Hartmut Häntze
Robust Kidney Abnormality Segmentation: A Validation Study of an AI-Based Framework
May 12, 2025Abstract:Kidney abnormality segmentation has important potential to enhance the clinical workflow, especially in settings requiring quantitative assessments. Kidney volume could serve as an important biomarker for renal diseases, with changes in volume correlating directly with kidney function. Currently, clinical practice often relies on subjective visual assessment for evaluating kidney size and abnormalities, including tumors and cysts, which are typically staged based on diameter, volume, and anatomical location. To support a more objective and reproducible approach, this research aims to develop a robust, thoroughly validated kidney abnormality segmentation algorithm, made publicly available for clinical and research use. We employ publicly available training datasets and leverage the state-of-the-art medical image segmentation framework nnU-Net. Validation is conducted using both proprietary and public test datasets, with segmentation performance quantified by Dice coefficient and the 95th percentile Hausdorff distance. Furthermore, we analyze robustness across subgroups based on patient sex, age, CT contrast phases, and tumor histologic subtypes. Our findings demonstrate that our segmentation algorithm, trained exclusively on publicly available data, generalizes effectively to external test sets and outperforms existing state-of-the-art models across all tested datasets. Subgroup analyses reveal consistent high performance, indicating strong robustness and reliability. The developed algorithm and associated code are publicly accessible at https://github.com/DIAGNijmegen/oncology-kidney-abnormality-segmentation.
Incorporating Anatomical Awareness for Enhanced Generalizability and Progression Prediction in Deep Learning-Based Radiographic Sacroiliitis Detection
May 12, 2024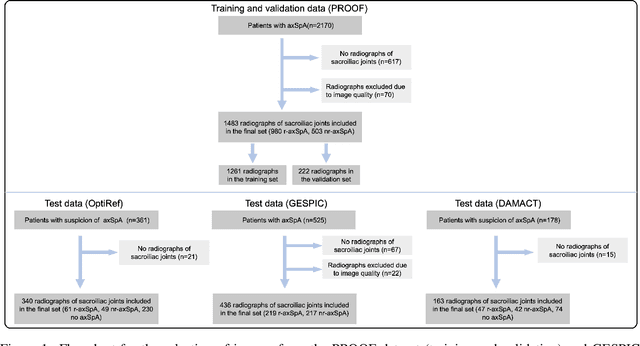
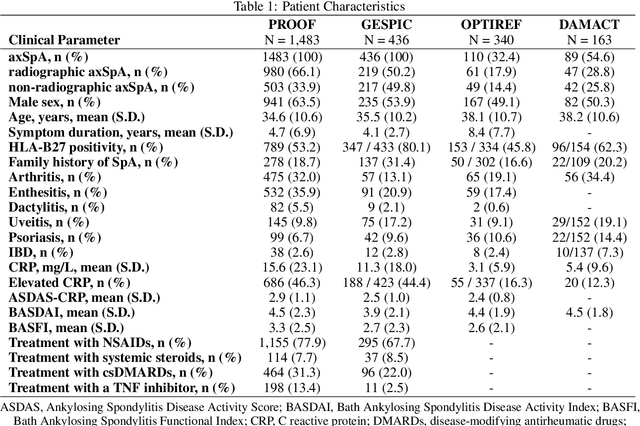
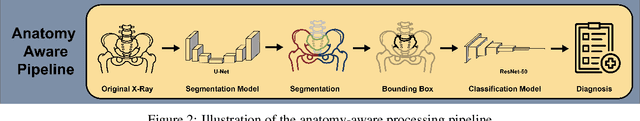

Abstract:Purpose: To examine whether incorporating anatomical awareness into a deep learning model can improve generalizability and enable prediction of disease progression. Methods: This retrospective multicenter study included conventional pelvic radiographs of 4 different patient cohorts focusing on axial spondyloarthritis (axSpA) collected at university and community hospitals. The first cohort, which consisted of 1483 radiographs, was split into training (n=1261) and validation (n=222) sets. The other cohorts comprising 436, 340, and 163 patients, respectively, were used as independent test datasets. For the second cohort, follow-up data of 311 patients was used to examine progression prediction capabilities. Two neural networks were trained, one on images cropped to the bounding box of the sacroiliac joints (anatomy-aware) and the other one on full radiographs. The performance of the models was compared using the area under the receiver operating characteristic curve (AUC), accuracy, sensitivity, and specificity. Results: On the three test datasets, the standard model achieved AUC scores of 0.853, 0.817, 0.947, with an accuracy of 0.770, 0.724, 0.850. Whereas the anatomy-aware model achieved AUC scores of 0.899, 0.846, 0.957, with an accuracy of 0.821, 0.744, 0.906, respectively. The patients who were identified as high risk by the anatomy aware model had an odds ratio of 2.16 (95% CI: 1.19, 3.86) for having progression of radiographic sacroiliitis within 2 years. Conclusion: Anatomical awareness can improve the generalizability of a deep learning model in detecting radiographic sacroiliitis. The model is published as fully open source alongside this study.
MRSegmentator: Robust Multi-Modality Segmentation of 40 Classes in MRI and CT Sequences
May 10, 2024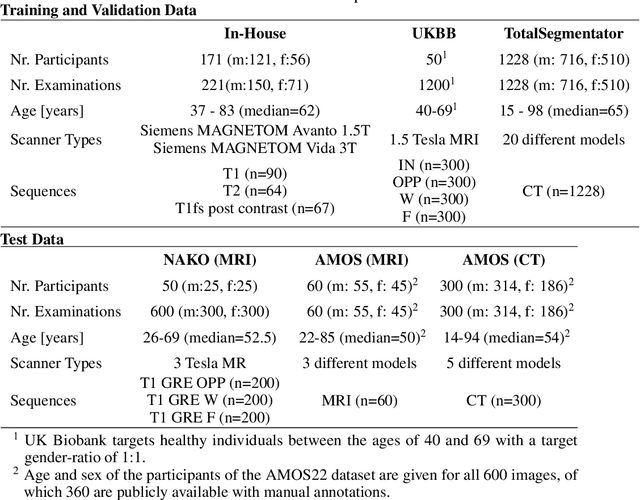
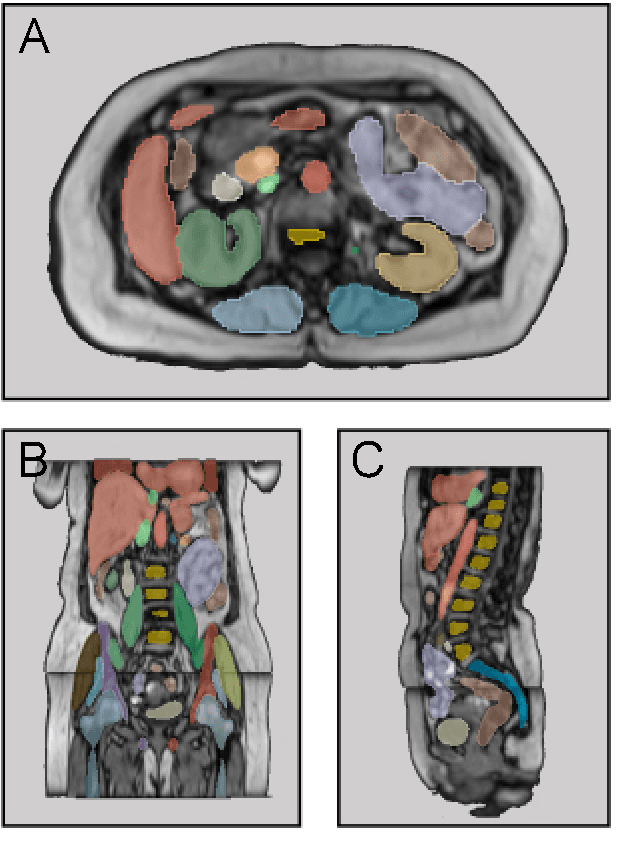
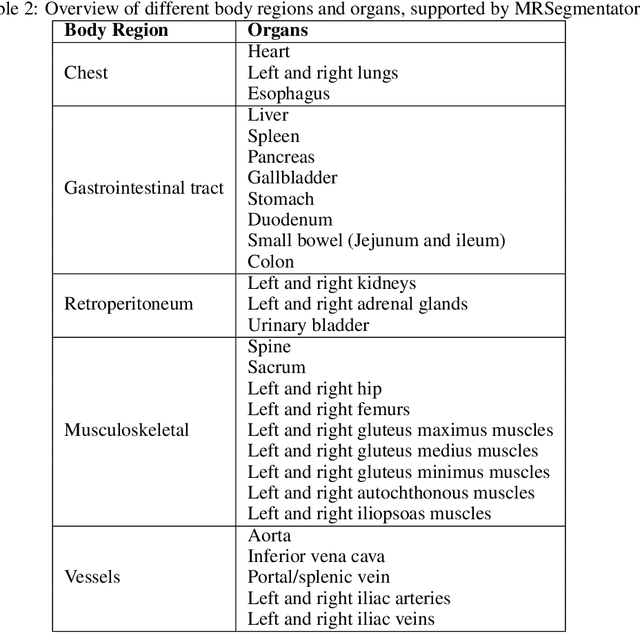
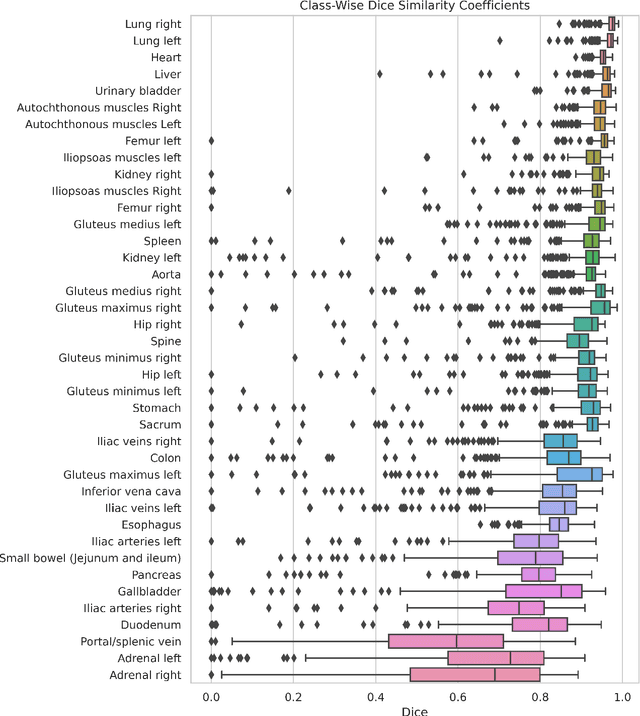
Abstract:Purpose: To introduce a deep learning model capable of multi-organ segmentation in MRI scans, offering a solution to the current limitations in MRI analysis due to challenges in resolution, standardized intensity values, and variability in sequences. Materials and Methods: he model was trained on 1,200 manually annotated MRI scans from the UK Biobank, 221 in-house MRI scans and 1228 CT scans, leveraging cross-modality transfer learning from CT segmentation models. A human-in-the-loop annotation workflow was employed to efficiently create high-quality segmentations. The model's performance was evaluated on NAKO and the AMOS22 dataset containing 600 and 60 MRI examinations. Dice Similarity Coefficient (DSC) and Hausdorff Distance (HD) was used to assess segmentation accuracy. The model will be open sourced. Results: The model showcased high accuracy in segmenting well-defined organs, achieving Dice Similarity Coefficient (DSC) scores of 0.97 for the right and left lungs, and 0.95 for the heart. It also demonstrated robustness in organs like the liver (DSC: 0.96) and kidneys (DSC: 0.95 left, 0.95 right), which present more variability. However, segmentation of smaller and complex structures such as the portal and splenic veins (DSC: 0.54) and adrenal glands (DSC: 0.65 left, 0.61 right) revealed the need for further model optimization. Conclusion: The proposed model is a robust, tool for accurate segmentation of 40 anatomical structures in MRI and CT images. By leveraging cross-modality learning and interactive annotation, the model achieves strong performance and generalizability across diverse datasets, making it a valuable resource for researchers and clinicians. It is open source and can be downloaded from https://github.com/hhaentze/MRSegmentator.
Improve Cross-Modality Segmentation by Treating MRI Images as Inverted CT Scans
May 04, 2024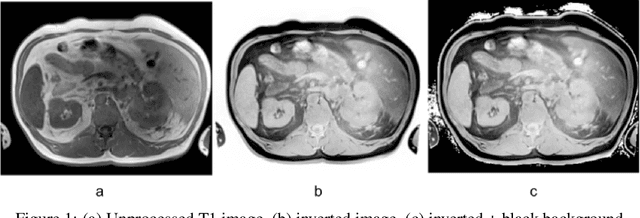
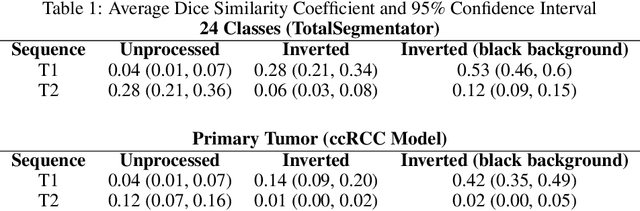

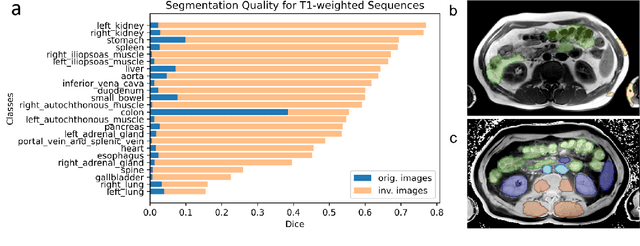
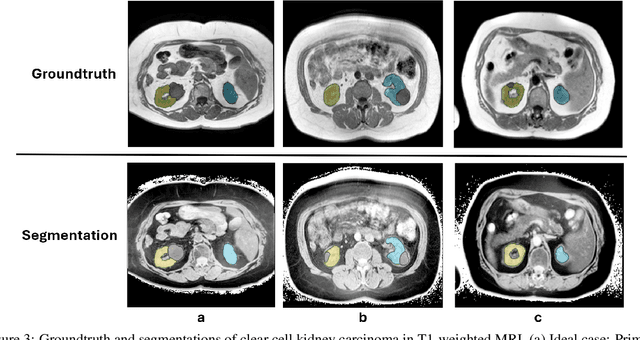
Abstract:Computed tomography (CT) segmentation models frequently include classes that are not currently supported by magnetic resonance imaging (MRI) segmentation models. In this study, we show that a simple image inversion technique can significantly improve the segmentation quality of CT segmentation models on MRI data, by using the TotalSegmentator model, applied to T1-weighted MRI images, as example. Image inversion is straightforward to implement and does not require dedicated graphics processing units (GPUs), thus providing a quick alternative to complex deep modality-transfer models for generating segmentation masks for MRI data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge