Keelin Murphy
Nodule detection and generation on chest X-rays: NODE21 Challenge
Jan 04, 2024



Abstract:Pulmonary nodules may be an early manifestation of lung cancer, the leading cause of cancer-related deaths among both men and women. Numerous studies have established that deep learning methods can yield high-performance levels in the detection of lung nodules in chest X-rays. However, the lack of gold-standard public datasets slows down the progression of the research and prevents benchmarking of methods for this task. To address this, we organized a public research challenge, NODE21, aimed at the detection and generation of lung nodules in chest X-rays. While the detection track assesses state-of-the-art nodule detection systems, the generation track determines the utility of nodule generation algorithms to augment training data and hence improve the performance of the detection systems. This paper summarizes the results of the NODE21 challenge and performs extensive additional experiments to examine the impact of the synthetically generated nodule training images on the detection algorithm performance.
Learn2Reg: comprehensive multi-task medical image registration challenge, dataset and evaluation in the era of deep learning
Dec 23, 2021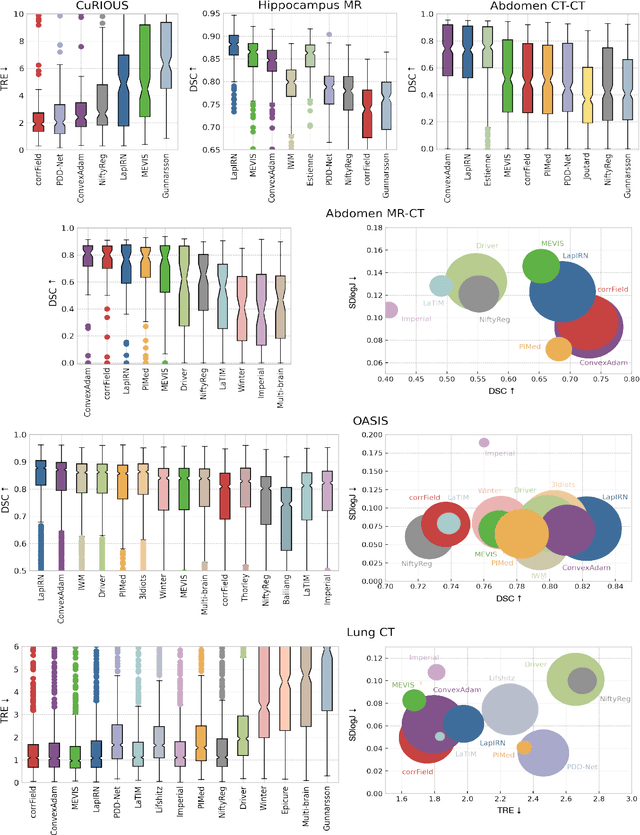
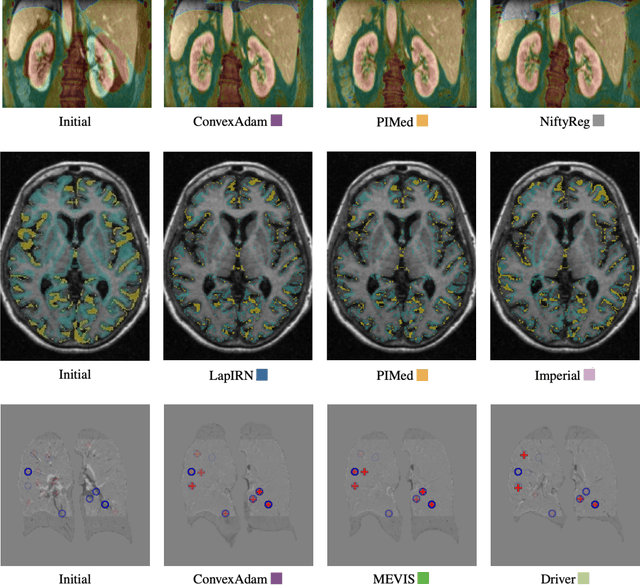
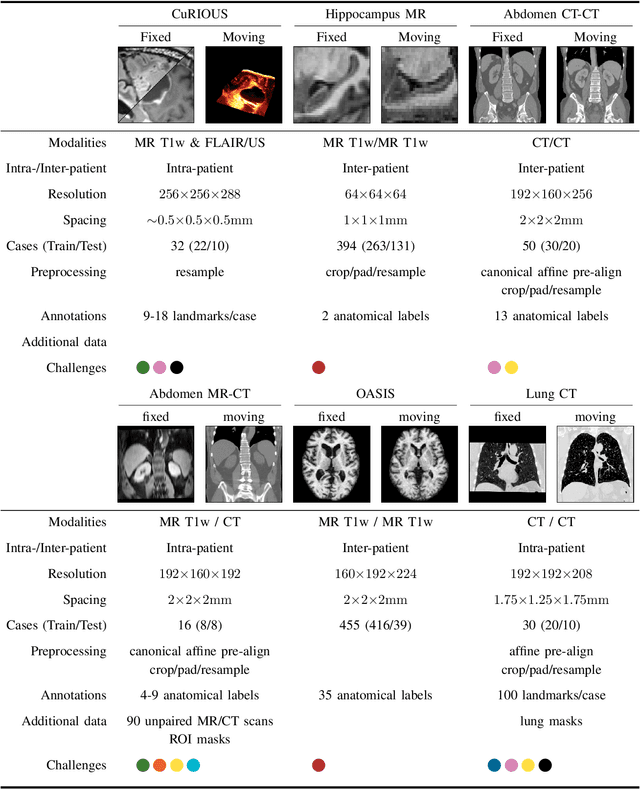
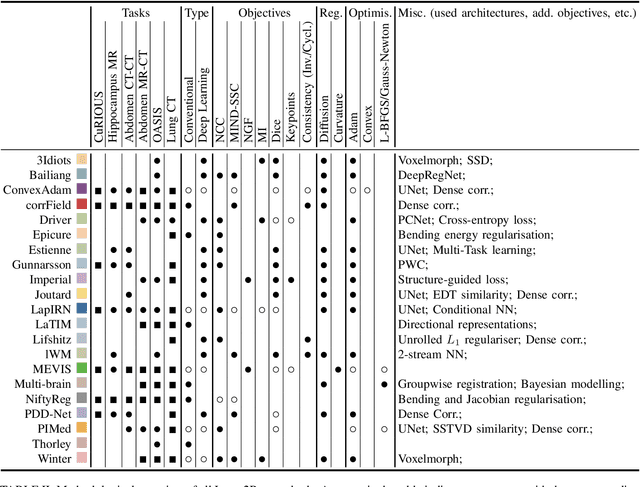
Abstract:Image registration is a fundamental medical image analysis task, and a wide variety of approaches have been proposed. However, only a few studies have comprehensively compared medical image registration approaches on a wide range of clinically relevant tasks, in part because of the lack of availability of such diverse data. This limits the development of registration methods, the adoption of research advances into practice, and a fair benchmark across competing approaches. The Learn2Reg challenge addresses these limitations by providing a multi-task medical image registration benchmark for comprehensive characterisation of deformable registration algorithms. A continuous evaluation will be possible at https://learn2reg.grand-challenge.org. Learn2Reg covers a wide range of anatomies (brain, abdomen, and thorax), modalities (ultrasound, CT, MR), availability of annotations, as well as intra- and inter-patient registration evaluation. We established an easily accessible framework for training and validation of 3D registration methods, which enabled the compilation of results of over 65 individual method submissions from more than 20 unique teams. We used a complementary set of metrics, including robustness, accuracy, plausibility, and runtime, enabling unique insight into the current state-of-the-art of medical image registration. This paper describes datasets, tasks, evaluation methods and results of the challenge, and the results of further analysis of transferability to new datasets, the importance of label supervision, and resulting bias.
Automated Estimation of Total Lung Volume using Chest Radiographs and Deep Learning
May 03, 2021
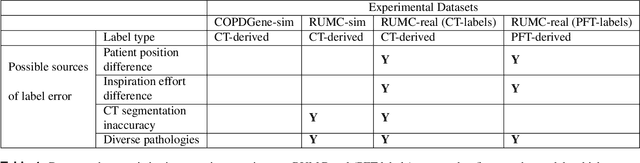
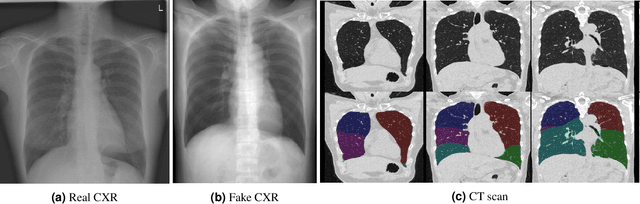
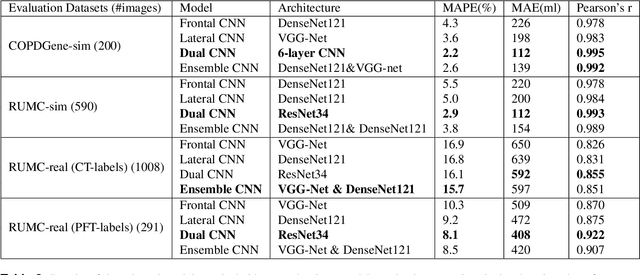
Abstract:Total lung volume is an important quantitative biomarker and is used for the assessment of restrictive lung diseases. In this study, we investigate the performance of several deep-learning approaches for automated measurement of total lung volume from chest radiographs. 7621 posteroanterior and lateral view chest radiographs (CXR) were collected from patients with chest CT available. Similarly, 928 CXR studies were chosen from patients with pulmonary function test (PFT) results. The reference total lung volume was calculated from lung segmentation on CT or PFT data, respectively. This dataset was used to train deep-learning architectures to predict total lung volume from chest radiographs. The experiments were constructed in a step-wise fashion with increasing complexity to demonstrate the effect of training with CT-derived labels only and the sources of error. The optimal models were tested on 291 CXR studies with reference lung volume obtained from PFT. The optimal deep-learning regression model showed an MAE of 408 ml and a MAPE of 8.1\% and Pearson's r = 0.92 using both frontal and lateral chest radiographs as input. CT-derived labels were useful for pre-training but the optimal performance was obtained by fine-tuning the network with PFT-derived labels. We demonstrate, for the first time, that state-of-the-art deep learning solutions can accurately measure total lung volume from plain chest radiographs. The proposed model can be used to obtain total lung volume from routinely acquired chest radiographs at no additional cost and could be a useful tool to identify trends over time in patients referred regularly for chest x-rays.
Deep Learning with robustness to missing data: A novel approach to the detection of COVID-19
Mar 25, 2021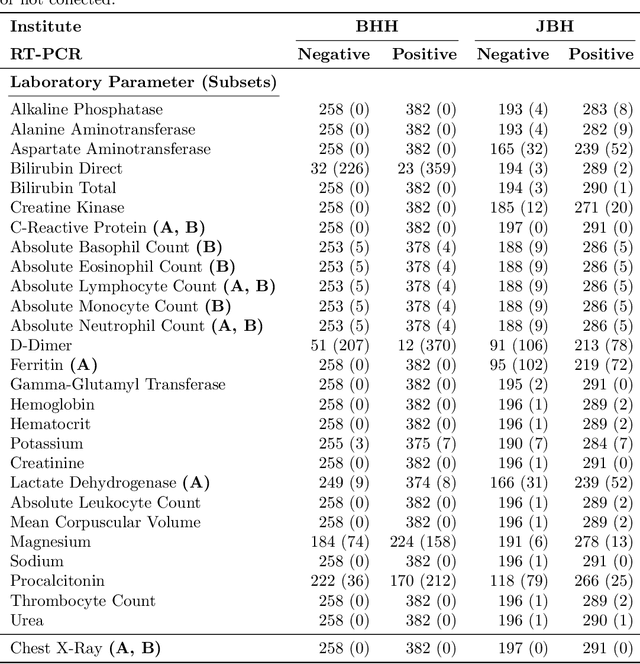
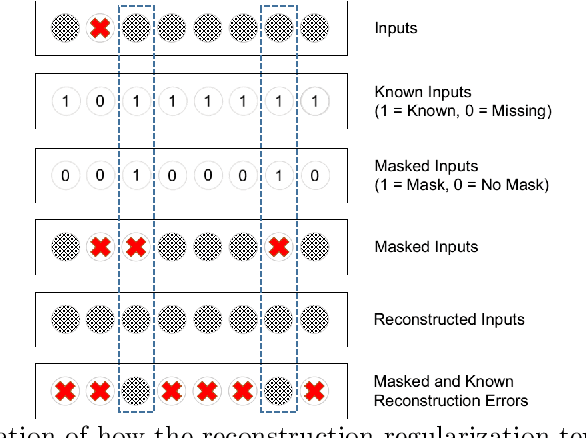
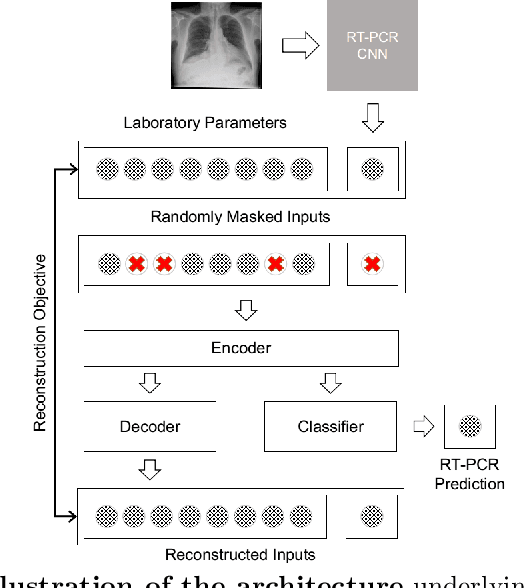
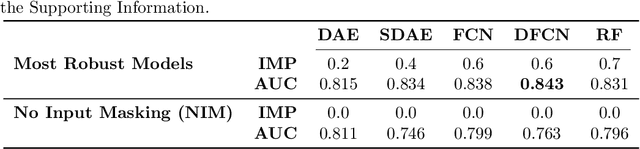
Abstract:In the context of the current global pandemic and the limitations of the RT-PCR test, we propose a novel deep learning architecture, DFCN, (Denoising Fully Connected Network) for the detection of COVID-19 using laboratory tests and chest x-rays. Since medical facilities around the world differ enormously in what laboratory tests or chest imaging may be available, DFCN is designed to be robust to missing input data. An ablation study extensively evaluates the performance benefits of the DFCN architecture as well as its robustness to missing inputs. Data from 1088 patients with confirmed RT-PCR results are obtained from two independent medical facilities. The data collected includes results from 27 laboratory tests and a chest x-ray scored by a deep learning network. Training and test datasets are defined based on the source medical facility. Data is made publicly available. The performance of DFCN in predicting the RT-PCR result is compared with 3 related architectures as well as a Random Forest baseline. All models are trained with varying levels of masked input data to encourage robustness to missing inputs. Missing data is simulated at test time by masking inputs randomly. Using area under the receiver operating curve (AUC) as a metric, DFCN outperforms all other models with statistical significance using random subsets of input data with 2-27 available inputs. When all 28 inputs are available DFCN obtains an AUC of 0.924, higher than achieved by any other model. Furthermore, with clinically meaningful subsets of parameters consisting of just 6 and 7 inputs respectively, DFCN also achieves higher AUCs than any other model, with values of 0.909 and 0.919.
Deep Learning for Chest X-ray Analysis: A Survey
Mar 15, 2021



Abstract:Recent advances in deep learning have led to a promising performance in many medical image analysis tasks. As the most commonly performed radiological exam, chest radiographs are a particularly important modality for which a variety of applications have been researched. The release of multiple, large, publicly available chest X-ray datasets in recent years has encouraged research interest and boosted the number of publications. In this paper, we review all studies using deep learning on chest radiographs, categorizing works by task: image-level prediction (classification and regression), segmentation, localization, image generation and domain adaptation. Commercially available applications are detailed, and a comprehensive discussion of the current state of the art and potential future directions are provided.
FRODO: Free rejection of out-of-distribution samples: application to chest x-ray analysis
Jul 02, 2019
Abstract:In this work, we propose a method to reject out-of-distribution samples which can be adapted to any network architecture and requires no additional training data. Publicly available chest x-ray data (38,353 images) is used to train a standard ResNet-50 model to detect emphysema. Feature activations of intermediate layers are used as descriptors defining the training data distribution. A novel metric, FRODO, is measured by using the Mahalanobis distance of a new test sample to the training data distribution. The method is tested using a held-out test dataset of 21,176 chest x-rays (in-distribution) and a set of 14,821 out-of-distribution x-ray images of incorrect orientation or anatomy. In classifying test samples as in or out-of distribution, our method achieves an AUC score of 0.99.
Computer aided detection of tuberculosis on chest radiographs: An evaluation of the CAD4TB v6 system
Mar 08, 2019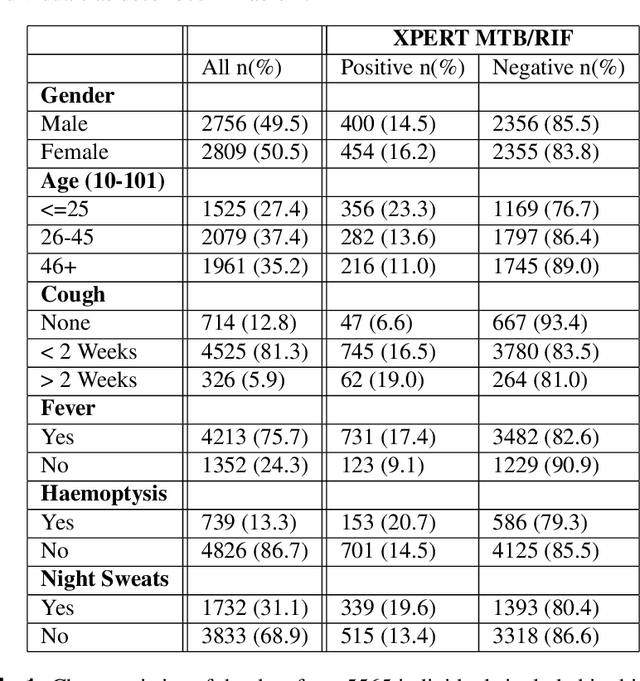
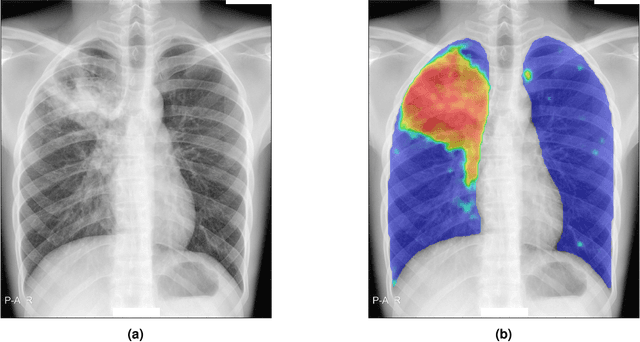
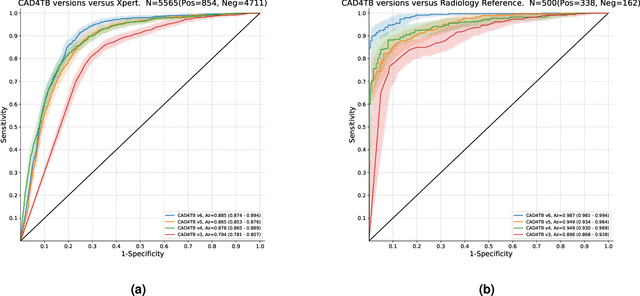
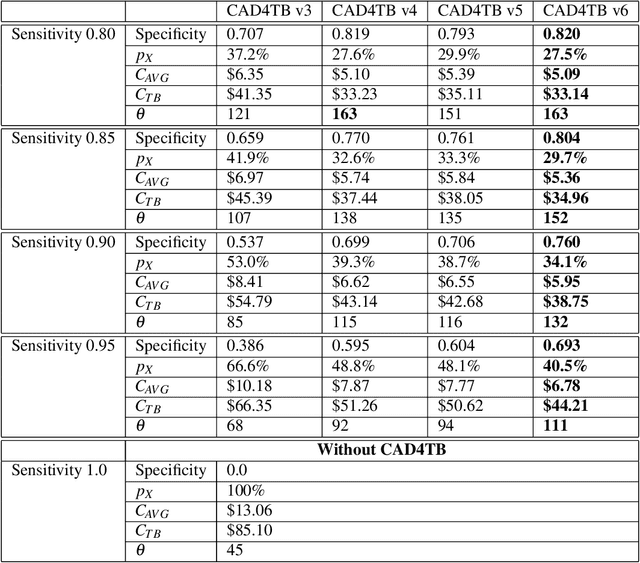
Abstract:There is a growing interest in the automated analysis of chest X-Ray (CXR) as a sensitive and inexpensive means of screening susceptible populations for pulmonary tuberculosis. In this work we evaluate the latest version of CAD4TB, a software platform designed for this purpose. Version 6 of CAD4TB was released in 2018 and is here tested on an independent dataset of 5565 CXR images with GeneXpert (Xpert) sputum test results available (854 Xpert positive subjects). A subset of 500 subjects (50% Xpert positive) was reviewed and annotated by 5 expert observers independently to obtain a radiological reference standard. The latest version of CAD4TB is found to outperform all previous versions in terms of area under receiver operating curve (ROC) with respect to both Xpert and radiological reference standards. Improvements with respect to Xpert are most apparent at high sensitivity levels with a specificity of 76% obtained at 90% sensitivity. When compared with the radiological reference standard, CAD4TB v6 also outperformed previous versions by a considerable margin and achieved 98% specificity at 90% sensitivity. No substantial difference was found between the performance of CAD4TB v6 and any of the various expert observers against the Xpert reference standard. A cost and efficiency analysis on this dataset demonstrates that in a standard clinical situation, operating at 90% sensitivity, users of CAD4TB v6 can process 132 subjects per day at an average cost per screen of \$5.95 per subject, while users of version 3 process only 85 subjects per day at a cost of \$8.41 per subject. At all tested operating points version 6 is shown to be more efficient and cost effective than any other version.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge