Marcel Bengs
Nodule detection and generation on chest X-rays: NODE21 Challenge
Jan 04, 2024



Abstract:Pulmonary nodules may be an early manifestation of lung cancer, the leading cause of cancer-related deaths among both men and women. Numerous studies have established that deep learning methods can yield high-performance levels in the detection of lung nodules in chest X-rays. However, the lack of gold-standard public datasets slows down the progression of the research and prevents benchmarking of methods for this task. To address this, we organized a public research challenge, NODE21, aimed at the detection and generation of lung nodules in chest X-rays. While the detection track assesses state-of-the-art nodule detection systems, the generation track determines the utility of nodule generation algorithms to augment training data and hence improve the performance of the detection systems. This paper summarizes the results of the NODE21 challenge and performs extensive additional experiments to examine the impact of the synthetically generated nodule training images on the detection algorithm performance.
Supervised Contrastive Learning to Classify Paranasal Anomalies in the Maxillary Sinus
Sep 05, 2022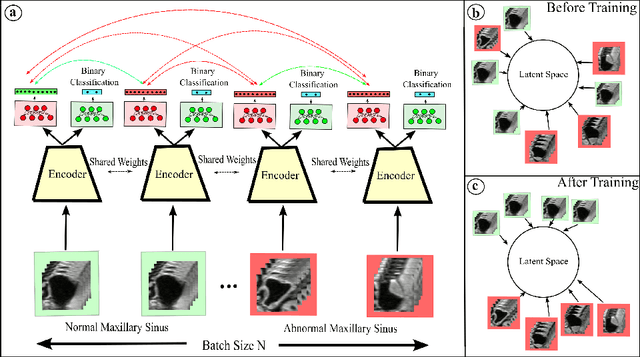
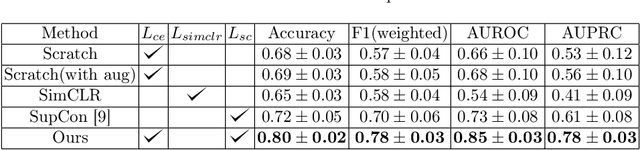
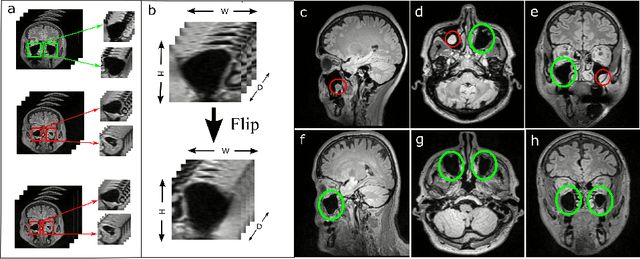
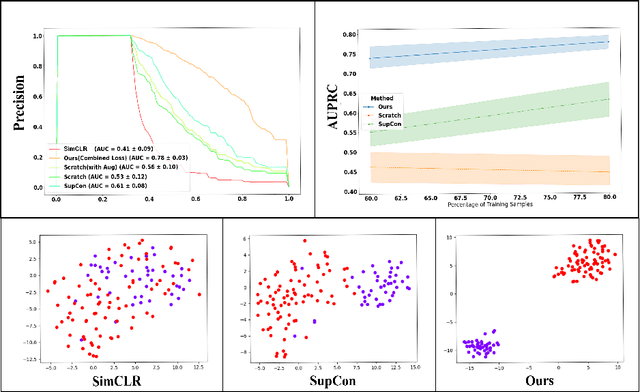
Abstract:Using deep learning techniques, anomalies in the paranasal sinus system can be detected automatically in MRI images and can be further analyzed and classified based on their volume, shape and other parameters like local contrast. However due to limited training data, traditional supervised learning methods often fail to generalize. Existing deep learning methods in paranasal anomaly classification have been used to diagnose at most one anomaly. In our work, we consider three anomalies. Specifically, we employ a 3D CNN to separate maxillary sinus volumes without anomalies from maxillary sinus volumes with anomalies. To learn robust representations from a small labelled dataset, we propose a novel learning paradigm that combines contrastive loss and cross-entropy loss. Particularly, we use a supervised contrastive loss that encourages embeddings of maxillary sinus volumes with and without anomaly to form two distinct clusters while the cross-entropy loss encourages the 3D CNN to maintain its discriminative ability. We report that optimising with both losses is advantageous over optimising with only one loss. We also find that our training strategy leads to label efficiency. With our method, a 3D CNN classifier achieves an AUROC of 0.85 while a 3D CNN classifier optimised with cross-entropy loss achieves an AUROC of 0.66.
Ultrasound Shear Wave Elasticity Imaging with Spatio-Temporal Deep Learning
Apr 28, 2022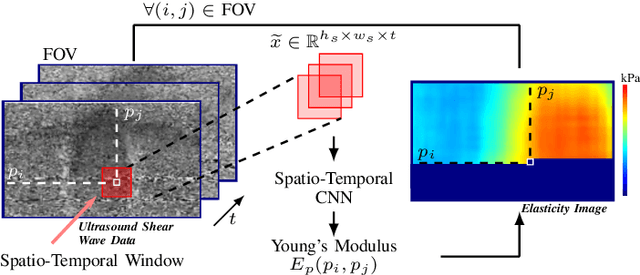
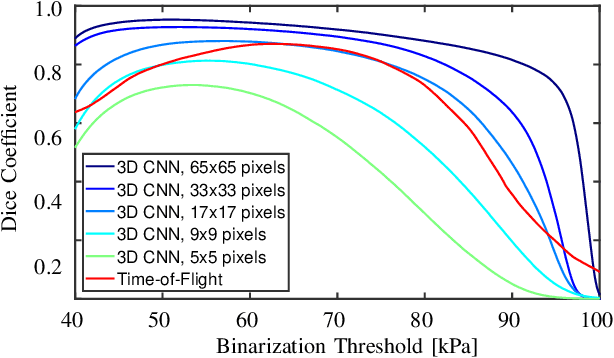
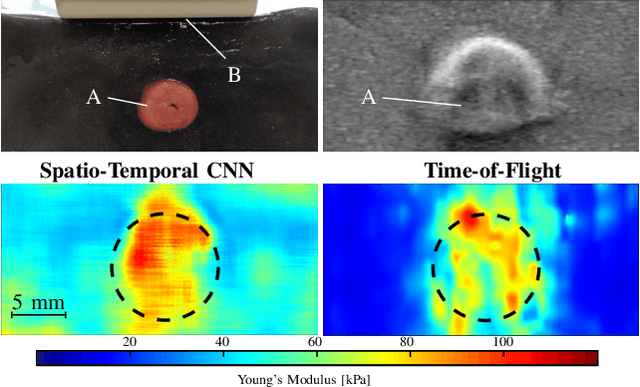
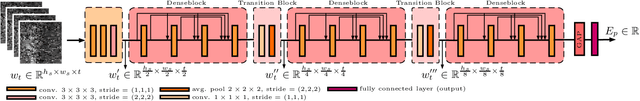
Abstract:Ultrasound shear wave elasticity imaging is a valuable tool for quantifying the elastic properties of tissue. Typically, the shear wave velocity is derived and mapped to an elasticity value, which neglects information such as the shape of the propagating shear wave or push sequence characteristics. We present 3D spatio-temporal CNNs for fast local elasticity estimation from ultrasound data. This approach is based on retrieving elastic properties from shear wave propagation within small local regions. A large training data set is acquired with a robot from homogeneous gelatin phantoms ranging from 17.42 kPa to 126.05 kPa with various push locations. The results show that our approach can estimate elastic properties on a pixelwise basis with a mean absolute error of 5.01+-4.37 kPa. Furthermore, we estimate local elasticity independent of the push location and can even perform accurate estimates inside the push region. For phantoms with embedded inclusions, we report a 53.93% lower MAE (7.50 kPa) and on the background of 85.24% (1.64 kPa) compared to a conventional shear wave method. Overall, our method offers fast local estimations of elastic properties with small spatio-temporal window sizes.
Unsupervised Anomaly Detection in 3D Brain MRI using Deep Learning with impured training data
Apr 12, 2022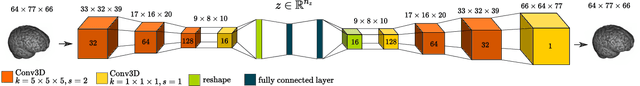
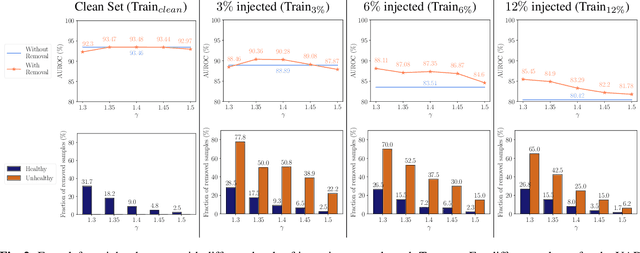
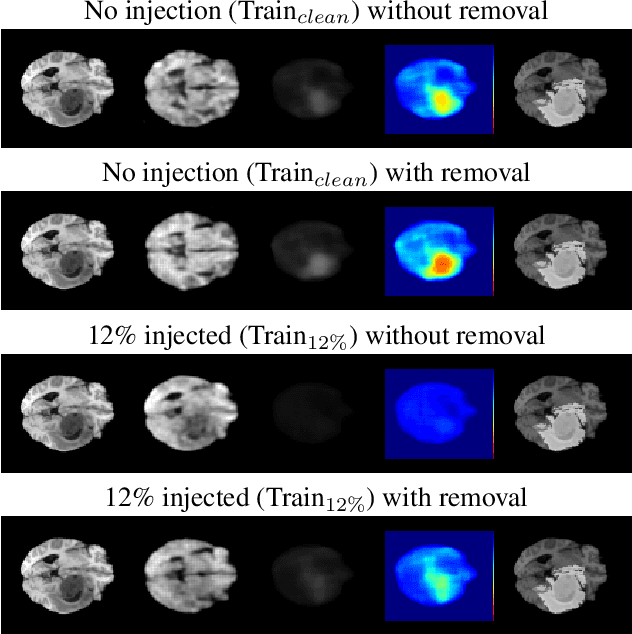
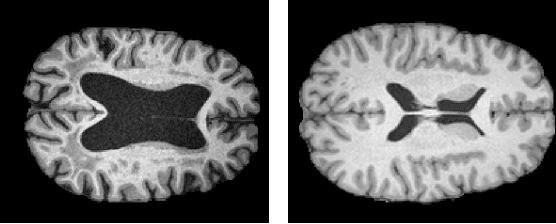
Abstract:The detection of lesions in magnetic resonance imaging (MRI)-scans of human brains remains challenging, time-consuming and error-prone. Recently, unsupervised anomaly detection (UAD) methods have shown promising results for this task. These methods rely on training data sets that solely contain healthy samples. Compared to supervised approaches, this significantly reduces the need for an extensive amount of labeled training data. However, data labelling remains error-prone. We study how unhealthy samples within the training data affect anomaly detection performance for brain MRI-scans. For our evaluations, we consider three publicly available data sets and use autoencoders (AE) as a well-established baseline method for UAD. We systematically evaluate the effect of impured training data by injecting different quantities of unhealthy samples to our training set of healthy samples from T1-weighted MRI-scans. We evaluate a method to identify falsely labeled samples directly during training based on the reconstruction error of the AE. Our results show that training with impured data decreases the UAD performance notably even with few falsely labeled samples. By performing outlier removal directly during training based on the reconstruction-loss, we demonstrate that falsely labeled data can be detected and removed to mitigate the effect of falsely labeled data. Overall, we highlight the importance of clean data sets for UAD in brain MRI and demonstrate an approach for detecting falsely labeled data directly during training.
Unsupervised Anomaly Detection in 3D Brain MRI using Deep Learning with Multi-Task Brain Age Prediction
Jan 31, 2022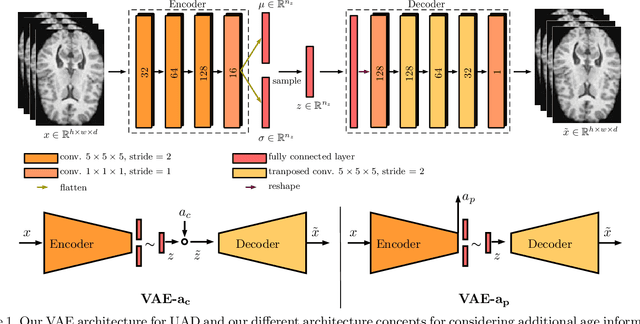

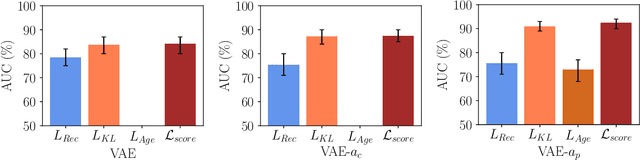
Abstract:Lesion detection in brain Magnetic Resonance Images (MRIs) remains a challenging task. MRIs are typically read and interpreted by domain experts, which is a tedious and time-consuming process. Recently, unsupervised anomaly detection (UAD) in brain MRI with deep learning has shown promising results to provide a quick, initial assessment. So far, these methods only rely on the visual appearance of healthy brain anatomy for anomaly detection. Another biomarker for abnormal brain development is the deviation between the brain age and the chronological age, which is unexplored in combination with UAD. We propose deep learning for UAD in 3D brain MRI considering additional age information. We analyze the value of age information during training, as an additional anomaly score, and systematically study several architecture concepts. Based on our analysis, we propose a novel deep learning approach for UAD with multi-task age prediction. We use clinical T1-weighted MRIs of 1735 healthy subjects and the publicly available BraTs 2019 data set for our study. Our novel approach significantly improves UAD performance with an AUC of 92.60% compared to an AUC-score of 84.37% using previous approaches without age information.
Multi-Scale Input Strategies for Medulloblastoma Tumor Classification using Deep Transfer Learning
Sep 14, 2021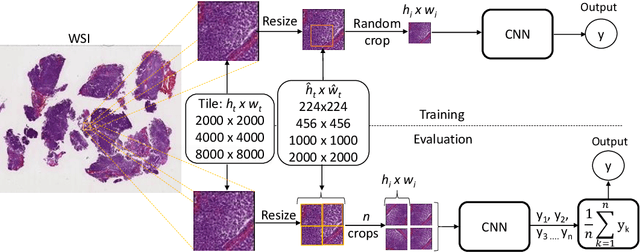
Abstract:Medulloblastoma (MB) is a primary central nervous system tumor and the most common malignant brain cancer among children. Neuropathologists perform microscopic inspection of histopathological tissue slides under a microscope to assess the severity of the tumor. This is a time-consuming task and often infused with observer variability. Recently, pre-trained convolutional neural networks (CNN) have shown promising results for MB subtype classification. Typically, high-resolution images are divided into smaller tiles for classification, while the size of the tiles has not been systematically evaluated. We study the impact of tile size and input strategy and classify the two major histopathological subtypes-Classic and Demoplastic/Nodular. To this end, we use recently proposed EfficientNets and evaluate tiles with increasing size combined with various downsampling scales. Our results demonstrate using large input tiles pixels followed by intermediate downsampling and patch cropping significantly improves MB classification performance. Our top-performing method achieves the AUC-ROC value of 90.90\% compared to 84.53\% using the previous approach with smaller input tiles.
3-Dimensional Deep Learning with Spatial Erasing for Unsupervised Anomaly Segmentation in Brain MRI
Sep 14, 2021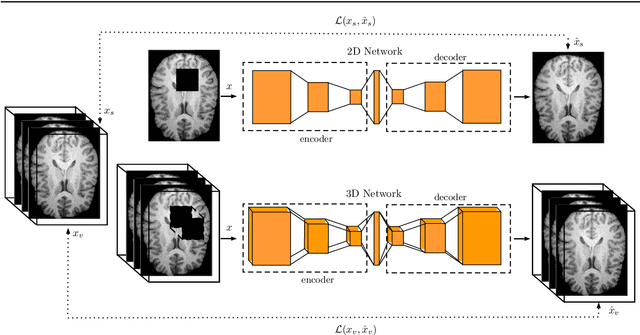
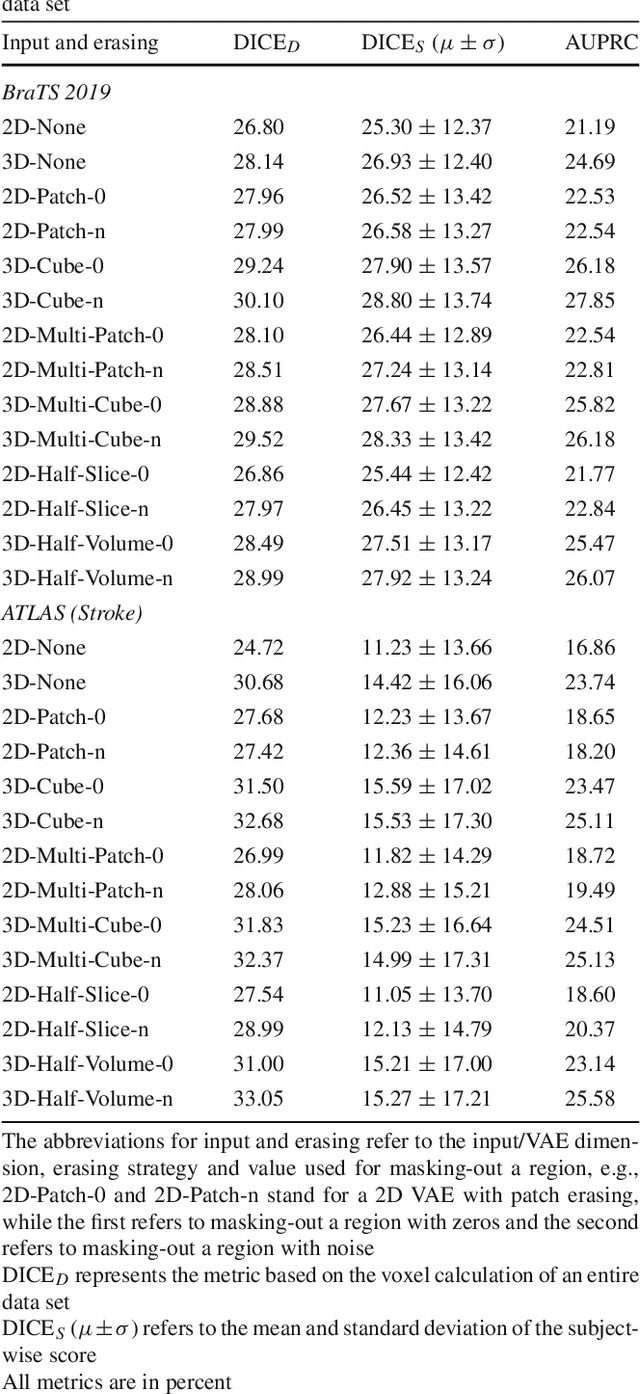
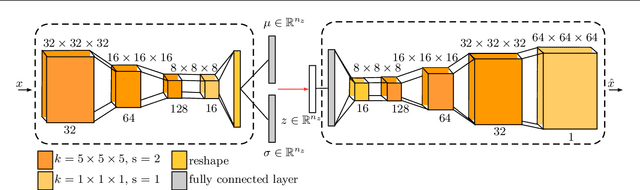
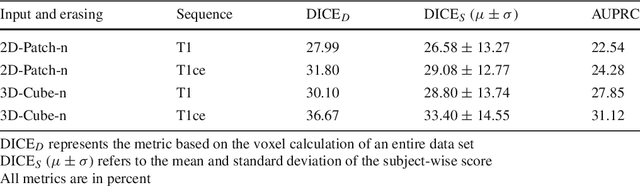
Abstract:Purpose. Brain Magnetic Resonance Images (MRIs) are essential for the diagnosis of neurological diseases. Recently, deep learning methods for unsupervised anomaly detection (UAD) have been proposed for the analysis of brain MRI. These methods rely on healthy brain MRIs and eliminate the requirement of pixel-wise annotated data compared to supervised deep learning. While a wide range of methods for UAD have been proposed, these methods are mostly 2D and only learn from MRI slices, disregarding that brain lesions are inherently 3D and the spatial context of MRI volumes remains unexploited. Methods. We investigate whether using increased spatial context by using MRI volumes combined with spatial erasing leads to improved unsupervised anomaly segmentation performance compared to learning from slices. We evaluate and compare 2D variational autoencoder (VAE) to their 3D counterpart, propose 3D input erasing, and systemically study the impact of the data set size on the performance. Results. Using two publicly available segmentation data sets for evaluation, 3D VAE outperform their 2D counterpart, highlighting the advantage of volumetric context. Also, our 3D erasing methods allow for further performance improvements. Our best performing 3D VAE with input erasing leads to an average DICE score of 31.40% compared to 25.76% for the 2D VAE. Conclusions. We propose 3D deep learning methods for UAD in brain MRI combined with 3D erasing and demonstrate that 3D methods clearly outperform their 2D counterpart for anomaly segmentation. Also, our spatial erasing method allows for further performance improvements and reduces the requirement for large data sets.
Medulloblastoma Tumor Classification using Deep Transfer Learning with Multi-Scale EfficientNets
Sep 10, 2021Abstract:Medulloblastoma (MB) is the most common malignant brain tumor in childhood. The diagnosis is generally based on the microscopic evaluation of histopathological tissue slides. However, visual-only assessment of histopathological patterns is a tedious and time-consuming task and is also affected by observer variability. Hence, automated MB tumor classification could assist pathologists by promoting consistency and robust quantification. Recently, convolutional neural networks (CNNs) have been proposed for this task, while transfer learning has shown promising results. In this work, we propose an end-to-end MB tumor classification and explore transfer learning with various input sizes and matching network dimensions. We focus on differentiating between the histological subtypes classic and desmoplastic/nodular. For this purpose, we systematically evaluate recently proposed EfficientNets, which uniformly scale all dimensions of a CNN. Using a data set with 161 cases, we demonstrate that pre-trained EfficientNets with larger input resolutions lead to significant performance improvements compared to commonly used pre-trained CNN architectures. Also, we highlight the importance of transfer learning, when using such large architectures. Overall, our best performing method achieves an F1-Score of 80.1%.
4D Spatio-Temporal Convolutional Networks for Object Position Estimation in OCT Volumes
Jul 02, 2020


Abstract:Tracking and localizing objects is a central problem in computer-assisted surgery. Optical coherence tomography (OCT) can be employed as an optical tracking system, due to its high spatial and temporal resolution. Recently, 3D convolutional neural networks (CNNs) have shown promising performance for pose estimation of a marker object using single volumetric OCT images. While this approach relied on spatial information only, OCT allows for a temporal stream of OCT image volumes capturing the motion of an object at high volumes rates. In this work, we systematically extend 3D CNNs to 4D spatio-temporal CNNs to evaluate the impact of additional temporal information for marker object tracking. Across various architectures, our results demonstrate that using a stream of OCT volumes and employing 4D spatio-temporal convolutions leads to a 30% lower mean absolute error compared to single volume processing with 3D CNNs.
Spectral-Spatial Recurrent-Convolutional Networks for In-Vivo Hyperspectral Tumor Type Classification
Jul 02, 2020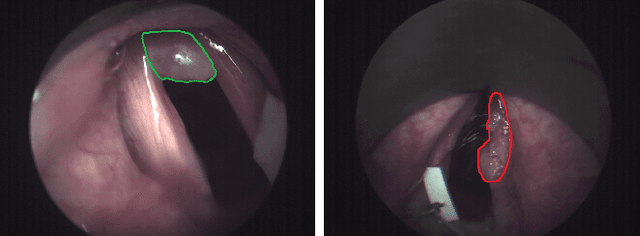
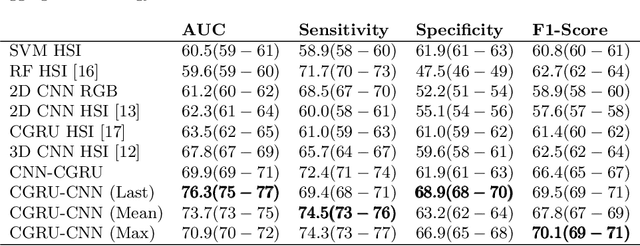
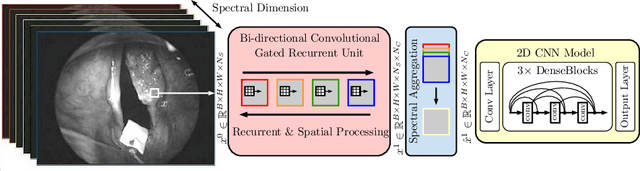
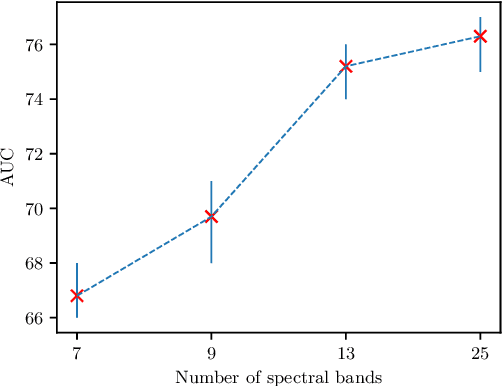
Abstract:Early detection of cancerous tissue is crucial for long-term patient survival. In the head and neck region, a typical diagnostic procedure is an endoscopic intervention where a medical expert manually assesses tissue using RGB camera images. While healthy and tumor regions are generally easier to distinguish, differentiating benign and malignant tumors is very challenging. This requires an invasive biopsy, followed by histological evaluation for diagnosis. Also, during tumor resection, tumor margins need to be verified by histological analysis. To avoid unnecessary tissue resection, a non-invasive, image-based diagnostic tool would be very valuable. Recently, hyperspectral imaging paired with deep learning has been proposed for this task, demonstrating promising results on ex-vivo specimens. In this work, we demonstrate the feasibility of in-vivo tumor type classification using hyperspectral imaging and deep learning. We analyze the value of using multiple hyperspectral bands compared to conventional RGB images and we study several machine learning models' ability to make use of the additional spectral information. Based on our insights, we address spectral and spatial processing using recurrent-convolutional models for effective spectral aggregating and spatial feature learning. Our best model achieves an AUC of 76.3%, significantly outperforming previous conventional and deep learning methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge