Gabriel Arenas
SynStitch: a Self-Supervised Learning Network for Ultrasound Image Stitching Using Synthetic Training Pairs and Indirect Supervision
Nov 11, 2024
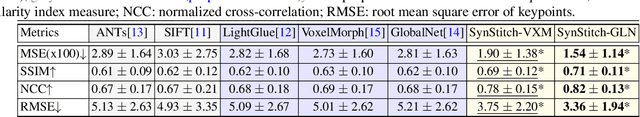
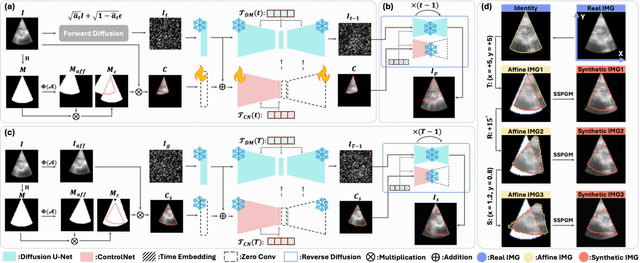

Abstract:Ultrasound (US) image stitching can expand the field-of-view (FOV) by combining multiple US images from varied probe positions. However, registering US images with only partially overlapping anatomical contents is a challenging task. In this work, we introduce SynStitch, a self-supervised framework designed for 2DUS stitching. SynStitch consists of a synthetic stitching pair generation module (SSPGM) and an image stitching module (ISM). SSPGM utilizes a patch-conditioned ControlNet to generate realistic 2DUS stitching pairs with known affine matrix from a single input image. ISM then utilizes this synthetic paired data to learn 2DUS stitching in a supervised manner. Our framework was evaluated against multiple leading methods on a kidney ultrasound dataset, demonstrating superior 2DUS stitching performance through both qualitative and quantitative analyses. The code will be made public upon acceptance of the paper.
PRISM Lite: A lightweight model for interactive 3D placenta segmentation in ultrasound
Aug 09, 2024Abstract:Placenta volume measured from 3D ultrasound (3DUS) images is an important tool for tracking the growth trajectory and is associated with pregnancy outcomes. Manual segmentation is the gold standard, but it is time-consuming and subjective. Although fully automated deep learning algorithms perform well, they do not always yield high-quality results for each case. Interactive segmentation models could address this issue. However, there is limited work on interactive segmentation models for the placenta. Despite their segmentation accuracy, these methods may not be feasible for clinical use as they require relatively large computational power which may be especially prohibitive in low-resource environments, or on mobile devices. In this paper, we propose a lightweight interactive segmentation model aiming for clinical use to interactively segment the placenta from 3DUS images in real-time. The proposed model adopts the segmentation from our fully automated model for initialization and is designed in a human-in-the-loop manner to achieve iterative improvements. The Dice score and normalized surface Dice are used as evaluation metrics. The results show that our model can achieve superior performance in segmentation compared to state-of-the-art models while using significantly fewer parameters. Additionally, the proposed model is much faster for inference and robust to poor initial masks. The code is available at https://github.com/MedICL-VU/PRISM-placenta.
Interactive Segmentation Model for Placenta Segmentation from 3D Ultrasound images
Jul 10, 2024



Abstract:Placenta volume measurement from 3D ultrasound images is critical for predicting pregnancy outcomes, and manual annotation is the gold standard. However, such manual annotation is expensive and time-consuming. Automated segmentation algorithms can often successfully segment the placenta, but these methods may not consistently produce robust segmentations suitable for practical use. Recently, inspired by the Segment Anything Model (SAM), deep learning-based interactive segmentation models have been widely applied in the medical imaging domain. These models produce a segmentation from visual prompts provided to indicate the target region, which may offer a feasible solution for practical use. However, none of these models are specifically designed for interactively segmenting 3D ultrasound images, which remain challenging due to the inherent noise of this modality. In this paper, we evaluate publicly available state-of-the-art 3D interactive segmentation models in contrast to a human-in-the-loop approach for the placenta segmentation task. The Dice score, normalized surface Dice, averaged symmetric surface distance, and 95-percent Hausdorff distance are used as evaluation metrics. We consider a Dice score of 0.95 a successful segmentation. Our results indicate that the human-in-the-loop segmentation model reaches this standard. Moreover, we assess the efficiency of the human-in-the-loop model as a function of the amount of prompts. Our results demonstrate that the human-in-the-loop model is both effective and efficient for interactive placenta segmentation. The code is available at \url{https://github.com/MedICL-VU/PRISM-placenta}.
False Negative/Positive Control for SAM on Noisy Medical Images
Aug 20, 2023Abstract:The Segment Anything Model (SAM) is a recently developed all-range foundation model for image segmentation. It can use sparse manual prompts such as bounding boxes to generate pixel-level segmentation in natural images but struggles in medical images such as low-contrast, noisy ultrasound images. We propose a refined test-phase prompt augmentation technique designed to improve SAM's performance in medical image segmentation. The method couples multi-box prompt augmentation and an aleatoric uncertainty-based false-negative (FN) and false-positive (FP) correction (FNPC) strategy. We evaluate the method on two ultrasound datasets and show improvement in SAM's performance and robustness to inaccurate prompts, without the necessity for further training or tuning. Moreover, we present the Single-Slice-to-Volume (SS2V) method, enabling 3D pixel-level segmentation using only the bounding box annotation from a single 2D slice. Our results allow efficient use of SAM in even noisy, low-contrast medical images. The source code will be released soon.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge