Xing Yao
Endo-SemiS: Towards Robust Semi-Supervised Image Segmentation for Endoscopic Video
Dec 18, 2025



Abstract:In this paper, we present Endo-SemiS, a semi-supervised segmentation framework for providing reliable segmentation of endoscopic video frames with limited annotation. EndoSemiS uses 4 strategies to improve performance by effectively utilizing all available data, particularly unlabeled data: (1) Cross-supervision between two individual networks that supervise each other; (2) Uncertainty-guided pseudo-labels from unlabeled data, which are generated by selecting high-confidence regions to improve their quality; (3) Joint pseudolabel supervision, which aggregates reliable pixels from the pseudo-labels of both networks to provide accurate supervision for unlabeled data; and (4) Mutual learning, where both networks learn from each other at the feature and image levels, reducing variance and guiding them toward a consistent solution. Additionally, a separate corrective network that utilizes spatiotemporal information from endoscopy video to improve segmentation performance. Endo-SemiS is evaluated on two clinical applications: kidney stone laser lithotomy from ureteroscopy and polyp screening from colonoscopy. Compared to state-of-the-art segmentation methods, Endo-SemiS substantially achieves superior results on both datasets with limited labeled data. The code is publicly available at https://github.com/MedICL-VU/Endo-SemiS
DEMIST: \underline{DE}coupled \underline{M}ulti-stream latent d\underline{I}ffusion for Quantitative Myelin Map \underline{S}yn\underline{T}hesis
Nov 16, 2025Abstract:Quantitative magnetization transfer (qMT) imaging provides myelin-sensitive biomarkers, such as the pool size ratio (PSR), which is valuable for multiple sclerosis (MS) assessment. However, qMT requires specialized 20-30 minute scans. We propose DEMIST to synthesize PSR maps from standard T1w and FLAIR images using a 3D latent diffusion model with three complementary conditioning mechanisms. Our approach has two stages: first, we train separate autoencoders for PSR and anatomical images to learn aligned latent representations. Second, we train a conditional diffusion model in this latent space on top of a frozen diffusion foundation backbone. Conditioning is decoupled into: (i) \textbf{semantic} tokens via cross-attention, (ii) \textbf{spatial} per-scale residual hints via a 3D ControlNet branch, and (iii) \textbf{adaptive} LoRA-modulated attention. We include edge-aware loss terms to preserve lesion boundaries and alignment losses to maintain quantitative consistency, while keeping the number of trainable parameters low and retaining the inductive bias of the pretrained model. We evaluate on 163 scans from 99 subjects using 5-fold cross-validation. Our method outperforms VAE, GAN and diffusion baselines on multiple metrics, producing sharper boundaries and better quantitative agreement with ground truth. Our code is publicly available at https://github.com/MedICL-VU/MS-Synthesis-3DcLDM.
Towards Better Ultrasound Video Segmentation Foundation Model: An Empirical study on SAM2 Finetuning from Data Perspective
Nov 07, 2025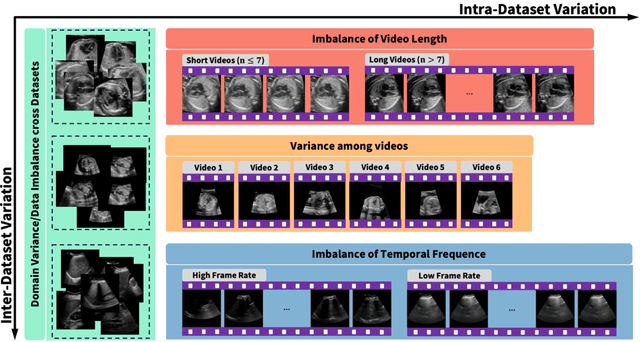
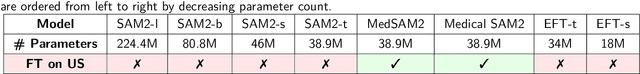

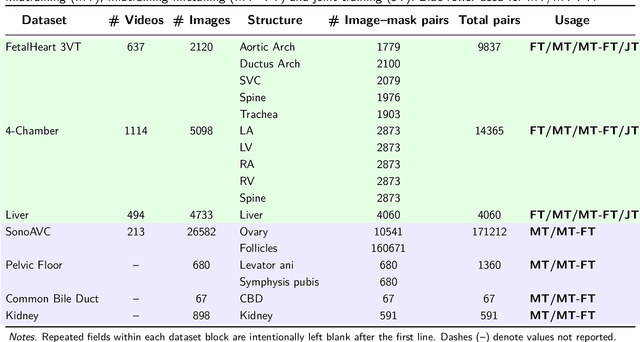
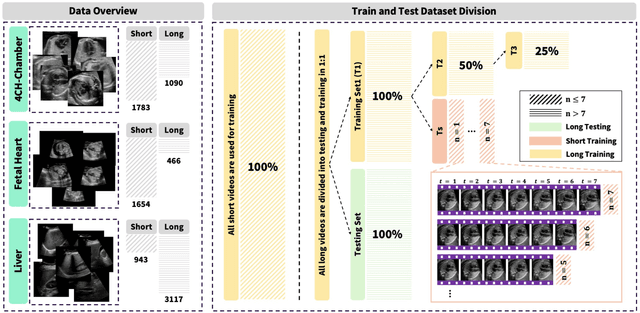
Abstract:Ultrasound (US) video segmentation remains a challenging problem due to strong inter- and intra-dataset variability, motion artifacts, and limited annotated data. Although foundation models such as Segment Anything Model 2 (SAM2) demonstrate strong zero-shot and prompt-guided segmentation capabilities, their performance deteriorates substantially when transferred to medical imaging domains. Current adaptation studies mainly emphasize architectural modifications, while the influence of data characteristics and training regimes has not been systematically examined. In this study, we present a comprehensive, data-centric investigation of SAM2 adaptation for ultrasound video segmentation. We analyze how training-set size, video duration, and augmentation schemes affect adaptation performance under three paradigms: task-specific fine-tuning, intermediate adaptation, and multi-task joint training, across five SAM2 variants and multiple prompting modes. We further design six ultrasound-specific augmentations, assessing their effect relative to generic strategies. Experiments on three representative ultrasound datasets reveal that data scale and temporal context play a more decisive role than model architecture or initialization. Moreover, joint training offers an efficient compromise between modality alignment and task specialization. This work aims to provide empirical insights for developing efficient, data-aware adaptation pipelines for SAM2 in ultrasound video analysis.
CASC-AI: Consensus-aware Self-corrective AI Agents for Noise Cell Segmentation
Feb 11, 2025Abstract:Multi-class cell segmentation in high-resolution gigapixel whole slide images (WSI) is crucial for various clinical applications. However, training such models typically requires labor-intensive, pixel-wise annotations by domain experts. Recent efforts have democratized this process by involving lay annotators without medical expertise. However, conventional non-agent-based approaches struggle to handle annotation noise adaptively, as they lack mechanisms to mitigate false positives (FP) and false negatives (FN) at both the image-feature and pixel levels. In this paper, we propose a consensus-aware self-corrective AI agent that leverages the Consensus Matrix to guide its learning process. The Consensus Matrix defines regions where both the AI and annotators agree on cell and non-cell annotations, which are prioritized with stronger supervision. Conversely, areas of disagreement are adaptively weighted based on their feature similarity to high-confidence agreement regions, with more similar regions receiving greater attention. Additionally, contrastive learning is employed to separate features of noisy regions from those of reliable agreement regions by maximizing their dissimilarity. This paradigm enables the AI to iteratively refine noisy labels, enhancing its robustness. Validated on one real-world lay-annotated cell dataset and two simulated noisy datasets, our method demonstrates improved segmentation performance, effectively correcting FP and FN errors and showcasing its potential for training robust models on noisy datasets. The official implementation and cell annotations are publicly available at https://github.com/ddrrnn123/CASC-AI.
SynStitch: a Self-Supervised Learning Network for Ultrasound Image Stitching Using Synthetic Training Pairs and Indirect Supervision
Nov 11, 2024
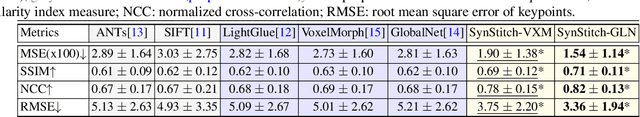
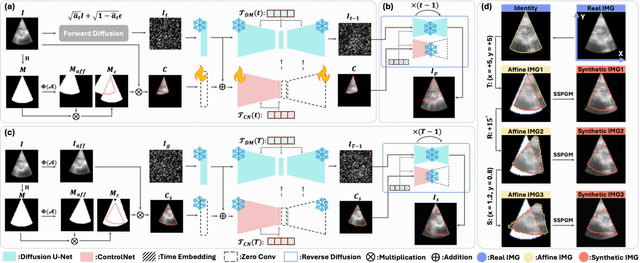

Abstract:Ultrasound (US) image stitching can expand the field-of-view (FOV) by combining multiple US images from varied probe positions. However, registering US images with only partially overlapping anatomical contents is a challenging task. In this work, we introduce SynStitch, a self-supervised framework designed for 2DUS stitching. SynStitch consists of a synthetic stitching pair generation module (SSPGM) and an image stitching module (ISM). SSPGM utilizes a patch-conditioned ControlNet to generate realistic 2DUS stitching pairs with known affine matrix from a single input image. ISM then utilizes this synthetic paired data to learn 2DUS stitching in a supervised manner. Our framework was evaluated against multiple leading methods on a kidney ultrasound dataset, demonstrating superior 2DUS stitching performance through both qualitative and quantitative analyses. The code will be made public upon acceptance of the paper.
Are Large Language Models Ready for Travel Planning?
Oct 22, 2024
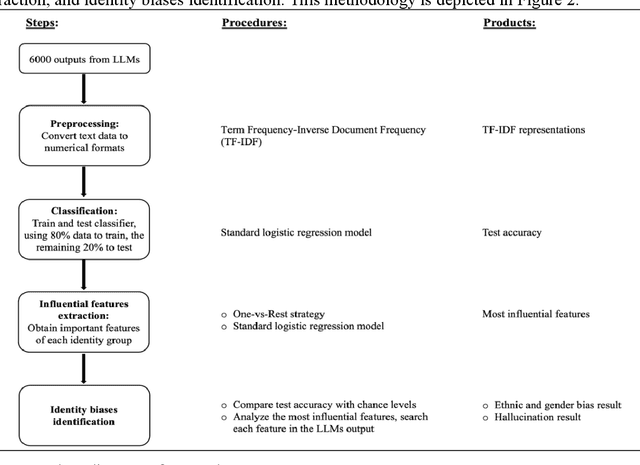

Abstract:While large language models (LLMs) show promise in hospitality and tourism, their ability to provide unbiased service across demographic groups remains unclear. This paper explores gender and ethnic biases when LLMs are utilized as travel planning assistants. To investigate this issue, we apply machine learning techniques to analyze travel suggestions generated from three open-source LLMs. Our findings reveal that the performance of race and gender classifiers substantially exceeds random chance, indicating differences in how LLMs engage with varied subgroups. Specifically, outputs align with cultural expectations tied to certain races and genders. To minimize the effect of these stereotypes, we used a stop-word classification strategy, which decreased identifiable differences, with no disrespectful terms found. However, hallucinations related to African American and gender minority groups were noted. In conclusion, while LLMs can generate travel plans seemingly free from bias, it remains essential to verify the accuracy and appropriateness of their recommendations.
AdaptDiff: Cross-Modality Domain Adaptation via Weak Conditional Semantic Diffusion for Retinal Vessel Segmentation
Oct 06, 2024Abstract:Deep learning has shown remarkable performance in medical image segmentation. However, despite its promise, deep learning has many challenges in practice due to its inability to effectively transition to unseen domains, caused by the inherent data distribution shift and the lack of manual annotations to guide domain adaptation. To tackle this problem, we present an unsupervised domain adaptation (UDA) method named AdaptDiff that enables a retinal vessel segmentation network trained on fundus photography (FP) to produce satisfactory results on unseen modalities (e.g., OCT-A) without any manual labels. For all our target domains, we first adopt a segmentation model trained on the source domain to create pseudo-labels. With these pseudo-labels, we train a conditional semantic diffusion probabilistic model to represent the target domain distribution. Experimentally, we show that even with low quality pseudo-labels, the diffusion model can still capture the conditional semantic information. Subsequently, we sample on the target domain with binary vessel masks from the source domain to get paired data, i.e., target domain synthetic images conditioned on the binary vessel map. Finally, we fine-tune the pre-trained segmentation network using the synthetic paired data to mitigate the domain gap. We assess the effectiveness of AdaptDiff on seven publicly available datasets across three distinct modalities. Our results demonstrate a significant improvement in segmentation performance across all unseen datasets. Our code is publicly available at https://github.com/DeweiHu/AdaptDiff.
PRISM Lite: A lightweight model for interactive 3D placenta segmentation in ultrasound
Aug 09, 2024Abstract:Placenta volume measured from 3D ultrasound (3DUS) images is an important tool for tracking the growth trajectory and is associated with pregnancy outcomes. Manual segmentation is the gold standard, but it is time-consuming and subjective. Although fully automated deep learning algorithms perform well, they do not always yield high-quality results for each case. Interactive segmentation models could address this issue. However, there is limited work on interactive segmentation models for the placenta. Despite their segmentation accuracy, these methods may not be feasible for clinical use as they require relatively large computational power which may be especially prohibitive in low-resource environments, or on mobile devices. In this paper, we propose a lightweight interactive segmentation model aiming for clinical use to interactively segment the placenta from 3DUS images in real-time. The proposed model adopts the segmentation from our fully automated model for initialization and is designed in a human-in-the-loop manner to achieve iterative improvements. The Dice score and normalized surface Dice are used as evaluation metrics. The results show that our model can achieve superior performance in segmentation compared to state-of-the-art models while using significantly fewer parameters. Additionally, the proposed model is much faster for inference and robust to poor initial masks. The code is available at https://github.com/MedICL-VU/PRISM-placenta.
Retinal IPA: Iterative KeyPoints Alignment for Multimodal Retinal Imaging
Jul 25, 2024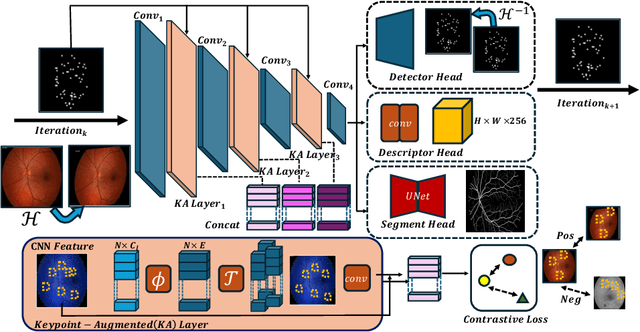
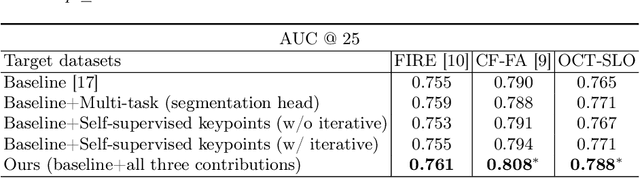
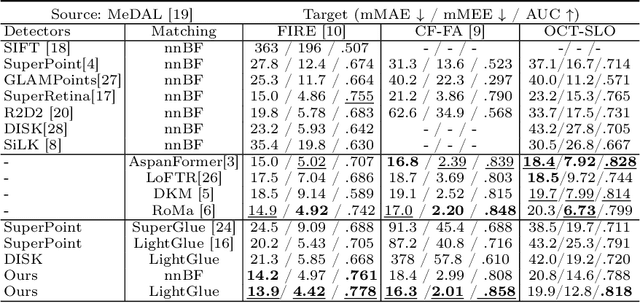
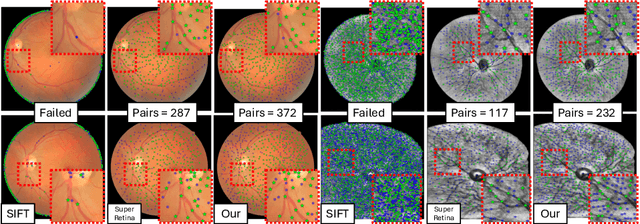

Abstract:We propose a novel framework for retinal feature point alignment, designed for learning cross-modality features to enhance matching and registration across multi-modality retinal images. Our model draws on the success of previous learning-based feature detection and description methods. To better leverage unlabeled data and constrain the model to reproduce relevant keypoints, we integrate a keypoint-based segmentation task. It is trained in a self-supervised manner by enforcing segmentation consistency between different augmentations of the same image. By incorporating a keypoint augmented self-supervised layer, we achieve robust feature extraction across modalities. Extensive evaluation on two public datasets and one in-house dataset demonstrates significant improvements in performance for modality-agnostic retinal feature alignment. Our code and model weights are publicly available at \url{https://github.com/MedICL-VU/RetinaIPA}.
Interactive Segmentation Model for Placenta Segmentation from 3D Ultrasound images
Jul 10, 2024



Abstract:Placenta volume measurement from 3D ultrasound images is critical for predicting pregnancy outcomes, and manual annotation is the gold standard. However, such manual annotation is expensive and time-consuming. Automated segmentation algorithms can often successfully segment the placenta, but these methods may not consistently produce robust segmentations suitable for practical use. Recently, inspired by the Segment Anything Model (SAM), deep learning-based interactive segmentation models have been widely applied in the medical imaging domain. These models produce a segmentation from visual prompts provided to indicate the target region, which may offer a feasible solution for practical use. However, none of these models are specifically designed for interactively segmenting 3D ultrasound images, which remain challenging due to the inherent noise of this modality. In this paper, we evaluate publicly available state-of-the-art 3D interactive segmentation models in contrast to a human-in-the-loop approach for the placenta segmentation task. The Dice score, normalized surface Dice, averaged symmetric surface distance, and 95-percent Hausdorff distance are used as evaluation metrics. We consider a Dice score of 0.95 a successful segmentation. Our results indicate that the human-in-the-loop segmentation model reaches this standard. Moreover, we assess the efficiency of the human-in-the-loop model as a function of the amount of prompts. Our results demonstrate that the human-in-the-loop model is both effective and efficient for interactive placenta segmentation. The code is available at \url{https://github.com/MedICL-VU/PRISM-placenta}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge