Lining Yu
KPIs 2024 Challenge: Advancing Glomerular Segmentation from Patch- to Slide-Level
Feb 11, 2025
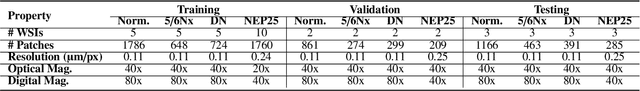
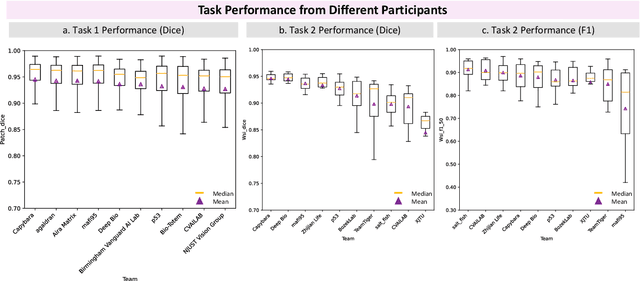

Abstract:Chronic kidney disease (CKD) is a major global health issue, affecting over 10% of the population and causing significant mortality. While kidney biopsy remains the gold standard for CKD diagnosis and treatment, the lack of comprehensive benchmarks for kidney pathology segmentation hinders progress in the field. To address this, we organized the Kidney Pathology Image Segmentation (KPIs) Challenge, introducing a dataset that incorporates preclinical rodent models of CKD with over 10,000 annotated glomeruli from 60+ Periodic Acid Schiff (PAS)-stained whole slide images. The challenge includes two tasks, patch-level segmentation and whole slide image segmentation and detection, evaluated using the Dice Similarity Coefficient (DSC) and F1-score. By encouraging innovative segmentation methods that adapt to diverse CKD models and tissue conditions, the KPIs Challenge aims to advance kidney pathology analysis, establish new benchmarks, and enable precise, large-scale quantification for disease research and diagnosis.
Towards Fine-grained Renal Vasculature Segmentation: Full-Scale Hierarchical Learning with FH-Seg
Feb 07, 2025Abstract:Accurate fine-grained segmentation of the renal vasculature is critical for nephrological analysis, yet it faces challenges due to diverse and insufficiently annotated images. Existing methods struggle to accurately segment intricate regions of the renal vasculature, such as the inner and outer walls, arteries and lesions. In this paper, we introduce FH-Seg, a Full-scale Hierarchical Learning Framework designed for comprehensive segmentation of the renal vasculature. Specifically, FH-Seg employs full-scale skip connections that merge detailed anatomical information with contextual semantics across scales, effectively bridging the gap between structural and pathological contexts. Additionally, we implement a learnable hierarchical soft attention gates to adaptively reduce interference from non-core information, enhancing the focus on critical vascular features. To advance research on renal pathology segmentation, we also developed a Large Renal Vasculature (LRV) dataset, which contains 16,212 fine-grained annotated images of 5,600 renal arteries. Extensive experiments on the LRV dataset demonstrate FH-Seg's superior accuracies (71.23% Dice, 73.06% F1), outperforming Omni-Seg by 2.67 and 2.13 percentage points respectively. Code is available at: https://github.com/hrlblab/FH-seg.
Glo-In-One-v2: Holistic Identification of Glomerular Cells, Tissues, and Lesions in Human and Mouse Histopathology
Nov 25, 2024Abstract:Segmenting glomerular intraglomerular tissue and lesions traditionally depends on detailed morphological evaluations by expert nephropathologists, a labor-intensive process susceptible to interobserver variability. Our group previously developed the Glo-In-One toolkit for integrated detection and segmentation of glomeruli. In this study, we leverage the Glo-In-One toolkit to version 2 with fine-grained segmentation capabilities, curating 14 distinct labels for tissue regions, cells, and lesions across a dataset of 23,529 annotated glomeruli across human and mouse histopathology data. To our knowledge, this dataset is among the largest of its kind to date.In this study, we present a single dynamic head deep learning architecture designed to segment 14 classes within partially labeled images of human and mouse pathology data. Our model was trained using a training set derived from 368 annotated kidney whole-slide images (WSIs) to identify 5 key intraglomerular tissues covering Bowman's capsule, glomerular tuft, mesangium, mesangial cells, and podocytes. Additionally, the network segments 9 glomerular lesion classes including adhesion, capsular drop, global sclerosis, hyalinosis, mesangial lysis, microaneurysm, nodular sclerosis, mesangial expansion, and segmental sclerosis. The glomerulus segmentation model achieved a decent performance compared with baselines, and achieved a 76.5 % average Dice Similarity Coefficient (DSC). Additional, transfer learning from rodent to human for glomerular lesion segmentation model has enhanced the average segmentation accuracy across different types of lesions by more than 3 %, as measured by Dice scores. The Glo-In-One-v2 model and trained weight have been made publicly available at https: //github.com/hrlblab/Glo-In-One_v2.
Adapting Mouse Pathological Model to Human Glomerular Lesion Segmentation
Jul 25, 2024Abstract:Moving from animal models to human applications in preclinical research encompasses a broad spectrum of disciplines in medical science. A fundamental element in the development of new drugs, treatments, diagnostic methods, and in deepening our understanding of disease processes is the accurate measurement of kidney tissues. Past studies have demonstrated the viability of translating glomeruli segmentation techniques from mouse models to human applications. Yet, these investigations tend to neglect the complexities involved in segmenting pathological glomeruli affected by different lesions. Such lesions present a wider range of morphological variations compared to healthy glomerular tissue, which are arguably more valuable than normal glomeruli in clinical practice. Furthermore, data on lesions from animal models can be more readily scaled up from disease models and whole kidney biopsies. This brings up a question: ``\textit{Can a pathological segmentation model trained on mouse models be effectively applied to human patients?}" To answer this question, we introduced GLAM, a deep learning study for fine-grained segmentation of human kidney lesions using a mouse model, addressing mouse-to-human transfer learning, by evaluating different learning strategies for segmenting human pathological lesions using zero-shot transfer learning and hybrid learning by leveraging mouse samples. From the results, the hybrid learning model achieved superior performance.
PrPSeg: Universal Proposition Learning for Panoramic Renal Pathology Segmentation
Feb 29, 2024



Abstract:Understanding the anatomy of renal pathology is crucial for advancing disease diagnostics, treatment evaluation, and clinical research. The complex kidney system comprises various components across multiple levels, including regions (cortex, medulla), functional units (glomeruli, tubules), and cells (podocytes, mesangial cells in glomerulus). Prior studies have predominantly overlooked the intricate spatial interrelations among objects from clinical knowledge. In this research, we introduce a novel universal proposition learning approach, called panoramic renal pathology segmentation (PrPSeg), designed to segment comprehensively panoramic structures within kidney by integrating extensive knowledge of kidney anatomy. In this paper, we propose (1) the design of a comprehensive universal proposition matrix for renal pathology, facilitating the incorporation of classification and spatial relationships into the segmentation process; (2) a token-based dynamic head single network architecture, with the improvement of the partial label image segmentation and capability for future data enlargement; and (3) an anatomy loss function, quantifying the inter-object relationships across the kidney.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge