Dewei Hu
IntraStyler: Exemplar-based Style Synthesis for Cross-modality Domain Adaptation
Jan 01, 2026Abstract:Image-level domain alignment is the de facto approach for unsupervised domain adaptation, where unpaired image translation is used to minimize the domain gap. Prior studies mainly focus on the domain shift between the source and target domains, whereas the intra-domain variability remains under-explored. To address the latter, an effective strategy is to diversify the styles of the synthetic target domain data during image translation. However, previous methods typically require intra-domain variations to be pre-specified for style synthesis, which may be impractical. In this paper, we propose an exemplar-based style synthesis method named IntraStyler, which can capture diverse intra-domain styles without any prior knowledge. Specifically, IntraStyler uses an exemplar image to guide the style synthesis such that the output style matches the exemplar style. To extract the style-only features, we introduce a style encoder to learn styles discriminatively based on contrastive learning. We evaluate the proposed method on the largest public dataset for cross-modality domain adaptation, CrossMoDA 2023. Our experiments show the efficacy of our method in controllable style synthesis and the benefits of diverse synthetic data for downstream segmentation. Code is available at https://github.com/han-liu/IntraStyler.
SynStitch: a Self-Supervised Learning Network for Ultrasound Image Stitching Using Synthetic Training Pairs and Indirect Supervision
Nov 11, 2024
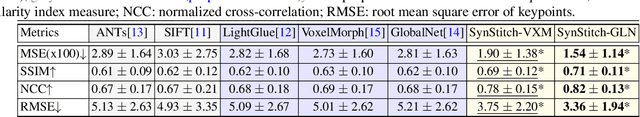
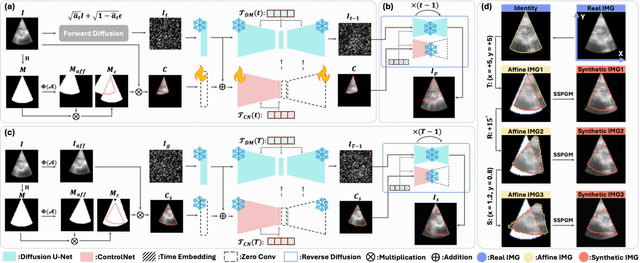

Abstract:Ultrasound (US) image stitching can expand the field-of-view (FOV) by combining multiple US images from varied probe positions. However, registering US images with only partially overlapping anatomical contents is a challenging task. In this work, we introduce SynStitch, a self-supervised framework designed for 2DUS stitching. SynStitch consists of a synthetic stitching pair generation module (SSPGM) and an image stitching module (ISM). SSPGM utilizes a patch-conditioned ControlNet to generate realistic 2DUS stitching pairs with known affine matrix from a single input image. ISM then utilizes this synthetic paired data to learn 2DUS stitching in a supervised manner. Our framework was evaluated against multiple leading methods on a kidney ultrasound dataset, demonstrating superior 2DUS stitching performance through both qualitative and quantitative analyses. The code will be made public upon acceptance of the paper.
AdaptDiff: Cross-Modality Domain Adaptation via Weak Conditional Semantic Diffusion for Retinal Vessel Segmentation
Oct 06, 2024Abstract:Deep learning has shown remarkable performance in medical image segmentation. However, despite its promise, deep learning has many challenges in practice due to its inability to effectively transition to unseen domains, caused by the inherent data distribution shift and the lack of manual annotations to guide domain adaptation. To tackle this problem, we present an unsupervised domain adaptation (UDA) method named AdaptDiff that enables a retinal vessel segmentation network trained on fundus photography (FP) to produce satisfactory results on unseen modalities (e.g., OCT-A) without any manual labels. For all our target domains, we first adopt a segmentation model trained on the source domain to create pseudo-labels. With these pseudo-labels, we train a conditional semantic diffusion probabilistic model to represent the target domain distribution. Experimentally, we show that even with low quality pseudo-labels, the diffusion model can still capture the conditional semantic information. Subsequently, we sample on the target domain with binary vessel masks from the source domain to get paired data, i.e., target domain synthetic images conditioned on the binary vessel map. Finally, we fine-tune the pre-trained segmentation network using the synthetic paired data to mitigate the domain gap. We assess the effectiveness of AdaptDiff on seven publicly available datasets across three distinct modalities. Our results demonstrate a significant improvement in segmentation performance across all unseen datasets. Our code is publicly available at https://github.com/DeweiHu/AdaptDiff.
Retinal IPA: Iterative KeyPoints Alignment for Multimodal Retinal Imaging
Jul 25, 2024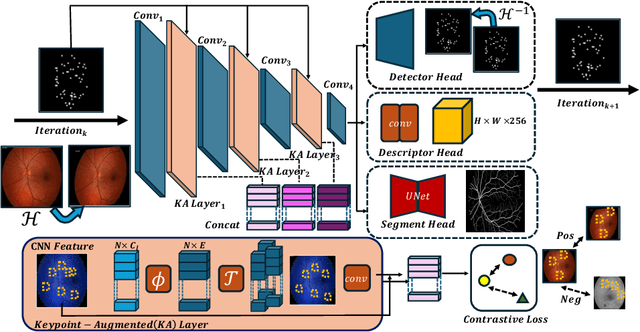
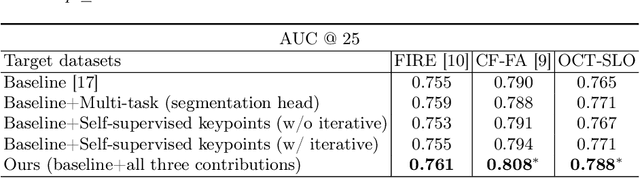
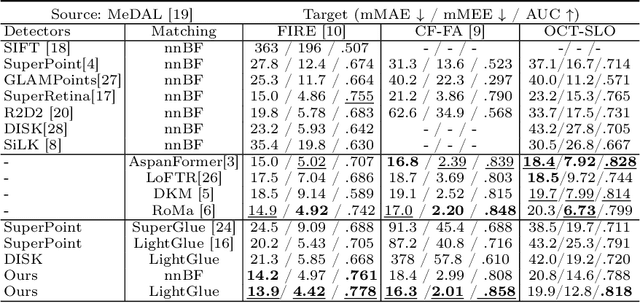
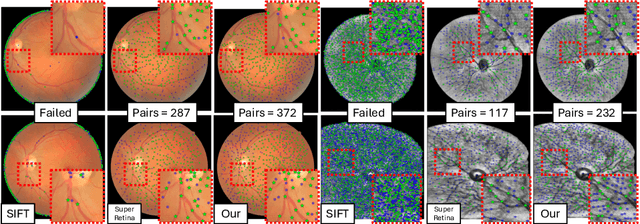

Abstract:We propose a novel framework for retinal feature point alignment, designed for learning cross-modality features to enhance matching and registration across multi-modality retinal images. Our model draws on the success of previous learning-based feature detection and description methods. To better leverage unlabeled data and constrain the model to reproduce relevant keypoints, we integrate a keypoint-based segmentation task. It is trained in a self-supervised manner by enforcing segmentation consistency between different augmentations of the same image. By incorporating a keypoint augmented self-supervised layer, we achieve robust feature extraction across modalities. Extensive evaluation on two public datasets and one in-house dataset demonstrates significant improvements in performance for modality-agnostic retinal feature alignment. Our code and model weights are publicly available at \url{https://github.com/MedICL-VU/RetinaIPA}.
PRISM: A Promptable and Robust Interactive Segmentation Model with Visual Prompts
Apr 23, 2024

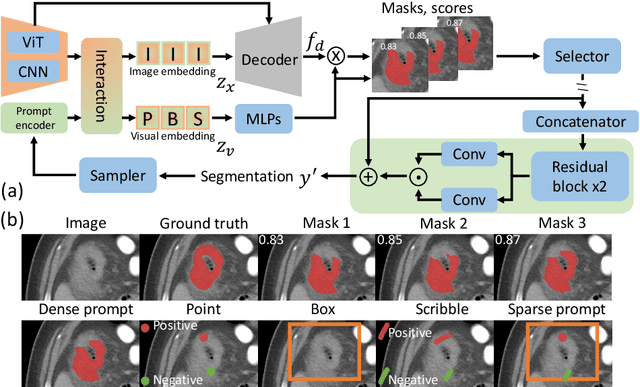

Abstract:In this paper, we present PRISM, a Promptable and Robust Interactive Segmentation Model, aiming for precise segmentation of 3D medical images. PRISM accepts various visual inputs, including points, boxes, and scribbles as sparse prompts, as well as masks as dense prompts. Specifically, PRISM is designed with four principles to achieve robustness: (1) Iterative learning. The model produces segmentations by using visual prompts from previous iterations to achieve progressive improvement. (2) Confidence learning. PRISM employs multiple segmentation heads per input image, each generating a continuous map and a confidence score to optimize predictions. (3) Corrective learning. Following each segmentation iteration, PRISM employs a shallow corrective refinement network to reassign mislabeled voxels. (4) Hybrid design. PRISM integrates hybrid encoders to better capture both the local and global information. Comprehensive validation of PRISM is conducted using four public datasets for tumor segmentation in the colon, pancreas, liver, and kidney, highlighting challenges caused by anatomical variations and ambiguous boundaries in accurate tumor identification. Compared to state-of-the-art methods, both with and without prompt engineering, PRISM significantly improves performance, achieving results that are close to human levels. The code is publicly available at https://github.com/MedICL-VU/PRISM.
Novel OCT mosaicking pipeline with Feature- and Pixel-based registration
Nov 21, 2023Abstract:High-resolution Optical Coherence Tomography (OCT) images are crucial for ophthalmology studies but are limited by their relatively narrow field of view (FoV). Image mosaicking is a technique for aligning multiple overlapping images to obtain a larger FoV. Current mosaicking pipelines often struggle with substantial noise and considerable displacement between the input sub-fields. In this paper, we propose a versatile pipeline for stitching multi-view OCT/OCTA \textit{en face} projection images. Our method combines the strengths of learning-based feature matching and robust pixel-based registration to align multiple images effectively. Furthermore, we advance the application of a trained foundational model, Segment Anything Model (SAM), to validate mosaicking results in an unsupervised manner. The efficacy of our pipeline is validated using an in-house dataset and a large public dataset, where our method shows superior performance in terms of both accuracy and computational efficiency. We also made our evaluation tool for image mosaicking and the corresponding pipeline publicly available at \url{https://github.com/MedICL-VU/OCT-mosaicking}.
Assessing Test-time Variability for Interactive 3D Medical Image Segmentation with Diverse Point Prompts
Nov 13, 2023Abstract:Interactive segmentation model leverages prompts from users to produce robust segmentation. This advancement is facilitated by prompt engineering, where interactive prompts serve as strong priors during test-time. However, this is an inherently subjective and hard-to-reproduce process. The variability in user expertise and inherently ambiguous boundaries in medical images can lead to inconsistent prompt selections, potentially affecting segmentation accuracy. This issue has not yet been extensively explored for medical imaging. In this paper, we assess the test-time variability for interactive medical image segmentation with diverse point prompts. For a given target region, the point is classified into three sub-regions: boundary, margin, and center. Our goal is to identify a straightforward and efficient approach for optimal prompt selection during test-time based on three considerations: (1) benefits of additional prompts, (2) effects of prompt placement, and (3) strategies for optimal prompt selection. We conduct extensive experiments on the public Medical Segmentation Decathlon dataset for challenging colon tumor segmentation task. We suggest an optimal strategy for prompt selection during test-time, supported by comprehensive results. The code is publicly available at https://github.com/MedICL-VU/variability
Promise:Prompt-driven 3D Medical Image Segmentation Using Pretrained Image Foundation Models
Nov 13, 2023Abstract:To address prevalent issues in medical imaging, such as data acquisition challenges and label availability, transfer learning from natural to medical image domains serves as a viable strategy to produce reliable segmentation results. However, several existing barriers between domains need to be broken down, including addressing contrast discrepancies, managing anatomical variability, and adapting 2D pretrained models for 3D segmentation tasks. In this paper, we propose ProMISe,a prompt-driven 3D medical image segmentation model using only a single point prompt to leverage knowledge from a pretrained 2D image foundation model. In particular, we use the pretrained vision transformer from the Segment Anything Model (SAM) and integrate lightweight adapters to extract depth-related (3D) spatial context without updating the pretrained weights. For robust results, a hybrid network with complementary encoders is designed, and a boundary-aware loss is proposed to achieve precise boundaries. We evaluate our model on two public datasets for colon and pancreas tumor segmentations, respectively. Compared to the state-of-the-art segmentation methods with and without prompt engineering, our proposed method achieves superior performance. The code is publicly available at https://github.com/MedICL-VU/ProMISe.
MAP: Domain Generalization via Meta-Learning on Anatomy-Consistent Pseudo-Modalities
Sep 03, 2023Abstract:Deep models suffer from limited generalization capability to unseen domains, which has severely hindered their clinical applicability. Specifically for the retinal vessel segmentation task, although the model is supposed to learn the anatomy of the target, it can be distracted by confounding factors like intensity and contrast. We propose Meta learning on Anatomy-consistent Pseudo-modalities (MAP), a method that improves model generalizability by learning structural features. We first leverage a feature extraction network to generate three distinct pseudo-modalities that share the vessel structure of the original image. Next, we use the episodic learning paradigm by selecting one of the pseudo-modalities as the meta-train dataset, and perform meta-testing on a continuous augmented image space generated through Dirichlet mixup of the remaining pseudo-modalities. Further, we introduce two loss functions that facilitate the model's focus on shape information by clustering the latent vectors obtained from images featuring identical vasculature. We evaluate our model on seven public datasets of various retinal imaging modalities and we conclude that MAP has substantially better generalizability. Our code is publically available at https://github.com/DeweiHu/MAP.
False Negative/Positive Control for SAM on Noisy Medical Images
Aug 20, 2023Abstract:The Segment Anything Model (SAM) is a recently developed all-range foundation model for image segmentation. It can use sparse manual prompts such as bounding boxes to generate pixel-level segmentation in natural images but struggles in medical images such as low-contrast, noisy ultrasound images. We propose a refined test-phase prompt augmentation technique designed to improve SAM's performance in medical image segmentation. The method couples multi-box prompt augmentation and an aleatoric uncertainty-based false-negative (FN) and false-positive (FP) correction (FNPC) strategy. We evaluate the method on two ultrasound datasets and show improvement in SAM's performance and robustness to inaccurate prompts, without the necessity for further training or tuning. Moreover, we present the Single-Slice-to-Volume (SS2V) method, enabling 3D pixel-level segmentation using only the bounding box annotation from a single 2D slice. Our results allow efficient use of SAM in even noisy, low-contrast medical images. The source code will be released soon.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge