Janis L. Vahldiek
Incorporating Anatomical Awareness for Enhanced Generalizability and Progression Prediction in Deep Learning-Based Radiographic Sacroiliitis Detection
May 12, 2024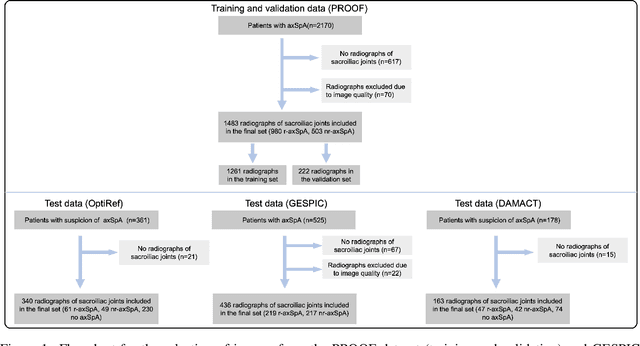
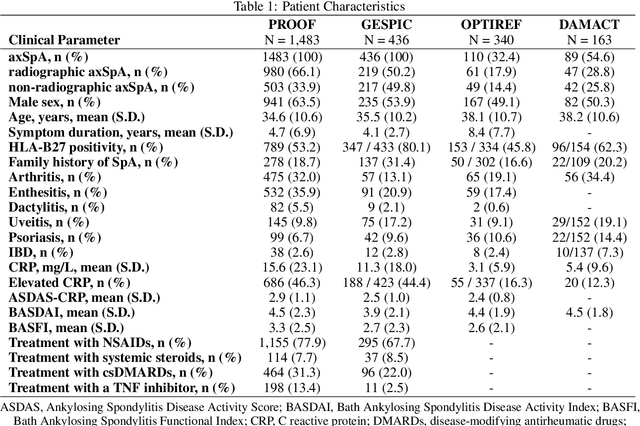
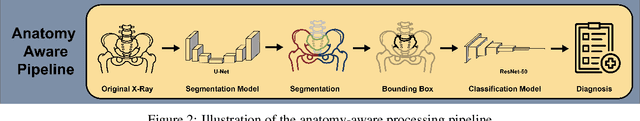

Abstract:Purpose: To examine whether incorporating anatomical awareness into a deep learning model can improve generalizability and enable prediction of disease progression. Methods: This retrospective multicenter study included conventional pelvic radiographs of 4 different patient cohorts focusing on axial spondyloarthritis (axSpA) collected at university and community hospitals. The first cohort, which consisted of 1483 radiographs, was split into training (n=1261) and validation (n=222) sets. The other cohorts comprising 436, 340, and 163 patients, respectively, were used as independent test datasets. For the second cohort, follow-up data of 311 patients was used to examine progression prediction capabilities. Two neural networks were trained, one on images cropped to the bounding box of the sacroiliac joints (anatomy-aware) and the other one on full radiographs. The performance of the models was compared using the area under the receiver operating characteristic curve (AUC), accuracy, sensitivity, and specificity. Results: On the three test datasets, the standard model achieved AUC scores of 0.853, 0.817, 0.947, with an accuracy of 0.770, 0.724, 0.850. Whereas the anatomy-aware model achieved AUC scores of 0.899, 0.846, 0.957, with an accuracy of 0.821, 0.744, 0.906, respectively. The patients who were identified as high risk by the anatomy aware model had an odds ratio of 2.16 (95% CI: 1.19, 3.86) for having progression of radiographic sacroiliitis within 2 years. Conclusion: Anatomical awareness can improve the generalizability of a deep learning model in detecting radiographic sacroiliitis. The model is published as fully open source alongside this study.
3D U-Net for segmentation of COVID-19 associated pulmonary infiltrates using transfer learning: State-of-the-art results on affordable hardware
Jan 25, 2021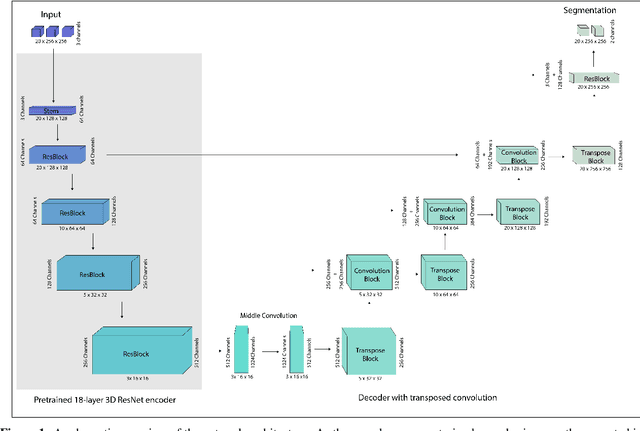
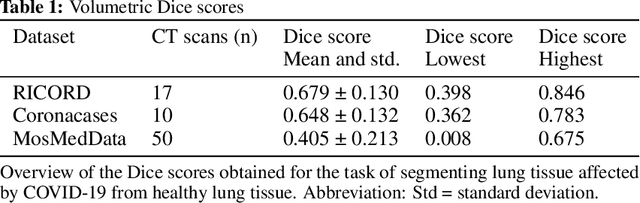

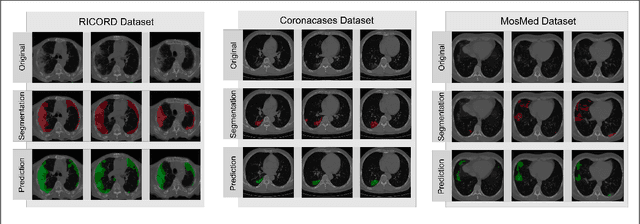
Abstract:Segmentation of pulmonary infiltrates can help assess severity of COVID-19, but manual segmentation is labor and time-intensive. Using neural networks to segment pulmonary infiltrates would enable automation of this task. However, training a 3D U-Net from computed tomography (CT) data is time- and resource-intensive. In this work, we therefore developed and tested a solution on how transfer learning can be used to train state-of-the-art segmentation models on limited hardware and in shorter time. We use the recently published RSNA International COVID-19 Open Radiology Database (RICORD) to train a fully three-dimensional U-Net architecture using an 18-layer 3D ResNet, pretrained on the Kinetics-400 dataset as encoder. The generalization of the model was then tested on two openly available datasets of patients with COVID-19, who received chest CTs (Corona Cases and MosMed datasets). Our model performed comparable to previously published 3D U-Net architectures, achieving a mean Dice score of 0.679 on the tuning dataset, 0.648 on the Coronacases dataset and 0.405 on the MosMed dataset. Notably, these results were achieved with shorter training time on a single GPU with less memory available than the GPUs used in previous studies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge