Alessa Hering
and on behalf of the UNICORN consortium
Tracking Cancer Through Text: Longitudinal Extraction From Radiology Reports Using Open-Source Large Language Models
Mar 11, 2026Abstract:Radiology reports capture crucial longitudinal information on tumor burden, treatment response, and disease progression, yet their unstructured narrative format complicates automated analysis. While large language models (LLMs) have advanced clinical text processing, most state-of-the-art systems remain proprietary, limiting their applicability in privacy-sensitive healthcare environments. We present a fully open-source, locally deployable pipeline for longitudinal information extraction from radiology reports, implemented using the llm_extractinator framework. The system applies the qwen2.5-72b model to extract and link target, non-target, and new lesion data across time points in accordance with RECIST criteria. Evaluation on 50 Dutch CT Thorax/Abdomen report pairs yielded high extraction performance, with attribute-level accuracies of 93.7% for target lesions, 94.9% for non-target lesions, and 94.0% for new lesions. The approach demonstrates that open-source LLMs can achieve clinically meaningful performance in multi-timepoint oncology tasks while ensuring data privacy and reproducibility. These results highlight the potential of locally deployable LLMs for scalable extraction of structured longitudinal data from routine clinical text.
Designing UNICORN: a Unified Benchmark for Imaging in Computational Pathology, Radiology, and Natural Language
Mar 03, 2026Abstract:Medical foundation models show promise to learn broadly generalizable features from large, diverse datasets. This could be the base for reliable cross-modality generalization and rapid adaptation to new, task-specific goals, with only a few task-specific examples. Yet, evidence for this is limited by the lack of public, standardized, and reproducible evaluation frameworks, as existing public benchmarks are often fragmented across task-, organ-, or modality-specific settings, limiting assessment of cross-task generalization. We introduce UNICORN, a public benchmark designed to systematically evaluate medical foundation models under a unified protocol. To isolate representation quality, we built the benchmark on a novel two-step framework that decouples model inference from task-specific evaluation based on standardized few-shot adaptation. As a central design choice, we constructed indirectly accessible sequestered test sets derived from clinically relevant cohorts, along with standardized evaluation code and a submission interface on an open benchmarking platform. Performance is aggregated into a single UNICORN Score, a new metric that we introduce to support direct comparison of foundation models across diverse medical domains, modalities, and task types. The UNICORN test dataset includes data from more than 2,400 patients, including over 3,700 vision cases and over 2,400 clinical reports collected from 17 institutions across eight countries. The benchmark spans eight anatomical regions and four imaging modalities. Both task-specific and aggregated leaderboards enable accessible, standardized, and reproducible evaluation. By standardizing multi-task, multi-modality assessment, UNICORN establishes a foundation for reproducible benchmarking of medical foundation models. Data, baseline methods, and the evaluation platform are publicly available via unicorn.grand-challenge.org.
Kidney Cancer Detection Using 3D-Based Latent Diffusion Models
Jan 09, 2026Abstract:In this work, we present a novel latent diffusion-based pipeline for 3D kidney anomaly detection on contrast-enhanced abdominal CT. The method combines Denoising Diffusion Probabilistic Models (DDPMs), Denoising Diffusion Implicit Models (DDIMs), and Vector-Quantized Generative Adversarial Networks (VQ-GANs). Unlike prior slice-wise approaches, our method operates directly on an image volume and leverages weak supervision with only case-level pseudo-labels. We benchmark our approach against state-of-the-art supervised segmentation and detection models. This study demonstrates the feasibility and promise of 3D latent diffusion for weakly supervised anomaly detection. While the current results do not yet match supervised baselines, they reveal key directions for improving reconstruction fidelity and lesion localization. Our findings provide an important step toward annotation-efficient, generative modeling of complex abdominal anatomy.
EvalBlocks: A Modular Pipeline for Rapidly Evaluating Foundation Models in Medical Imaging
Jan 07, 2026Abstract:Developing foundation models in medical imaging requires continuous monitoring of downstream performance. Researchers are burdened with tracking numerous experiments, design choices, and their effects on performance, often relying on ad-hoc, manual workflows that are inherently slow and error-prone. We introduce EvalBlocks, a modular, plug-and-play framework for efficient evaluation of foundation models during development. Built on Snakemake, EvalBlocks supports seamless integration of new datasets, foundation models, aggregation methods, and evaluation strategies. All experiments and results are tracked centrally and are reproducible with a single command, while efficient caching and parallel execution enable scalable use on shared compute infrastructure. Demonstrated on five state-of-the-art foundation models and three medical imaging classification tasks, EvalBlocks streamlines model evaluation, enabling researchers to iterate faster and focus on model innovation rather than evaluation logistics. The framework is released as open source software at https://github.com/DIAGNijmegen/eval-blocks.
ULS+: Data-driven Model Adaptation Enhances Lesion Segmentation
Jan 06, 2026Abstract:In this study, we present ULS+, an enhanced version of the Universal Lesion Segmentation (ULS) model. The original ULS model segments lesions across the whole body in CT scans given volumes of interest (VOIs) centered around a click-point. Since its release, several new public datasets have become available that can further improve model performance. ULS+ incorporates these additional datasets and uses smaller input image sizes, resulting in higher accuracy and faster inference. We compared ULS and ULS+ using the Dice score and robustness to click-point location on the ULS23 Challenge test data and a subset of the Longitudinal-CT dataset. In all comparisons, ULS+ significantly outperformed ULS. Additionally, ULS+ ranks first on the ULS23 Challenge test-phase leaderboard. By maintaining a cycle of data-driven updates and clinical validation, ULS+ establishes a foundation for robust and clinically relevant lesion segmentation models.
Fairness Evaluation of Risk Estimation Models for Lung Cancer Screening
Dec 23, 2025Abstract:Lung cancer is the leading cause of cancer-related mortality in adults worldwide. Screening high-risk individuals with annual low-dose CT (LDCT) can support earlier detection and reduce deaths, but widespread implementation may strain the already limited radiology workforce. AI models have shown potential in estimating lung cancer risk from LDCT scans. However, high-risk populations for lung cancer are diverse, and these models' performance across demographic groups remains an open question. In this study, we drew on the considerations on confounding factors and ethically significant biases outlined in the JustEFAB framework to evaluate potential performance disparities and fairness in two deep learning risk estimation models for lung cancer screening: the Sybil lung cancer risk model and the Venkadesh21 nodule risk estimator. We also examined disparities in the PanCan2b logistic regression model recommended in the British Thoracic Society nodule management guideline. Both deep learning models were trained on data from the US-based National Lung Screening Trial (NLST), and assessed on a held-out NLST validation set. We evaluated AUROC, sensitivity, and specificity across demographic subgroups, and explored potential confounding from clinical risk factors. We observed a statistically significant AUROC difference in Sybil's performance between women (0.88, 95% CI: 0.86, 0.90) and men (0.81, 95% CI: 0.78, 0.84, p < .001). At 90% specificity, Venkadesh21 showed lower sensitivity for Black (0.39, 95% CI: 0.23, 0.59) than White participants (0.69, 95% CI: 0.65, 0.73). These differences were not explained by available clinical confounders and thus may be classified as unfair biases according to JustEFAB. Our findings highlight the importance of improving and monitoring model performance across underrepresented subgroups, and further research on algorithmic fairness, in lung cancer screening.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2025:025
Beyond the LUMIR challenge: The pathway to foundational registration models
May 30, 2025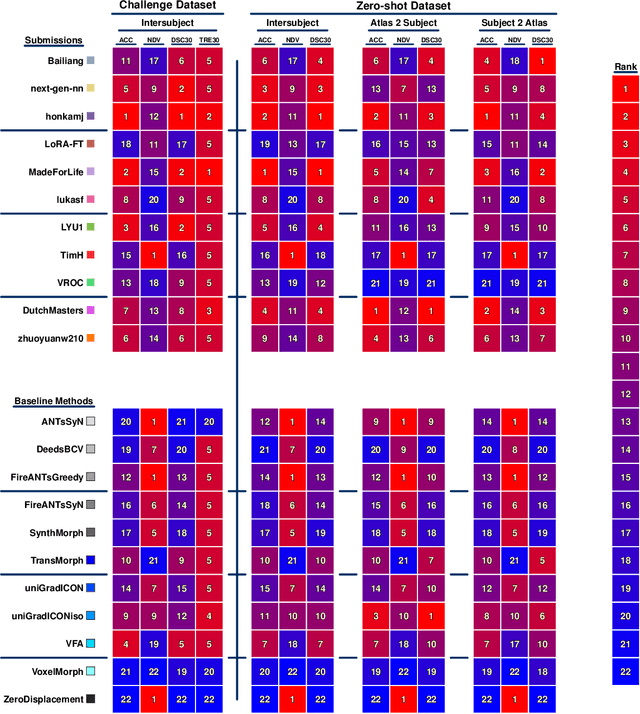
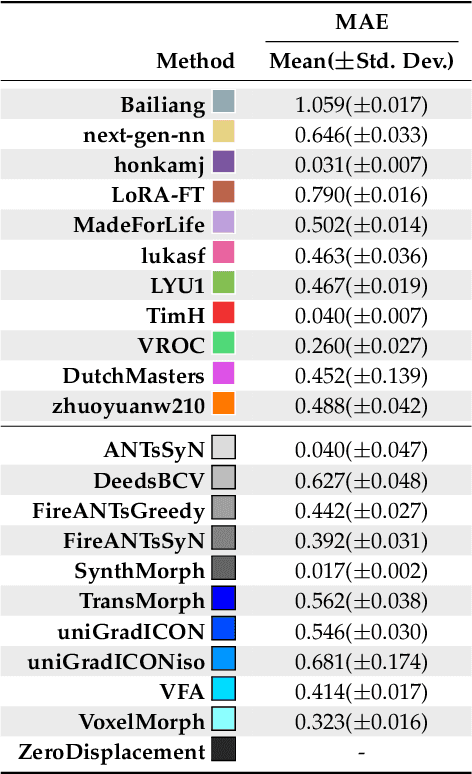
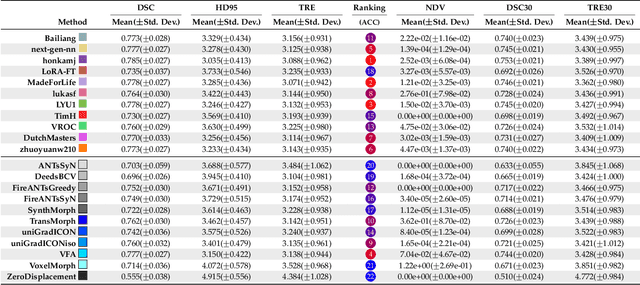

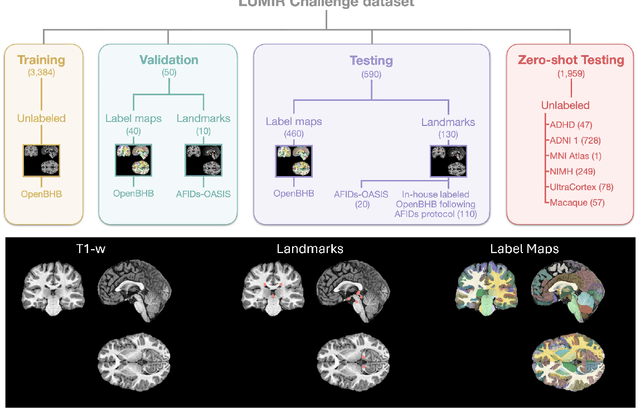
Abstract:Medical image challenges have played a transformative role in advancing the field, catalyzing algorithmic innovation and establishing new performance standards across diverse clinical applications. Image registration, a foundational task in neuroimaging pipelines, has similarly benefited from the Learn2Reg initiative. Building on this foundation, we introduce the Large-scale Unsupervised Brain MRI Image Registration (LUMIR) challenge, a next-generation benchmark designed to assess and advance unsupervised brain MRI registration. Distinct from prior challenges that leveraged anatomical label maps for supervision, LUMIR removes this dependency by providing over 4,000 preprocessed T1-weighted brain MRIs for training without any label maps, encouraging biologically plausible deformation modeling through self-supervision. In addition to evaluating performance on 590 held-out test subjects, LUMIR introduces a rigorous suite of zero-shot generalization tasks, spanning out-of-domain imaging modalities (e.g., FLAIR, T2-weighted, T2*-weighted), disease populations (e.g., Alzheimer's disease), acquisition protocols (e.g., 9.4T MRI), and species (e.g., macaque brains). A total of 1,158 subjects and over 4,000 image pairs were included for evaluation. Performance was assessed using both segmentation-based metrics (Dice coefficient, 95th percentile Hausdorff distance) and landmark-based registration accuracy (target registration error). Across both in-domain and zero-shot tasks, deep learning-based methods consistently achieved state-of-the-art accuracy while producing anatomically plausible deformation fields. The top-performing deep learning-based models demonstrated diffeomorphic properties and inverse consistency, outperforming several leading optimization-based methods, and showing strong robustness to most domain shifts, the exception being a drop in performance on out-of-domain contrasts.
Robust Kidney Abnormality Segmentation: A Validation Study of an AI-Based Framework
May 12, 2025Abstract:Kidney abnormality segmentation has important potential to enhance the clinical workflow, especially in settings requiring quantitative assessments. Kidney volume could serve as an important biomarker for renal diseases, with changes in volume correlating directly with kidney function. Currently, clinical practice often relies on subjective visual assessment for evaluating kidney size and abnormalities, including tumors and cysts, which are typically staged based on diameter, volume, and anatomical location. To support a more objective and reproducible approach, this research aims to develop a robust, thoroughly validated kidney abnormality segmentation algorithm, made publicly available for clinical and research use. We employ publicly available training datasets and leverage the state-of-the-art medical image segmentation framework nnU-Net. Validation is conducted using both proprietary and public test datasets, with segmentation performance quantified by Dice coefficient and the 95th percentile Hausdorff distance. Furthermore, we analyze robustness across subgroups based on patient sex, age, CT contrast phases, and tumor histologic subtypes. Our findings demonstrate that our segmentation algorithm, trained exclusively on publicly available data, generalizes effectively to external test sets and outperforms existing state-of-the-art models across all tested datasets. Subgroup analyses reveal consistent high performance, indicating strong robustness and reliability. The developed algorithm and associated code are publicly accessible at https://github.com/DIAGNijmegen/oncology-kidney-abnormality-segmentation.
OncoReg: Medical Image Registration for Oncological Challenges
Apr 01, 2025Abstract:In modern cancer research, the vast volume of medical data generated is often underutilised due to challenges related to patient privacy. The OncoReg Challenge addresses this issue by enabling researchers to develop and validate image registration methods through a two-phase framework that ensures patient privacy while fostering the development of more generalisable AI models. Phase one involves working with a publicly available dataset, while phase two focuses on training models on a private dataset within secure hospital networks. OncoReg builds upon the foundation established by the Learn2Reg Challenge by incorporating the registration of interventional cone-beam computed tomography (CBCT) with standard planning fan-beam CT (FBCT) images in radiotherapy. Accurate image registration is crucial in oncology, particularly for dynamic treatment adjustments in image-guided radiotherapy, where precise alignment is necessary to minimise radiation exposure to healthy tissues while effectively targeting tumours. This work details the methodology and data behind the OncoReg Challenge and provides a comprehensive analysis of the competition entries and results. Findings reveal that feature extraction plays a pivotal role in this registration task. A new method emerging from this challenge demonstrated its versatility, while established approaches continue to perform comparably to newer techniques. Both deep learning and classical approaches still play significant roles in image registration, with the combination of methods - particularly in feature extraction - proving most effective.
Divide to Conquer: A Field Decomposition Approach for Multi-Organ Whole-Body CT Image Registration
Mar 28, 2025Abstract:Image registration is an essential technique for the analysis of Computed Tomography (CT) images in clinical practice. However, existing methodologies are predominantly tailored to a specific organ of interest and often exhibit lower performance on other organs, thus limiting their generalizability and applicability. Multi-organ registration addresses these limitations, but the simultaneous alignment of multiple organs with diverse shapes, sizes and locations requires a highly complex deformation field with a multi-layer composition of individual deformations. This study introduces a novel field decomposition approach to address the high complexity of deformations in multi-organ whole-body CT image registration. The proposed method is trained and evaluated on a longitudinal dataset of 691 patients, each with two CT images obtained at distinct time points. These scans fully encompass the thoracic, abdominal, and pelvic regions. Two baseline registration methods are selected for this study: one based on optimization techniques and another based on deep learning. Experimental results demonstrate that the proposed approach outperforms baseline methods in handling complex deformations in multi-organ whole-body CT image registration.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge