Felix J. Dorfner
MHub.ai: A Simple, Standardized, and Reproducible Platform for AI Models in Medical Imaging
Jan 15, 2026Abstract:Artificial intelligence (AI) has the potential to transform medical imaging by automating image analysis and accelerating clinical research. However, research and clinical use are limited by the wide variety of AI implementations and architectures, inconsistent documentation, and reproducibility issues. Here, we introduce MHub.ai, an open-source, container-based platform that standardizes access to AI models with minimal configuration, promoting accessibility and reproducibility in medical imaging. MHub.ai packages models from peer-reviewed publications into standardized containers that support direct processing of DICOM and other formats, provide a unified application interface, and embed structured metadata. Each model is accompanied by publicly available reference data that can be used to confirm model operation. MHub.ai includes an initial set of state-of-the-art segmentation, prediction, and feature extraction models for different modalities. The modular framework enables adaptation of any model and supports community contributions. We demonstrate the utility of the platform in a clinical use case through comparative evaluation of lung segmentation models. To further strengthen transparency and reproducibility, we publicly release the generated segmentations and evaluation metrics and provide interactive dashboards that allow readers to inspect individual cases and reproduce or extend our analysis. By simplifying model use, MHub.ai enables side-by-side benchmarking with identical execution commands and standardized outputs, and lowers the barrier to clinical translation.
DBT-DINO: Towards Foundation model based analysis of Digital Breast Tomosynthesis
Dec 15, 2025



Abstract:Foundation models have shown promise in medical imaging but remain underexplored for three-dimensional imaging modalities. No foundation model currently exists for Digital Breast Tomosynthesis (DBT), despite its use for breast cancer screening. To develop and evaluate a foundation model for DBT (DBT-DINO) across multiple clinical tasks and assess the impact of domain-specific pre-training. Self-supervised pre-training was performed using the DINOv2 methodology on over 25 million 2D slices from 487,975 DBT volumes from 27,990 patients. Three downstream tasks were evaluated: (1) breast density classification using 5,000 screening exams; (2) 5-year risk of developing breast cancer using 106,417 screening exams; and (3) lesion detection using 393 annotated volumes. For breast density classification, DBT-DINO achieved an accuracy of 0.79 (95\% CI: 0.76--0.81), outperforming both the MetaAI DINOv2 baseline (0.73, 95\% CI: 0.70--0.76, p<.001) and DenseNet-121 (0.74, 95\% CI: 0.71--0.76, p<.001). For 5-year breast cancer risk prediction, DBT-DINO achieved an AUROC of 0.78 (95\% CI: 0.76--0.80) compared to DINOv2's 0.76 (95\% CI: 0.74--0.78, p=.57). For lesion detection, DINOv2 achieved a higher average sensitivity of 0.67 (95\% CI: 0.60--0.74) compared to DBT-DINO with 0.62 (95\% CI: 0.53--0.71, p=.60). DBT-DINO demonstrated better performance on cancerous lesions specifically with a detection rate of 78.8\% compared to Dinov2's 77.3\%. Using a dataset of unprecedented size, we developed DBT-DINO, the first foundation model for DBT. DBT-DINO demonstrated strong performance on breast density classification and cancer risk prediction. However, domain-specific pre-training showed variable benefits on the detection task, with ImageNet baseline outperforming DBT-DINO on general lesion detection, indicating that localized detection tasks require further methodological development.
Scalable Drift Monitoring in Medical Imaging AI
Oct 17, 2024



Abstract:The integration of artificial intelligence (AI) into medical imaging has advanced clinical diagnostics but poses challenges in managing model drift and ensuring long-term reliability. To address these challenges, we develop MMC+, an enhanced framework for scalable drift monitoring, building upon the CheXstray framework that introduced real-time drift detection for medical imaging AI models using multi-modal data concordance. This work extends the original framework's methodologies, providing a more scalable and adaptable solution for real-world healthcare settings and offers a reliable and cost-effective alternative to continuous performance monitoring addressing limitations of both continuous and periodic monitoring methods. MMC+ introduces critical improvements to the original framework, including more robust handling of diverse data streams, improved scalability with the integration of foundation models like MedImageInsight for high-dimensional image embeddings without site-specific training, and the introduction of uncertainty bounds to better capture drift in dynamic clinical environments. Validated with real-world data from Massachusetts General Hospital during the COVID-19 pandemic, MMC+ effectively detects significant data shifts and correlates them with model performance changes. While not directly predicting performance degradation, MMC+ serves as an early warning system, indicating when AI systems may deviate from acceptable performance bounds and enabling timely interventions. By emphasizing the importance of monitoring diverse data streams and evaluating data shifts alongside model performance, this work contributes to the broader adoption and integration of AI solutions in clinical settings.
Biomedical Large Languages Models Seem not to be Superior to Generalist Models on Unseen Medical Data
Aug 25, 2024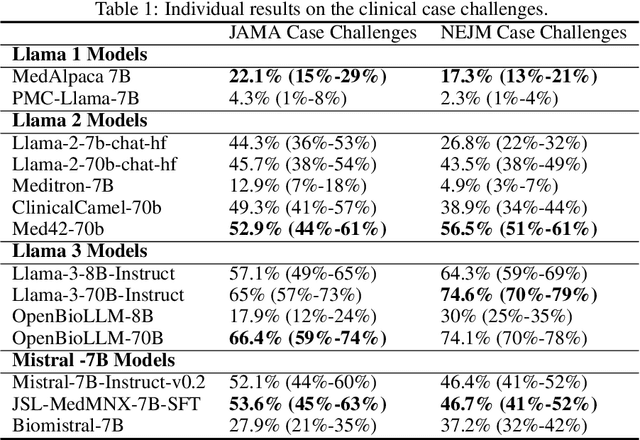
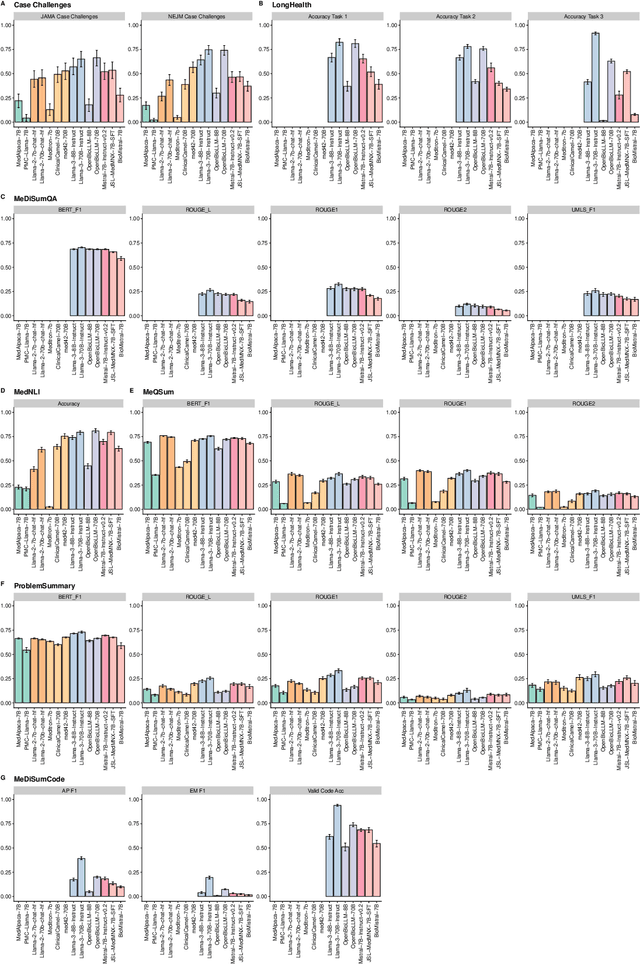
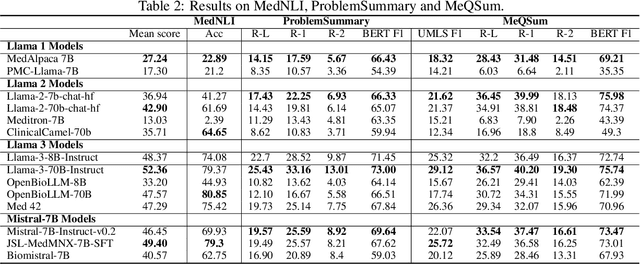
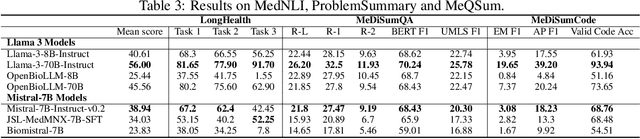
Abstract:Large language models (LLMs) have shown potential in biomedical applications, leading to efforts to fine-tune them on domain-specific data. However, the effectiveness of this approach remains unclear. This study evaluates the performance of biomedically fine-tuned LLMs against their general-purpose counterparts on a variety of clinical tasks. We evaluated their performance on clinical case challenges from the New England Journal of Medicine (NEJM) and the Journal of the American Medical Association (JAMA) and on several clinical tasks (e.g., information extraction, document summarization, and clinical coding). Using benchmarks specifically chosen to be likely outside the fine-tuning datasets of biomedical models, we found that biomedical LLMs mostly perform inferior to their general-purpose counterparts, especially on tasks not focused on medical knowledge. While larger models showed similar performance on case tasks (e.g., OpenBioLLM-70B: 66.4% vs. Llama-3-70B-Instruct: 65% on JAMA cases), smaller biomedical models showed more pronounced underperformance (e.g., OpenBioLLM-8B: 30% vs. Llama-3-8B-Instruct: 64.3% on NEJM cases). Similar trends were observed across the CLUE (Clinical Language Understanding Evaluation) benchmark tasks, with general-purpose models often performing better on text generation, question answering, and coding tasks. Our results suggest that fine-tuning LLMs to biomedical data may not provide the expected benefits and may potentially lead to reduced performance, challenging prevailing assumptions about domain-specific adaptation of LLMs and highlighting the need for more rigorous evaluation frameworks in healthcare AI. Alternative approaches, such as retrieval-augmented generation, may be more effective in enhancing the biomedical capabilities of LLMs without compromising their general knowledge.
Incorporating Anatomical Awareness for Enhanced Generalizability and Progression Prediction in Deep Learning-Based Radiographic Sacroiliitis Detection
May 12, 2024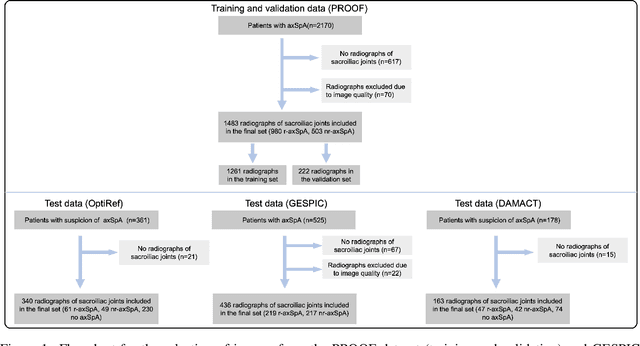
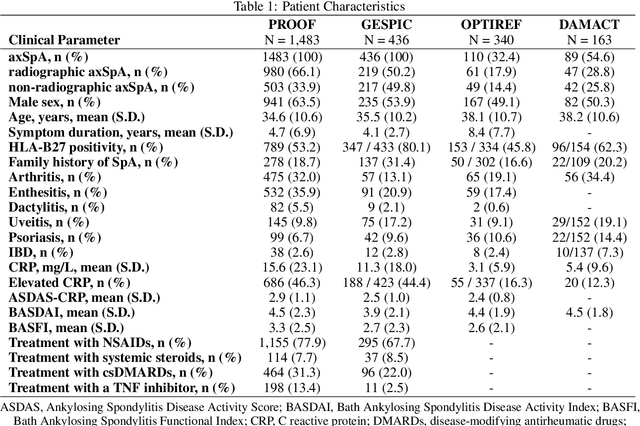
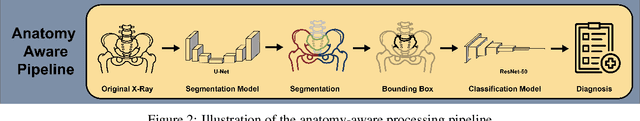

Abstract:Purpose: To examine whether incorporating anatomical awareness into a deep learning model can improve generalizability and enable prediction of disease progression. Methods: This retrospective multicenter study included conventional pelvic radiographs of 4 different patient cohorts focusing on axial spondyloarthritis (axSpA) collected at university and community hospitals. The first cohort, which consisted of 1483 radiographs, was split into training (n=1261) and validation (n=222) sets. The other cohorts comprising 436, 340, and 163 patients, respectively, were used as independent test datasets. For the second cohort, follow-up data of 311 patients was used to examine progression prediction capabilities. Two neural networks were trained, one on images cropped to the bounding box of the sacroiliac joints (anatomy-aware) and the other one on full radiographs. The performance of the models was compared using the area under the receiver operating characteristic curve (AUC), accuracy, sensitivity, and specificity. Results: On the three test datasets, the standard model achieved AUC scores of 0.853, 0.817, 0.947, with an accuracy of 0.770, 0.724, 0.850. Whereas the anatomy-aware model achieved AUC scores of 0.899, 0.846, 0.957, with an accuracy of 0.821, 0.744, 0.906, respectively. The patients who were identified as high risk by the anatomy aware model had an odds ratio of 2.16 (95% CI: 1.19, 3.86) for having progression of radiographic sacroiliitis within 2 years. Conclusion: Anatomical awareness can improve the generalizability of a deep learning model in detecting radiographic sacroiliitis. The model is published as fully open source alongside this study.
MRSegmentator: Robust Multi-Modality Segmentation of 40 Classes in MRI and CT Sequences
May 10, 2024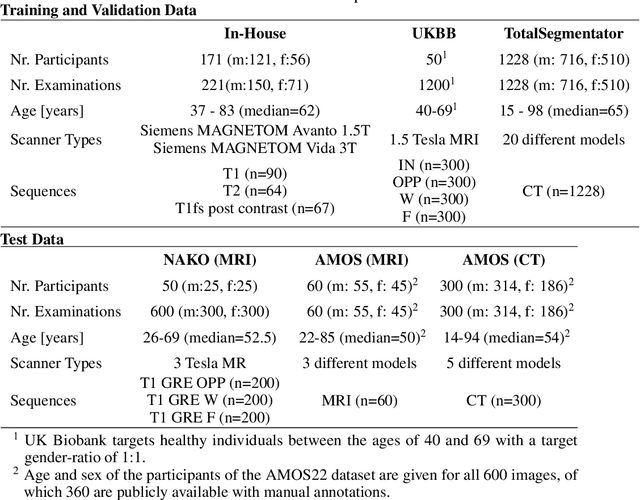
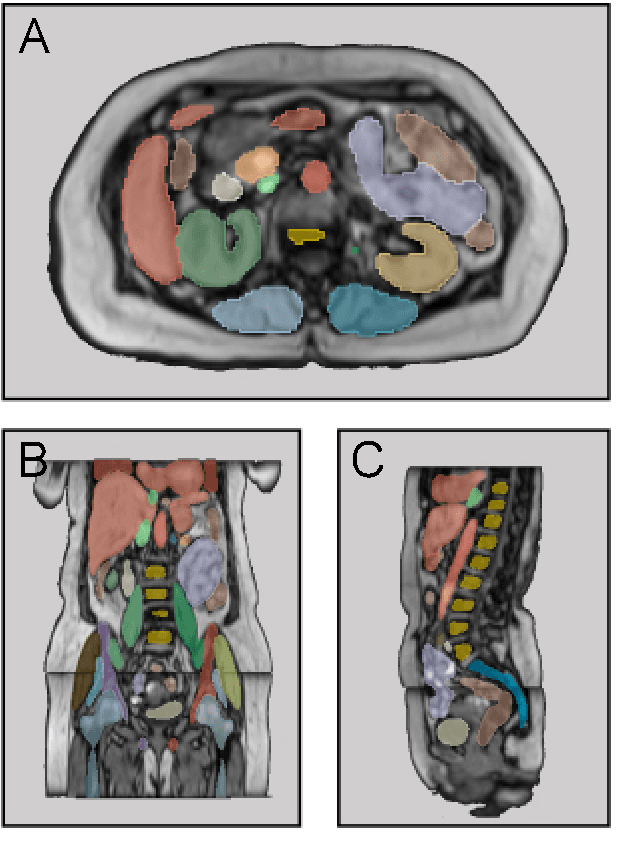
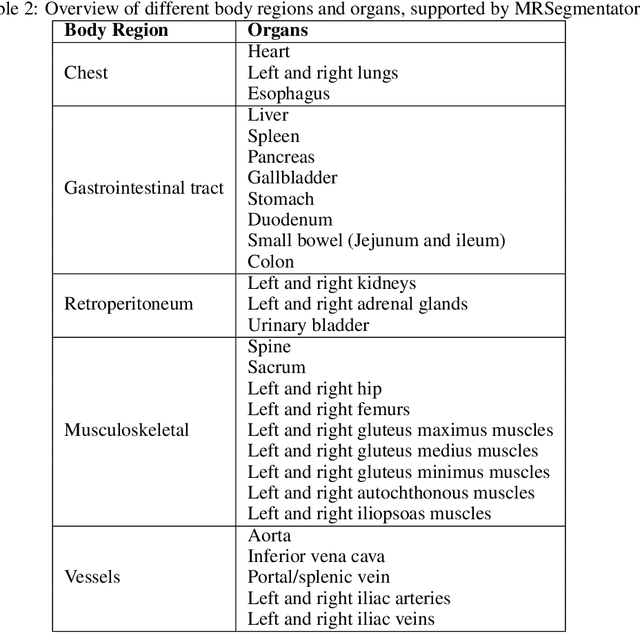
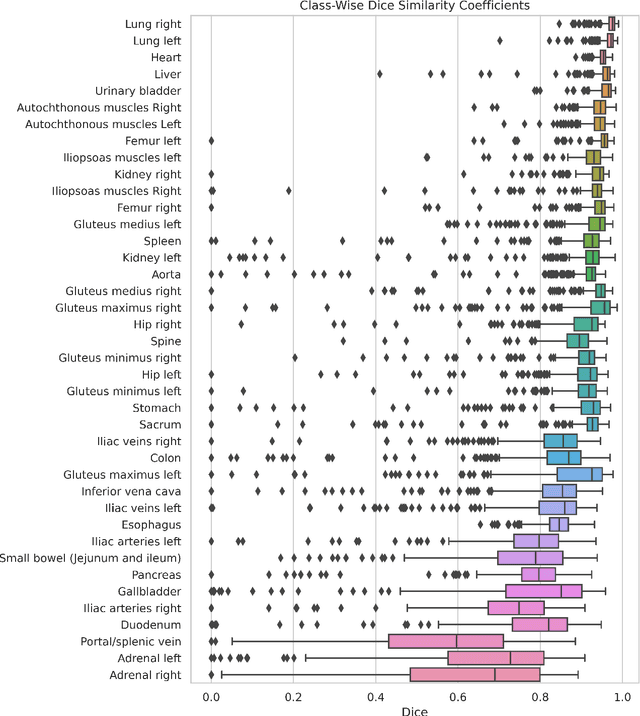
Abstract:Purpose: To introduce a deep learning model capable of multi-organ segmentation in MRI scans, offering a solution to the current limitations in MRI analysis due to challenges in resolution, standardized intensity values, and variability in sequences. Materials and Methods: he model was trained on 1,200 manually annotated MRI scans from the UK Biobank, 221 in-house MRI scans and 1228 CT scans, leveraging cross-modality transfer learning from CT segmentation models. A human-in-the-loop annotation workflow was employed to efficiently create high-quality segmentations. The model's performance was evaluated on NAKO and the AMOS22 dataset containing 600 and 60 MRI examinations. Dice Similarity Coefficient (DSC) and Hausdorff Distance (HD) was used to assess segmentation accuracy. The model will be open sourced. Results: The model showcased high accuracy in segmenting well-defined organs, achieving Dice Similarity Coefficient (DSC) scores of 0.97 for the right and left lungs, and 0.95 for the heart. It also demonstrated robustness in organs like the liver (DSC: 0.96) and kidneys (DSC: 0.95 left, 0.95 right), which present more variability. However, segmentation of smaller and complex structures such as the portal and splenic veins (DSC: 0.54) and adrenal glands (DSC: 0.65 left, 0.61 right) revealed the need for further model optimization. Conclusion: The proposed model is a robust, tool for accurate segmentation of 40 anatomical structures in MRI and CT images. By leveraging cross-modality learning and interactive annotation, the model achieves strong performance and generalizability across diverse datasets, making it a valuable resource for researchers and clinicians. It is open source and can be downloaded from https://github.com/hhaentze/MRSegmentator.
Improve Cross-Modality Segmentation by Treating MRI Images as Inverted CT Scans
May 04, 2024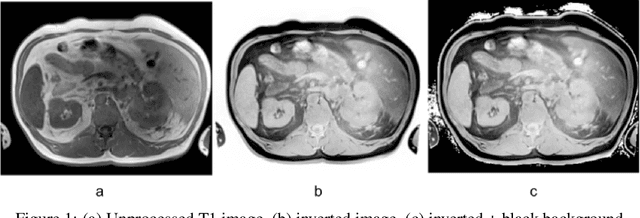
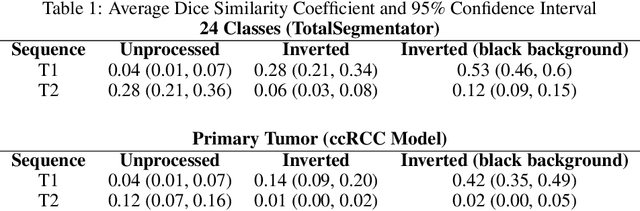

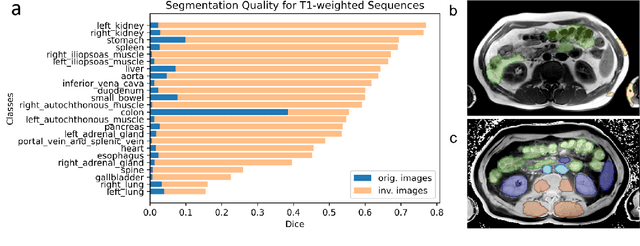
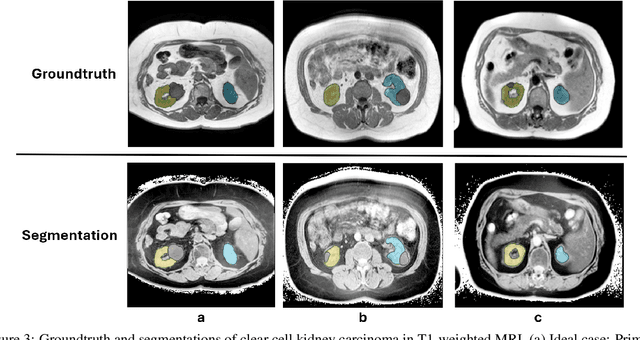
Abstract:Computed tomography (CT) segmentation models frequently include classes that are not currently supported by magnetic resonance imaging (MRI) segmentation models. In this study, we show that a simple image inversion technique can significantly improve the segmentation quality of CT segmentation models on MRI data, by using the TotalSegmentator model, applied to T1-weighted MRI images, as example. Image inversion is straightforward to implement and does not require dedicated graphics processing units (GPUs), thus providing a quick alternative to complex deep modality-transfer models for generating segmentation masks for MRI data.
Is Open-Source There Yet? A Comparative Study on Commercial and Open-Source LLMs in Their Ability to Label Chest X-Ray Reports
Feb 19, 2024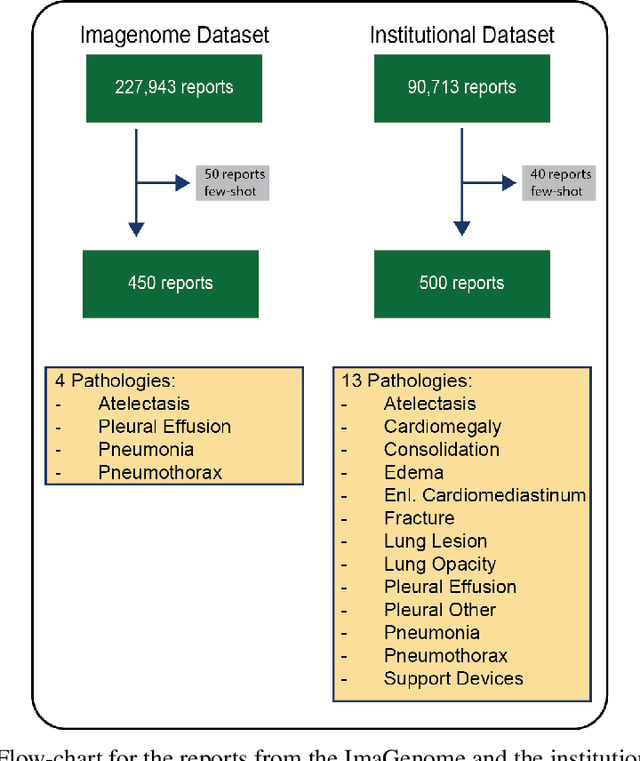
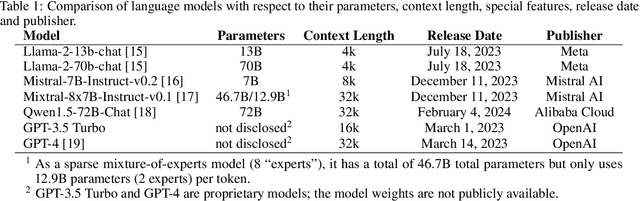
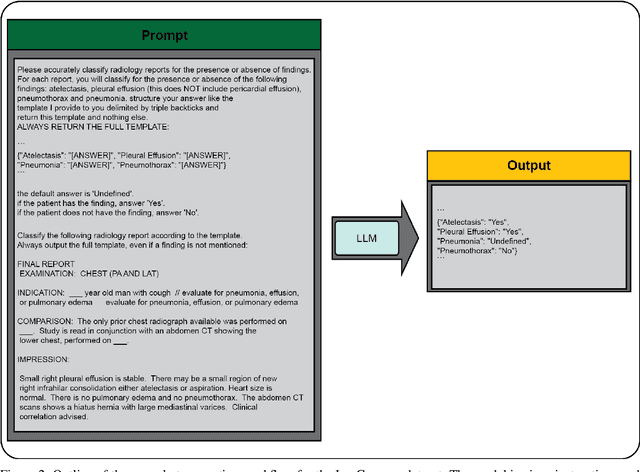
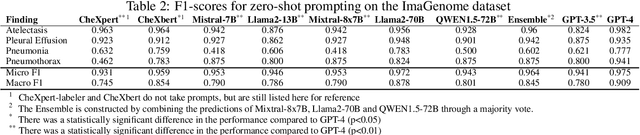
Abstract:Introduction: With the rapid advances in large language models (LLMs), there have been numerous new open source as well as commercial models. While recent publications have explored GPT-4 in its application to extracting information of interest from radiology reports, there has not been a real-world comparison of GPT-4 to different leading open-source models. Materials and Methods: Two different and independent datasets were used. The first dataset consists of 540 chest x-ray reports that were created at the Massachusetts General Hospital between July 2019 and July 2021. The second dataset consists of 500 chest x-ray reports from the ImaGenome dataset. We then compared the commercial models GPT-3.5 Turbo and GPT-4 from OpenAI to the open-source models Mistral-7B, Mixtral-8x7B, Llama2-13B, Llama2-70B, QWEN1.5-72B and CheXbert and CheXpert-labeler in their ability to accurately label the presence of multiple findings in x-ray text reports using different prompting techniques. Results: On the ImaGenome dataset, the best performing open-source model was Llama2-70B with micro F1-scores of 0.972 and 0.970 for zero- and few-shot prompts, respectively. GPT-4 achieved micro F1-scores of 0.975 and 0.984, respectively. On the institutional dataset, the best performing open-source model was QWEN1.5-72B with micro F1-scores of 0.952 and 0.965 for zero- and few-shot prompting, respectively. GPT-4 achieved micro F1-scores of 0.975 and 0.973, respectively. Conclusion: In this paper, we show that while GPT-4 is superior to open-source models in zero-shot report labeling, the implementation of few-shot prompting can bring open-source models on par with GPT-4. This shows that open-source models could be a performant and privacy preserving alternative to GPT-4 for the task of radiology report classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge