Mathias Prokop
and on behalf of the UNICORN consortium
Designing UNICORN: a Unified Benchmark for Imaging in Computational Pathology, Radiology, and Natural Language
Mar 03, 2026Abstract:Medical foundation models show promise to learn broadly generalizable features from large, diverse datasets. This could be the base for reliable cross-modality generalization and rapid adaptation to new, task-specific goals, with only a few task-specific examples. Yet, evidence for this is limited by the lack of public, standardized, and reproducible evaluation frameworks, as existing public benchmarks are often fragmented across task-, organ-, or modality-specific settings, limiting assessment of cross-task generalization. We introduce UNICORN, a public benchmark designed to systematically evaluate medical foundation models under a unified protocol. To isolate representation quality, we built the benchmark on a novel two-step framework that decouples model inference from task-specific evaluation based on standardized few-shot adaptation. As a central design choice, we constructed indirectly accessible sequestered test sets derived from clinically relevant cohorts, along with standardized evaluation code and a submission interface on an open benchmarking platform. Performance is aggregated into a single UNICORN Score, a new metric that we introduce to support direct comparison of foundation models across diverse medical domains, modalities, and task types. The UNICORN test dataset includes data from more than 2,400 patients, including over 3,700 vision cases and over 2,400 clinical reports collected from 17 institutions across eight countries. The benchmark spans eight anatomical regions and four imaging modalities. Both task-specific and aggregated leaderboards enable accessible, standardized, and reproducible evaluation. By standardizing multi-task, multi-modality assessment, UNICORN establishes a foundation for reproducible benchmarking of medical foundation models. Data, baseline methods, and the evaluation platform are publicly available via unicorn.grand-challenge.org.
ULS+: Data-driven Model Adaptation Enhances Lesion Segmentation
Jan 06, 2026Abstract:In this study, we present ULS+, an enhanced version of the Universal Lesion Segmentation (ULS) model. The original ULS model segments lesions across the whole body in CT scans given volumes of interest (VOIs) centered around a click-point. Since its release, several new public datasets have become available that can further improve model performance. ULS+ incorporates these additional datasets and uses smaller input image sizes, resulting in higher accuracy and faster inference. We compared ULS and ULS+ using the Dice score and robustness to click-point location on the ULS23 Challenge test data and a subset of the Longitudinal-CT dataset. In all comparisons, ULS+ significantly outperformed ULS. Additionally, ULS+ ranks first on the ULS23 Challenge test-phase leaderboard. By maintaining a cycle of data-driven updates and clinical validation, ULS+ establishes a foundation for robust and clinically relevant lesion segmentation models.
Robust Kidney Abnormality Segmentation: A Validation Study of an AI-Based Framework
May 12, 2025Abstract:Kidney abnormality segmentation has important potential to enhance the clinical workflow, especially in settings requiring quantitative assessments. Kidney volume could serve as an important biomarker for renal diseases, with changes in volume correlating directly with kidney function. Currently, clinical practice often relies on subjective visual assessment for evaluating kidney size and abnormalities, including tumors and cysts, which are typically staged based on diameter, volume, and anatomical location. To support a more objective and reproducible approach, this research aims to develop a robust, thoroughly validated kidney abnormality segmentation algorithm, made publicly available for clinical and research use. We employ publicly available training datasets and leverage the state-of-the-art medical image segmentation framework nnU-Net. Validation is conducted using both proprietary and public test datasets, with segmentation performance quantified by Dice coefficient and the 95th percentile Hausdorff distance. Furthermore, we analyze robustness across subgroups based on patient sex, age, CT contrast phases, and tumor histologic subtypes. Our findings demonstrate that our segmentation algorithm, trained exclusively on publicly available data, generalizes effectively to external test sets and outperforms existing state-of-the-art models across all tested datasets. Subgroup analyses reveal consistent high performance, indicating strong robustness and reliability. The developed algorithm and associated code are publicly accessible at https://github.com/DIAGNijmegen/oncology-kidney-abnormality-segmentation.
Divide to Conquer: A Field Decomposition Approach for Multi-Organ Whole-Body CT Image Registration
Mar 28, 2025Abstract:Image registration is an essential technique for the analysis of Computed Tomography (CT) images in clinical practice. However, existing methodologies are predominantly tailored to a specific organ of interest and often exhibit lower performance on other organs, thus limiting their generalizability and applicability. Multi-organ registration addresses these limitations, but the simultaneous alignment of multiple organs with diverse shapes, sizes and locations requires a highly complex deformation field with a multi-layer composition of individual deformations. This study introduces a novel field decomposition approach to address the high complexity of deformations in multi-organ whole-body CT image registration. The proposed method is trained and evaluated on a longitudinal dataset of 691 patients, each with two CT images obtained at distinct time points. These scans fully encompass the thoracic, abdominal, and pelvic regions. Two baseline registration methods are selected for this study: one based on optimization techniques and another based on deep learning. Experimental results demonstrate that the proposed approach outperforms baseline methods in handling complex deformations in multi-organ whole-body CT image registration.
MRSegmentator: Robust Multi-Modality Segmentation of 40 Classes in MRI and CT Sequences
May 10, 2024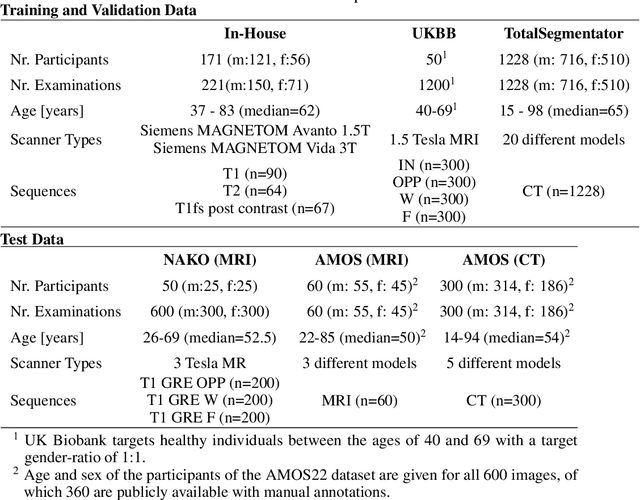
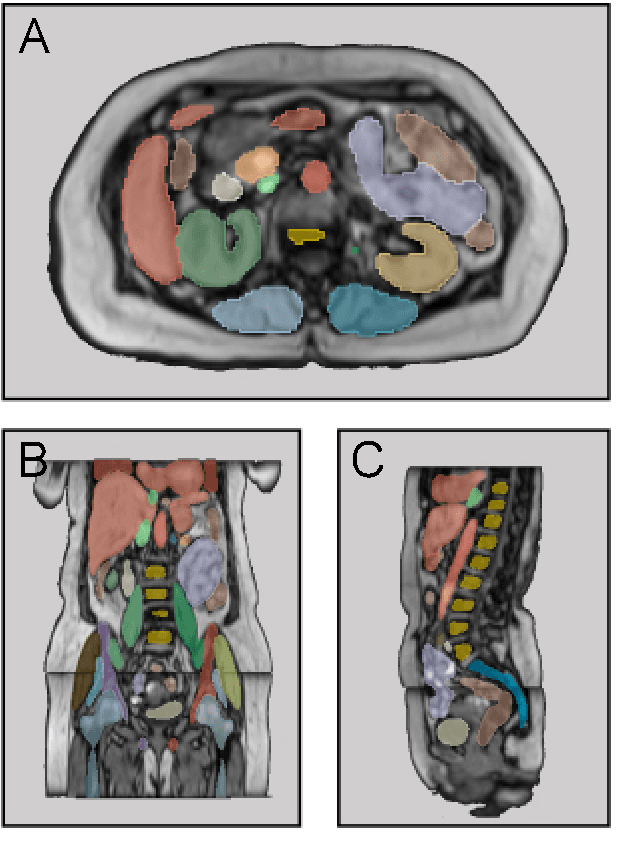
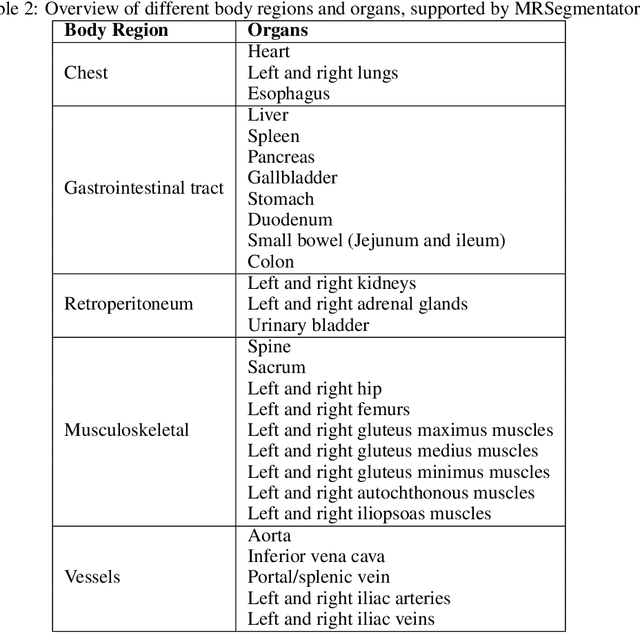
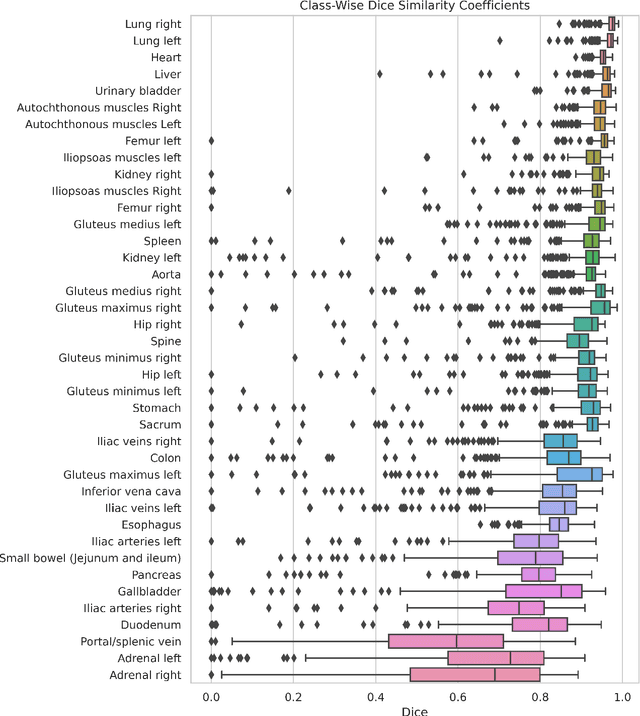
Abstract:Purpose: To introduce a deep learning model capable of multi-organ segmentation in MRI scans, offering a solution to the current limitations in MRI analysis due to challenges in resolution, standardized intensity values, and variability in sequences. Materials and Methods: he model was trained on 1,200 manually annotated MRI scans from the UK Biobank, 221 in-house MRI scans and 1228 CT scans, leveraging cross-modality transfer learning from CT segmentation models. A human-in-the-loop annotation workflow was employed to efficiently create high-quality segmentations. The model's performance was evaluated on NAKO and the AMOS22 dataset containing 600 and 60 MRI examinations. Dice Similarity Coefficient (DSC) and Hausdorff Distance (HD) was used to assess segmentation accuracy. The model will be open sourced. Results: The model showcased high accuracy in segmenting well-defined organs, achieving Dice Similarity Coefficient (DSC) scores of 0.97 for the right and left lungs, and 0.95 for the heart. It also demonstrated robustness in organs like the liver (DSC: 0.96) and kidneys (DSC: 0.95 left, 0.95 right), which present more variability. However, segmentation of smaller and complex structures such as the portal and splenic veins (DSC: 0.54) and adrenal glands (DSC: 0.65 left, 0.61 right) revealed the need for further model optimization. Conclusion: The proposed model is a robust, tool for accurate segmentation of 40 anatomical structures in MRI and CT images. By leveraging cross-modality learning and interactive annotation, the model achieves strong performance and generalizability across diverse datasets, making it a valuable resource for researchers and clinicians. It is open source and can be downloaded from https://github.com/hhaentze/MRSegmentator.
Transfer learning from a sparsely annotated dataset of 3D medical images
Nov 08, 2023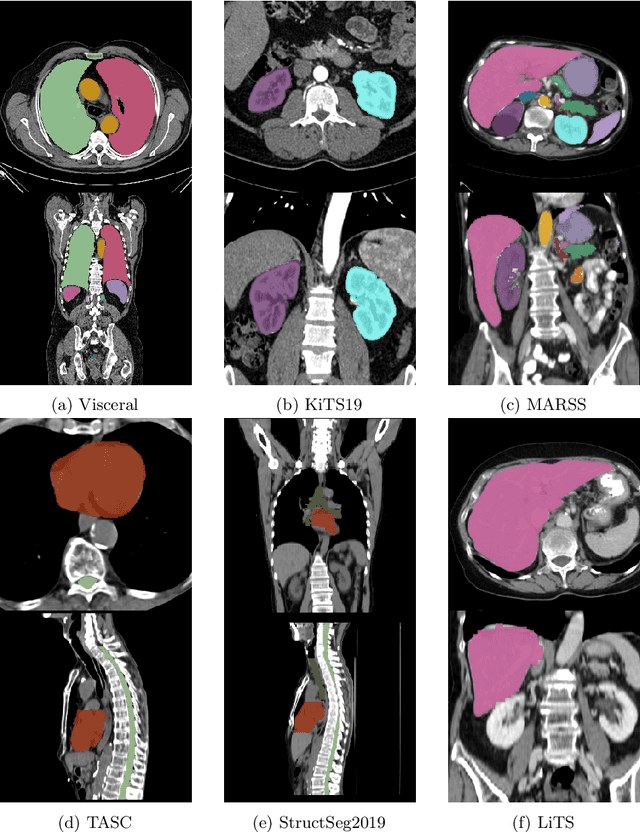
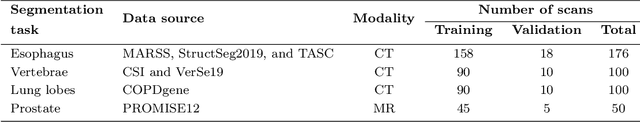
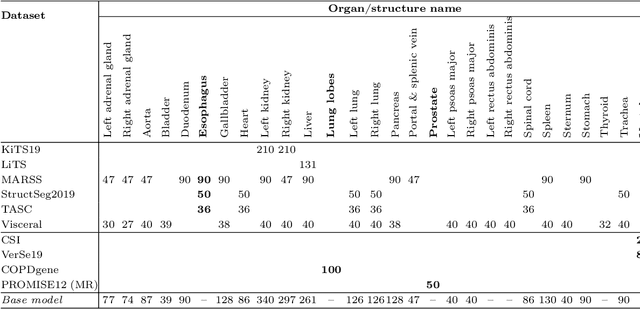
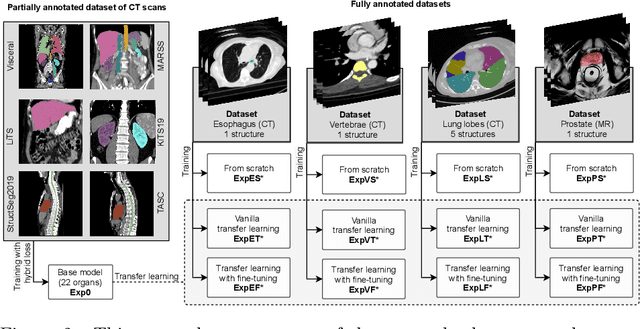

Abstract:Transfer learning leverages pre-trained model features from a large dataset to save time and resources when training new models for various tasks, potentially enhancing performance. Due to the lack of large datasets in the medical imaging domain, transfer learning from one medical imaging model to other medical imaging models has not been widely explored. This study explores the use of transfer learning to improve the performance of deep convolutional neural networks for organ segmentation in medical imaging. A base segmentation model (3D U-Net) was trained on a large and sparsely annotated dataset; its weights were used for transfer learning on four new down-stream segmentation tasks for which a fully annotated dataset was available. We analyzed the training set size's influence to simulate scarce data. The results showed that transfer learning from the base model was beneficial when small datasets were available, providing significant performance improvements; where fine-tuning the base model is more beneficial than updating all the network weights with vanilla transfer learning. Transfer learning with fine-tuning increased the performance by up to 0.129 (+28\%) Dice score than experiments trained from scratch, and on average 23 experiments increased the performance by 0.029 Dice score in the new segmentation tasks. The study also showed that cross-modality transfer learning using CT scans was beneficial. The findings of this study demonstrate the potential of transfer learning to improve the efficiency of annotation and increase the accessibility of accurate organ segmentation in medical imaging, ultimately leading to improved patient care. We made the network definition and weights publicly available to benefit other users and researchers.
Kidney abnormality segmentation in thorax-abdomen CT scans
Sep 06, 2023
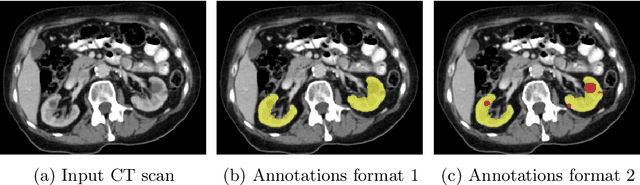


Abstract:In this study, we introduce a deep learning approach for segmenting kidney parenchyma and kidney abnormalities to support clinicians in identifying and quantifying renal abnormalities such as cysts, lesions, masses, metastases, and primary tumors. Our end-to-end segmentation method was trained on 215 contrast-enhanced thoracic-abdominal CT scans, with half of these scans containing one or more abnormalities. We began by implementing our own version of the original 3D U-Net network and incorporated four additional components: an end-to-end multi-resolution approach, a set of task-specific data augmentations, a modified loss function using top-$k$, and spatial dropout. Furthermore, we devised a tailored post-processing strategy. Ablation studies demonstrated that each of the four modifications enhanced kidney abnormality segmentation performance, while three out of four improved kidney parenchyma segmentation. Subsequently, we trained the nnUNet framework on our dataset. By ensembling the optimized 3D U-Net and the nnUNet with our specialized post-processing, we achieved marginally superior results. Our best-performing model attained Dice scores of 0.965 and 0.947 for segmenting kidney parenchyma in two test sets (20 scans without abnormalities and 30 with abnormalities), outperforming an independent human observer who scored 0.944 and 0.925, respectively. In segmenting kidney abnormalities within the 30 test scans containing them, the top-performing method achieved a Dice score of 0.585, while an independent second human observer reached a score of 0.664, suggesting potential for further improvement in computerized methods. All training data is available to the research community under a CC-BY 4.0 license on https://doi.org/10.5281/zenodo.8014289
Automated Estimation of Total Lung Volume using Chest Radiographs and Deep Learning
May 03, 2021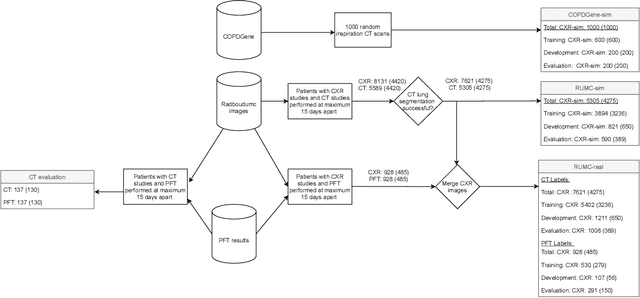
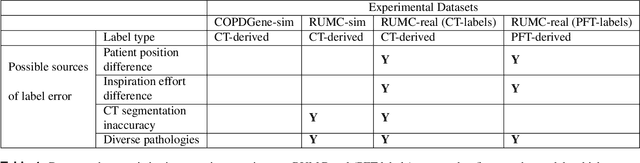
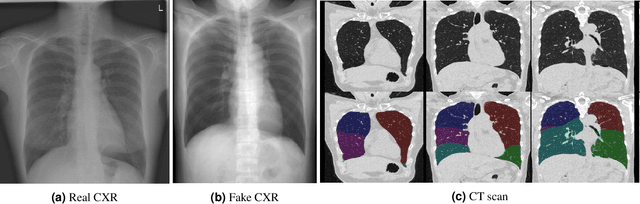
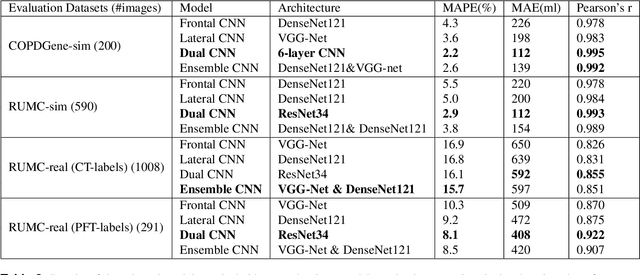
Abstract:Total lung volume is an important quantitative biomarker and is used for the assessment of restrictive lung diseases. In this study, we investigate the performance of several deep-learning approaches for automated measurement of total lung volume from chest radiographs. 7621 posteroanterior and lateral view chest radiographs (CXR) were collected from patients with chest CT available. Similarly, 928 CXR studies were chosen from patients with pulmonary function test (PFT) results. The reference total lung volume was calculated from lung segmentation on CT or PFT data, respectively. This dataset was used to train deep-learning architectures to predict total lung volume from chest radiographs. The experiments were constructed in a step-wise fashion with increasing complexity to demonstrate the effect of training with CT-derived labels only and the sources of error. The optimal models were tested on 291 CXR studies with reference lung volume obtained from PFT. The optimal deep-learning regression model showed an MAE of 408 ml and a MAPE of 8.1\% and Pearson's r = 0.92 using both frontal and lateral chest radiographs as input. CT-derived labels were useful for pre-training but the optimal performance was obtained by fine-tuning the network with PFT-derived labels. We demonstrate, for the first time, that state-of-the-art deep learning solutions can accurately measure total lung volume from plain chest radiographs. The proposed model can be used to obtain total lung volume from routinely acquired chest radiographs at no additional cost and could be a useful tool to identify trends over time in patients referred regularly for chest x-rays.
Validation, comparison, and combination of algorithms for automatic detection of pulmonary nodules in computed tomography images: the LUNA16 challenge
Jul 15, 2017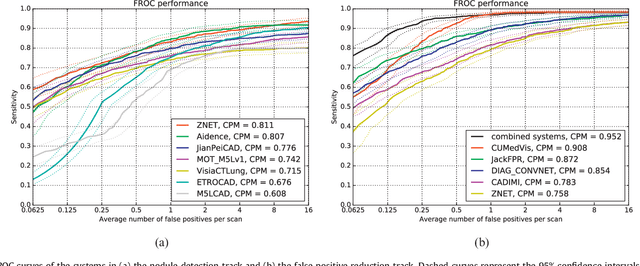
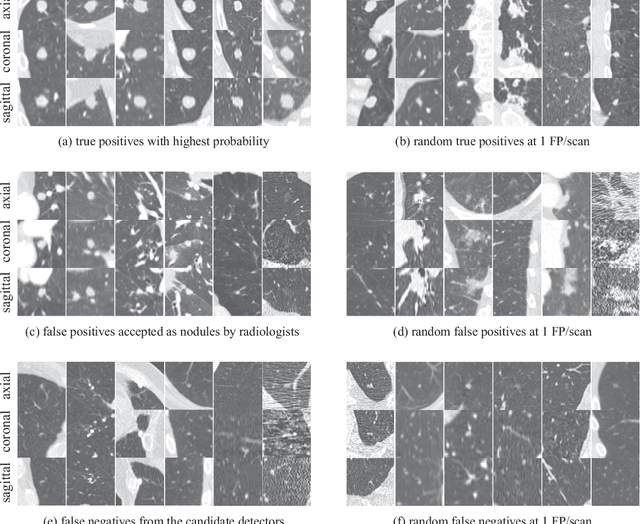
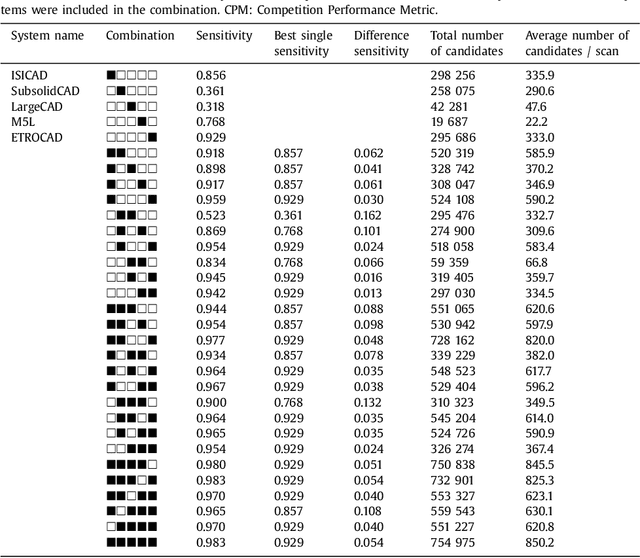

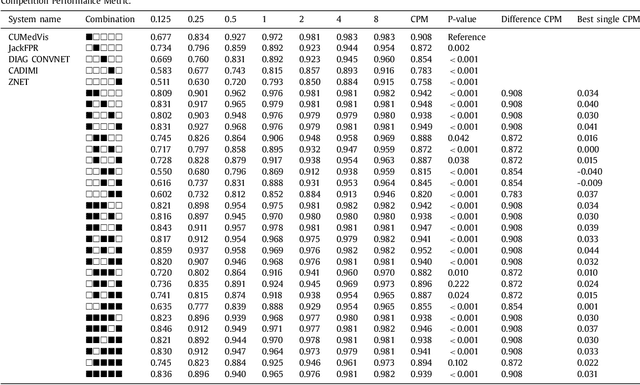
Abstract:Automatic detection of pulmonary nodules in thoracic computed tomography (CT) scans has been an active area of research for the last two decades. However, there have only been few studies that provide a comparative performance evaluation of different systems on a common database. We have therefore set up the LUNA16 challenge, an objective evaluation framework for automatic nodule detection algorithms using the largest publicly available reference database of chest CT scans, the LIDC-IDRI data set. In LUNA16, participants develop their algorithm and upload their predictions on 888 CT scans in one of the two tracks: 1) the complete nodule detection track where a complete CAD system should be developed, or 2) the false positive reduction track where a provided set of nodule candidates should be classified. This paper describes the setup of LUNA16 and presents the results of the challenge so far. Moreover, the impact of combining individual systems on the detection performance was also investigated. It was observed that the leading solutions employed convolutional networks and used the provided set of nodule candidates. The combination of these solutions achieved an excellent sensitivity of over 95% at fewer than 1.0 false positives per scan. This highlights the potential of combining algorithms to improve the detection performance. Our observer study with four expert readers has shown that the best system detects nodules that were missed by expert readers who originally annotated the LIDC-IDRI data. We released this set of additional nodules for further development of CAD systems.
Towards automatic pulmonary nodule management in lung cancer screening with deep learning
May 23, 2017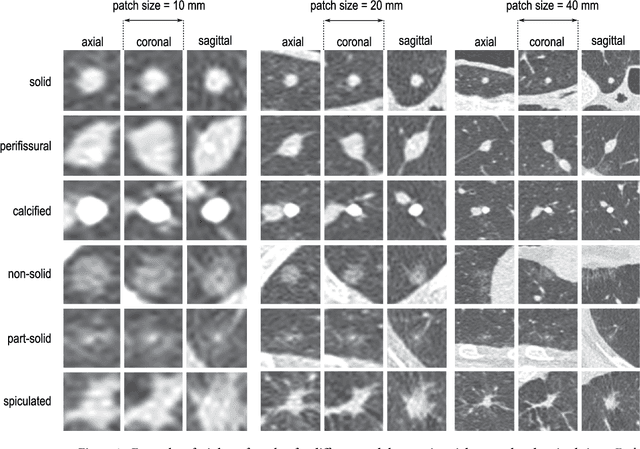
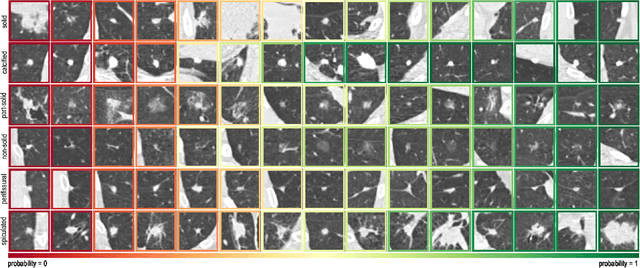
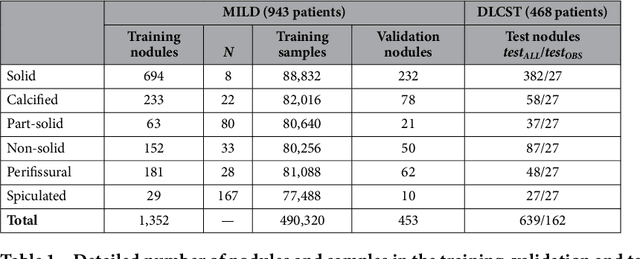
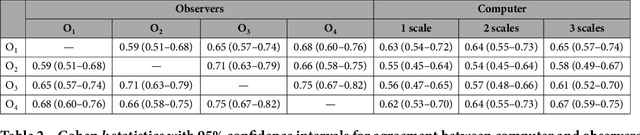

Abstract:The introduction of lung cancer screening programs will produce an unprecedented amount of chest CT scans in the near future, which radiologists will have to read in order to decide on a patient follow-up strategy. According to the current guidelines, the workup of screen-detected nodules strongly relies on nodule size and nodule type. In this paper, we present a deep learning system based on multi-stream multi-scale convolutional networks, which automatically classifies all nodule types relevant for nodule workup. The system processes raw CT data containing a nodule without the need for any additional information such as nodule segmentation or nodule size and learns a representation of 3D data by analyzing an arbitrary number of 2D views of a given nodule. The deep learning system was trained with data from the Italian MILD screening trial and validated on an independent set of data from the Danish DLCST screening trial. We analyze the advantage of processing nodules at multiple scales with a multi-stream convolutional network architecture, and we show that the proposed deep learning system achieves performance at classifying nodule type that surpasses the one of classical machine learning approaches and is within the inter-observer variability among four experienced human observers.
* Published on Scientific Reports
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge