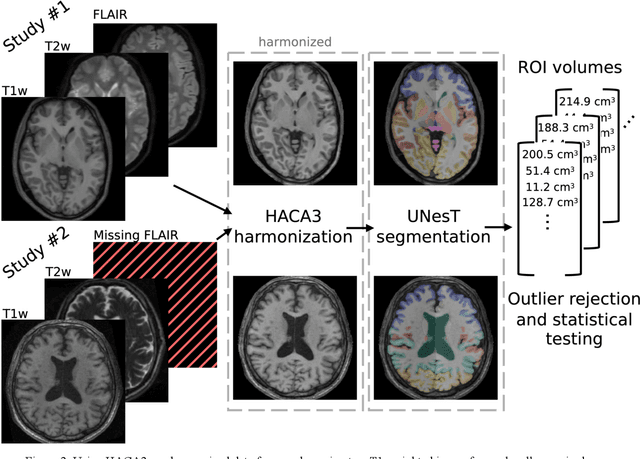
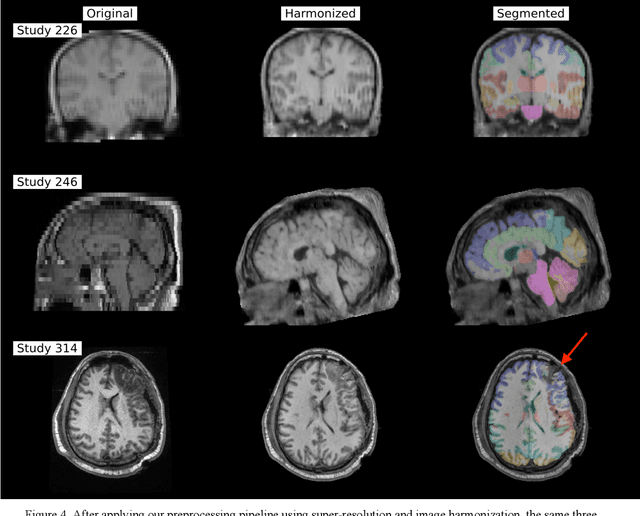
Abstract:Traumatic brain injury (TBI) is intrinsically heterogeneous, and typical clinical outcome measures like the Glasgow Coma Scale complicate this diversity. The large variability in severity and patient outcomes render it difficult to link structural damage to functional deficits. The Federal Interagency Traumatic Brain Injury Research (FITBIR) repository contains large-scale multi-site magnetic resonance imaging data of varying resolutions and acquisition parameters (25 shared studies with 7,693 sessions that have age, sex and TBI status defined - 5,811 TBI and 1,882 controls). To reveal shared pathways of injury of TBI through imaging, we analyzed T1-weighted images from these sessions by first harmonizing to a local dataset and segmenting 132 regions of interest (ROIs) in the brain. After running quality assurance, calculating the volumes of the ROIs, and removing outliers, we calculated the z-scores of volumes for all participants relative to the mean and standard deviation of the controls. We regressed out sex, age, and total brain volume with a multivariate linear regression, and we found significant differences in 37 ROIs between subjects with TBI and controls (p < 0.05 with independent t-tests with false discovery rate correction). We found that differences originated in 1) the brainstem, occipital pole and structures posterior to the orbit, 2) subcortical gray matter and insular cortex, and 3) cerebral and cerebellar white matter using independent component analysis and clustering the component loadings of those with TBI.
DeepFixel: Crossing white matter fiber identification through spherical convolutional neural networks
Nov 05, 2025Abstract:Diffusion-weighted magnetic resonance imaging allows for reconstruction of models for structural connectivity in the brain, such as fiber orientation distribution functions (ODFs) that describe the distribution, direction, and volume of white matter fiber bundles in a voxel. Crossing white matter fibers in voxels complicate analysis and can lead to errors in downstream tasks like tractography. We introduce one option for separating fiber ODFs by performing a nonlinear optimization to fit ODFs to the given data and penalizing terms that are not symmetric about the axis of the fiber. However, this optimization is non-convex and computationally infeasible across an entire image (approximately 1.01 x 106 ms per voxel). We introduce DeepFixel, a spherical convolutional neural network approximation for this nonlinear optimization. We model the probability distribution of fibers as a spherical mesh with higher angular resolution than a truncated spherical harmonic representation. To validate DeepFixel, we compare to the nonlinear optimization and a fixel-based separation algorithm of two-fiber and three-fiber ODFs. The median angular correlation coefficient is 1 (interquartile range of 0.00) using the nonlinear optimization algorithm, 0.988 (0.317) using a fiber bundle elements or "fixel"-based separation algorithm, and 0.973 (0.004) using DeepFixel. DeepFixel is more computationally efficient than the non-convex optimization (0.32 ms per voxel). DeepFixel's spherical mesh representation is successful at disentangling at smaller angular separations and smaller volume fractions than the fixel-based separation algorithm.
Lifespan Pancreas Morphology for Control vs Type 2 Diabetes using AI on Largescale Clinical Imaging
Aug 20, 2025Abstract:Purpose: Understanding how the pancreas changes is critical for detecting deviations in type 2 diabetes and other pancreatic disease. We measure pancreas size and shape using morphological measurements from ages 0 to 90. Our goals are to 1) identify reliable clinical imaging modalities for AI-based pancreas measurement, 2) establish normative morphological aging trends, and 3) detect potential deviations in type 2 diabetes. Approach: We analyzed a clinically acquired dataset of 2533 patients imaged with abdominal CT or MRI. We resampled the scans to 3mm isotropic resolution, segmented the pancreas using automated methods, and extracted 13 morphological pancreas features across the lifespan. First, we assessed CT and MRI measurements to determine which modalities provide consistent lifespan trends. Second, we characterized distributions of normative morphological patterns stratified by age group and sex. Third, we used GAMLSS regression to model pancreas morphology trends in 1350 patients matched for age, sex, and type 2 diabetes status to identify any deviations from normative aging associated with type 2 diabetes. Results: When adjusting for confounders, the aging trends for 10 of 13 morphological features were significantly different between patients with type 2 diabetes and non-diabetic controls (p < 0.05 after multiple comparisons corrections). Additionally, MRI appeared to yield different pancreas measurements than CT using our AI-based method. Conclusions: We provide lifespan trends demonstrating that the size and shape of the pancreas is altered in type 2 diabetes using 675 control patients and 675 diabetes patients. Moreover, our findings reinforce that the pancreas is smaller in type 2 diabetes. Additionally, we contribute a reference of lifespan pancreas morphology from a large cohort of non-diabetic control patients in a clinical setting.
Multipath cycleGAN for harmonization of paired and unpaired low-dose lung computed tomography reconstruction kernels
May 28, 2025Abstract:Reconstruction kernels in computed tomography (CT) affect spatial resolution and noise characteristics, introducing systematic variability in quantitative imaging measurements such as emphysema quantification. Choosing an appropriate kernel is therefore essential for consistent quantitative analysis. We propose a multipath cycleGAN model for CT kernel harmonization, trained on a mixture of paired and unpaired data from a low-dose lung cancer screening cohort. The model features domain-specific encoders and decoders with a shared latent space and uses discriminators tailored for each domain.We train the model on 42 kernel combinations using 100 scans each from seven representative kernels in the National Lung Screening Trial (NLST) dataset. To evaluate performance, 240 scans from each kernel are harmonized to a reference soft kernel, and emphysema is quantified before and after harmonization. A general linear model assesses the impact of age, sex, smoking status, and kernel on emphysema. We also evaluate harmonization from soft kernels to a reference hard kernel. To assess anatomical consistency, we compare segmentations of lung vessels, muscle, and subcutaneous adipose tissue generated by TotalSegmentator between harmonized and original images. Our model is benchmarked against traditional and switchable cycleGANs. For paired kernels, our approach reduces bias in emphysema scores, as seen in Bland-Altman plots (p<0.05). For unpaired kernels, harmonization eliminates confounding differences in emphysema (p>0.05). High Dice scores confirm preservation of muscle and fat anatomy, while lung vessel overlap remains reasonable. Overall, our shared latent space multipath cycleGAN enables robust harmonization across paired and unpaired CT kernels, improving emphysema quantification and preserving anatomical fidelity.
CASC-AI: Consensus-aware Self-corrective AI Agents for Noise Cell Segmentation
Feb 11, 2025Abstract:Multi-class cell segmentation in high-resolution gigapixel whole slide images (WSI) is crucial for various clinical applications. However, training such models typically requires labor-intensive, pixel-wise annotations by domain experts. Recent efforts have democratized this process by involving lay annotators without medical expertise. However, conventional non-agent-based approaches struggle to handle annotation noise adaptively, as they lack mechanisms to mitigate false positives (FP) and false negatives (FN) at both the image-feature and pixel levels. In this paper, we propose a consensus-aware self-corrective AI agent that leverages the Consensus Matrix to guide its learning process. The Consensus Matrix defines regions where both the AI and annotators agree on cell and non-cell annotations, which are prioritized with stronger supervision. Conversely, areas of disagreement are adaptively weighted based on their feature similarity to high-confidence agreement regions, with more similar regions receiving greater attention. Additionally, contrastive learning is employed to separate features of noisy regions from those of reliable agreement regions by maximizing their dissimilarity. This paradigm enables the AI to iteratively refine noisy labels, enhancing its robustness. Validated on one real-world lay-annotated cell dataset and two simulated noisy datasets, our method demonstrates improved segmentation performance, effectively correcting FP and FN errors and showcasing its potential for training robust models on noisy datasets. The official implementation and cell annotations are publicly available at https://github.com/ddrrnn123/CASC-AI.
Investigating the impact of kernel harmonization and deformable registration on inspiratory and expiratory chest CT images for people with COPD
Feb 07, 2025



Abstract:Paired inspiratory-expiratory CT scans enable the quantification of gas trapping due to small airway disease and emphysema by analyzing lung tissue motion in COPD patients. Deformable image registration of these scans assesses regional lung volumetric changes. However, variations in reconstruction kernels between paired scans introduce errors in quantitative analysis. This work proposes a two-stage pipeline to harmonize reconstruction kernels and perform deformable image registration using data acquired from the COPDGene study. We use a cycle generative adversarial network (GAN) to harmonize inspiratory scans reconstructed with a hard kernel (BONE) to match expiratory scans reconstructed with a soft kernel (STANDARD). We then deformably register the expiratory scans to inspiratory scans. We validate harmonization by measuring emphysema using a publicly available segmentation algorithm before and after harmonization. Results show harmonization significantly reduces emphysema measurement inconsistencies, decreasing median emphysema scores from 10.479% to 3.039%, with a reference median score of 1.305% from the STANDARD kernel as the target. Registration accuracy is evaluated via Dice overlap between emphysema regions on inspiratory, expiratory, and deformed images. The Dice coefficient between inspiratory emphysema masks and deformably registered emphysema masks increases significantly across registration stages (p<0.001). Additionally, we demonstrate that deformable registration is robust to kernel variations.
Sensitivity of quantitative diffusion MRI tractography and microstructure to anisotropic spatial sampling
Sep 26, 2024Abstract:Purpose: Diffusion weighted MRI (dMRI) and its models of neural structure provide insight into human brain organization and variations in white matter. A recent study by McMaster, et al. showed that complex graph measures of the connectome, the graphical representation of a tractogram, vary with spatial sampling changes, but biases introduced by anisotropic voxels in the process have not been well characterized. This study uses microstructural measures (fractional anisotropy and mean diffusivity) and white matter bundle properties (bundle volume, length, and surface area) to further understand the effect of anisotropic voxels on microstructure and tractography. Methods: The statistical significance of the selected measures derived from dMRI data were assessed by comparing three white matter bundles at different spatial resolutions with 44 subjects from the Human Connectome Project Young Adult dataset scan/rescan data using the Wilcoxon Signed Rank test. The original isotropic resolution (1.25 mm isotropic) was explored with six anisotropic resolutions with 0.25 mm incremental steps in the z dimension. Then, all generated resolutions were upsampled to 1.25 mm isotropic and 1 mm isotropic. Results: There were statistically significant differences between at least one microstructural and one bundle measure at every resolution (p less than or equal to 0.05, corrected for multiple comparisons). Cohen's d coefficient evaluated the effect size of anisotropic voxels on microstructure and tractography. Conclusion: Fractional anisotropy and mean diffusivity cannot be recovered with basic up sampling from low quality data with gold standard data. However, the bundle measures from tractogram become more repeatable when voxels are resampled to 1 mm isotropic.
Multi-Modality Conditioned Variational U-Net for Field-of-View Extension in Brain Diffusion MRI
Sep 20, 2024
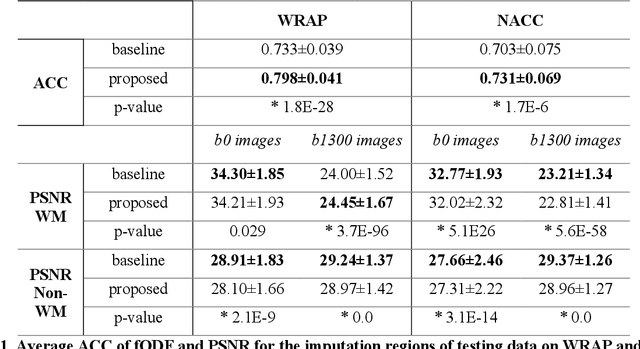

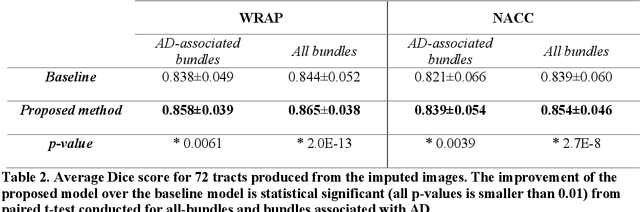
Abstract:An incomplete field-of-view (FOV) in diffusion magnetic resonance imaging (dMRI) can severely hinder the volumetric and bundle analyses of whole-brain white matter connectivity. Although existing works have investigated imputing the missing regions using deep generative models, it remains unclear how to specifically utilize additional information from paired multi-modality data and whether this can enhance the imputation quality and be useful for downstream tractography. To fill this gap, we propose a novel framework for imputing dMRI scans in the incomplete part of the FOV by integrating the learned diffusion features in the acquired part of the FOV to the complete brain anatomical structure. We hypothesize that by this design the proposed framework can enhance the imputation performance of the dMRI scans and therefore be useful for repairing whole-brain tractography in corrupted dMRI scans with incomplete FOV. We tested our framework on two cohorts from different sites with a total of 96 subjects and compared it with a baseline imputation method that treats the information from T1w and dMRI scans equally. The proposed framework achieved significant improvements in imputation performance, as demonstrated by angular correlation coefficient (p < 1E-5), and in downstream tractography accuracy, as demonstrated by Dice score (p < 0.01). Results suggest that the proposed framework improved imputation performance in dMRI scans by specifically utilizing additional information from paired multi-modality data, compared with the baseline method. The imputation achieved by the proposed framework enhances whole brain tractography, and therefore reduces the uncertainty when analyzing bundles associated with neurodegenerative.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge