Debayan Bhattacharya
Leveraging the Mahalanobis Distance to enhance Unsupervised Brain MRI Anomaly Detection
Jul 17, 2024

Abstract:Unsupervised Anomaly Detection (UAD) methods rely on healthy data distributions to identify anomalies as outliers. In brain MRI, a common approach is reconstruction-based UAD, where generative models reconstruct healthy brain MRIs, and anomalies are detected as deviations between input and reconstruction. However, this method is sensitive to imperfect reconstructions, leading to false positives that impede the segmentation. To address this limitation, we construct multiple reconstructions with probabilistic diffusion models. We then analyze the resulting distribution of these reconstructions using the Mahalanobis distance to identify anomalies as outliers. By leveraging information about normal variations and covariance of individual pixels within this distribution, we effectively refine anomaly scoring, leading to improved segmentation. Our experimental results demonstrate substantial performance improvements across various data sets. Specifically, compared to relying solely on single reconstructions, our approach achieves relative improvements of 15.9%, 35.4%, 48.0%, and 4.7% in terms of AUPRC for the BRATS21, ATLAS, MSLUB and WMH data sets, respectively.
Self-supervised learning for classifying paranasal anomalies in the maxillary sinus
Apr 29, 2024Abstract:Purpose: Paranasal anomalies, frequently identified in routine radiological screenings, exhibit diverse morphological characteristics. Due to the diversity of anomalies, supervised learning methods require large labelled dataset exhibiting diverse anomaly morphology. Self-supervised learning (SSL) can be used to learn representations from unlabelled data. However, there are no SSL methods designed for the downstream task of classifying paranasal anomalies in the maxillary sinus (MS). Methods: Our approach uses a 3D Convolutional Autoencoder (CAE) trained in an unsupervised anomaly detection (UAD) framework. Initially, we train the 3D CAE to reduce reconstruction errors when reconstructing normal maxillary sinus (MS) image. Then, this CAE is applied to an unlabelled dataset to generate coarse anomaly locations by creating residual MS images. Following this, a 3D Convolutional Neural Network (CNN) reconstructs these residual images, which forms our SSL task. Lastly, we fine-tune the encoder part of the 3D CNN on a labelled dataset of normal and anomalous MS images. Results: The proposed SSL technique exhibits superior performance compared to existing generic self-supervised methods, especially in scenarios with limited annotated data. When trained on just 10% of the annotated dataset, our method achieves an Area Under the Precision-Recall Curve (AUPRC) of 0.79 for the downstream classification task. This performance surpasses other methods, with BYOL attaining an AUPRC of 0.75, SimSiam at 0.74, SimCLR at 0.73 and Masked Autoencoding using SparK at 0.75. Conclusion: A self-supervised learning approach that inherently focuses on localizing paranasal anomalies proves to be advantageous, particularly when the subsequent task involves differentiating normal from anomalous maxillary sinuses. Access our code at https://github.com/mtec-tuhh/self-supervised-paranasal-anomaly
Diffusion Models with Ensembled Structure-Based Anomaly Scoring for Unsupervised Anomaly Detection
Mar 21, 2024


Abstract:Supervised deep learning techniques show promise in medical image analysis. However, they require comprehensive annotated data sets, which poses challenges, particularly for rare diseases. Consequently, unsupervised anomaly detection (UAD) emerges as a viable alternative for pathology segmentation, as only healthy data is required for training. However, recent UAD anomaly scoring functions often focus on intensity only and neglect structural differences, which impedes the segmentation performance. This work investigates the potential of Structural Similarity (SSIM) to bridge this gap. SSIM captures both intensity and structural disparities and can be advantageous over the classical $l1$ error. However, we show that there is more than one optimal kernel size for the SSIM calculation for different pathologies. Therefore, we investigate an adaptive ensembling strategy for various kernel sizes to offer a more pathology-agnostic scoring mechanism. We demonstrate that this ensembling strategy can enhance the performance of DMs and mitigate the sensitivity to different kernel sizes across varying pathologies, highlighting its promise for brain MRI anomaly detection.
PolypNextLSTM: A lightweight and fast polyp video segmentation network using ConvNext and ConvLSTM
Feb 28, 2024Abstract:Commonly employed in polyp segmentation, single image UNet architectures lack the temporal insight clinicians gain from video data in diagnosing polyps. To mirror clinical practices more faithfully, our proposed solution, PolypNextLSTM, leverages video-based deep learning, harnessing temporal information for superior segmentation performance with the least parameter overhead, making it possibly suitable for edge devices. PolypNextLSTM employs a UNet-like structure with ConvNext-Tiny as its backbone, strategically omitting the last two layers to reduce parameter overhead. Our temporal fusion module, a Convolutional Long Short Term Memory (ConvLSTM), effectively exploits temporal features. Our primary novelty lies in PolypNextLSTM, which stands out as the leanest in parameters and the fastest model, surpassing the performance of five state-of-the-art image and video-based deep learning models. The evaluation of the SUN-SEG dataset spans easy-to-detect and hard-to-detect polyp scenarios, along with videos containing challenging artefacts like fast motion and occlusion. Comparison against 5 image-based and 5 video-based models demonstrates PolypNextLSTM's superiority, achieving a Dice score of 0.7898 on the hard-to-detect polyp test set, surpassing image-based PraNet (0.7519) and video-based PNSPlusNet (0.7486). Notably, our model excels in videos featuring complex artefacts such as ghosting and occlusion. PolypNextLSTM, integrating pruned ConvNext-Tiny with ConvLSTM for temporal fusion, not only exhibits superior segmentation performance but also maintains the highest frames per speed among evaluated models. Access code here https://github.com/mtec-tuhh/PolypNextLSTM
Guided Reconstruction with Conditioned Diffusion Models for Unsupervised Anomaly Detection in Brain MRIs
Dec 07, 2023



Abstract:Unsupervised anomaly detection in Brain MRIs aims to identify abnormalities as outliers from a healthy training distribution. Reconstruction-based approaches that use generative models to learn to reconstruct healthy brain anatomy are commonly used for this task. Diffusion models are an emerging class of deep generative models that show great potential regarding reconstruction fidelity. However, they face challenges in preserving intensity characteristics in the reconstructed images, limiting their performance in anomaly detection. To address this challenge, we propose to condition the denoising mechanism of diffusion models with additional information about the image to reconstruct coming from a latent representation of the noise-free input image. This conditioning enables high-fidelity reconstruction of healthy brain structures while aligning local intensity characteristics of input-reconstruction pairs. We evaluate our method's reconstruction quality, domain adaptation features and finally segmentation performance on publicly available data sets with various pathologies. Using our proposed conditioning mechanism we can reduce the false-positive predictions and enable a more precise delineation of anomalies which significantly enhances the anomaly detection performance compared to established state-of-the-art approaches to unsupervised anomaly detection in brain MRI. Furthermore, our approach shows promise in domain adaptation across different MRI acquisitions and simulated contrasts, a crucial property of general anomaly detection methods.
An objective validation of polyp and instrument segmentation methods in colonoscopy through Medico 2020 polyp segmentation and MedAI 2021 transparency challenges
Jul 30, 2023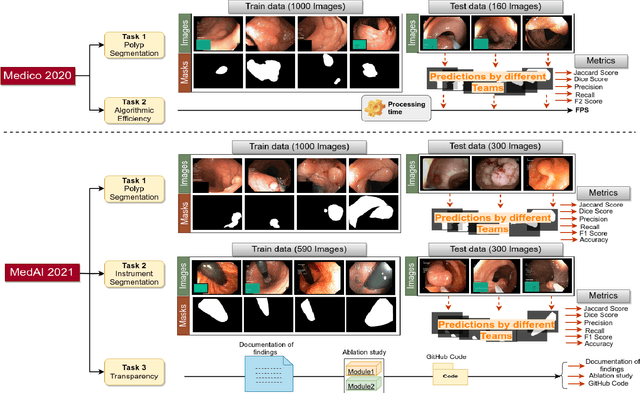
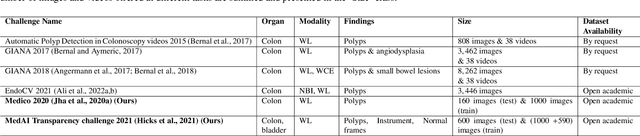
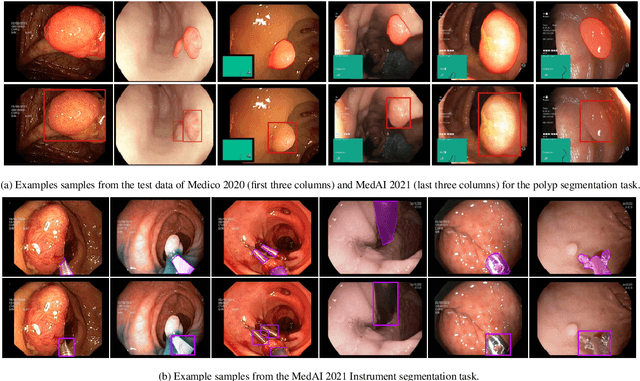
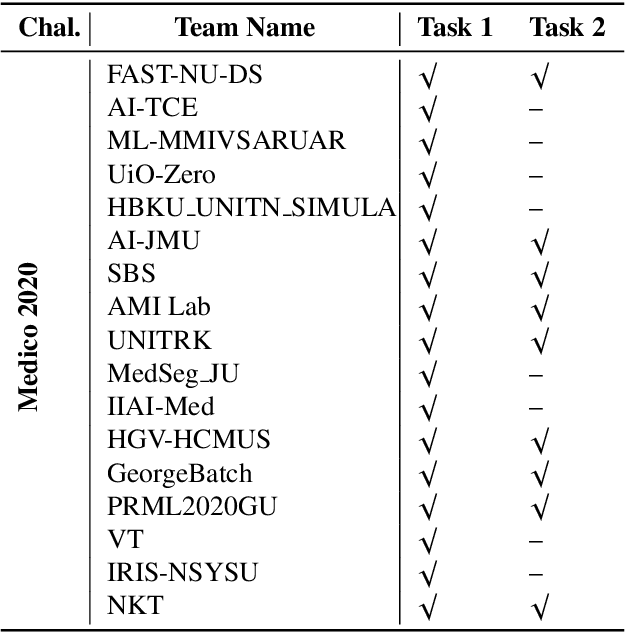
Abstract:Automatic analysis of colonoscopy images has been an active field of research motivated by the importance of early detection of precancerous polyps. However, detecting polyps during the live examination can be challenging due to various factors such as variation of skills and experience among the endoscopists, lack of attentiveness, and fatigue leading to a high polyp miss-rate. Deep learning has emerged as a promising solution to this challenge as it can assist endoscopists in detecting and classifying overlooked polyps and abnormalities in real time. In addition to the algorithm's accuracy, transparency and interpretability are crucial to explaining the whys and hows of the algorithm's prediction. Further, most algorithms are developed in private data, closed source, or proprietary software, and methods lack reproducibility. Therefore, to promote the development of efficient and transparent methods, we have organized the "Medico automatic polyp segmentation (Medico 2020)" and "MedAI: Transparency in Medical Image Segmentation (MedAI 2021)" competitions. We present a comprehensive summary and analyze each contribution, highlight the strength of the best-performing methods, and discuss the possibility of clinical translations of such methods into the clinic. For the transparency task, a multi-disciplinary team, including expert gastroenterologists, accessed each submission and evaluated the team based on open-source practices, failure case analysis, ablation studies, usability and understandability of evaluations to gain a deeper understanding of the models' credibility for clinical deployment. Through the comprehensive analysis of the challenge, we not only highlight the advancements in polyp and surgical instrument segmentation but also encourage qualitative evaluation for building more transparent and understandable AI-based colonoscopy systems.
Tissue Classification During Needle Insertion Using Self-Supervised Contrastive Learning and Optical Coherence Tomography
Apr 26, 2023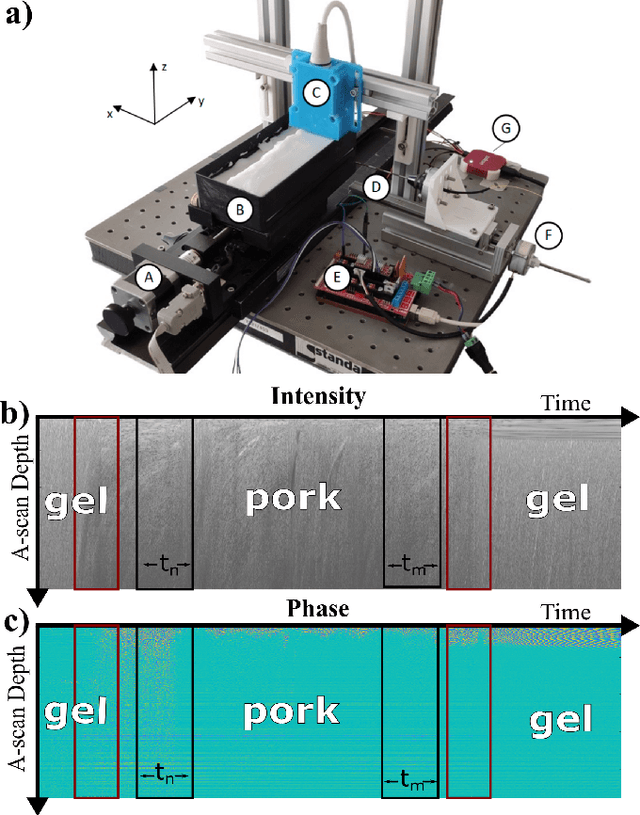
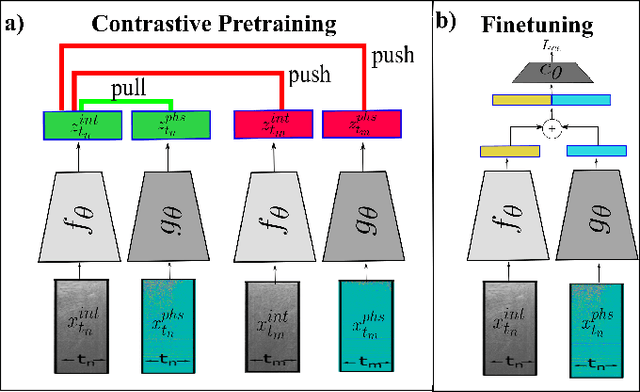
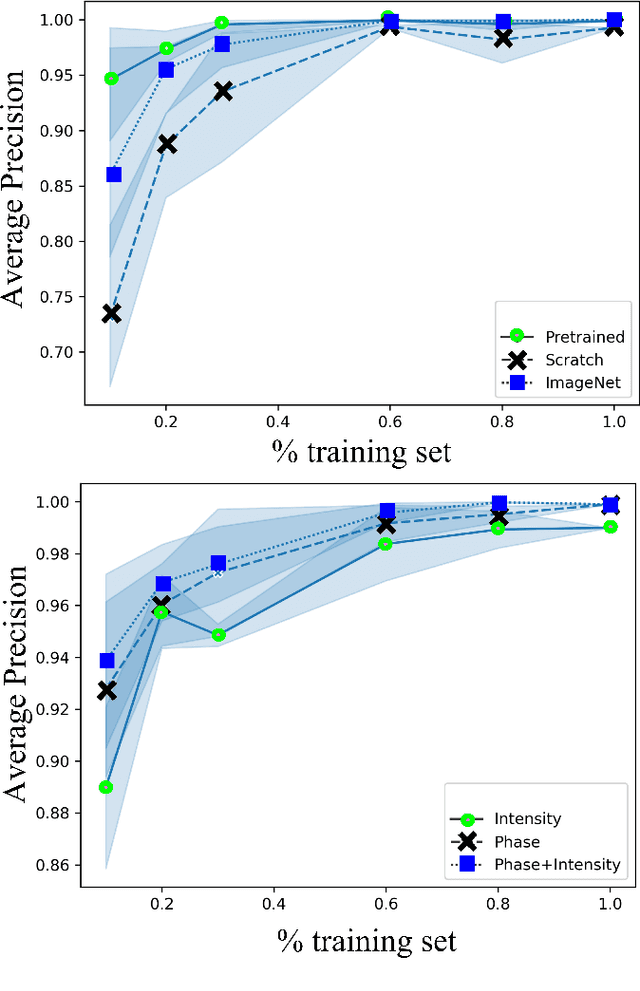
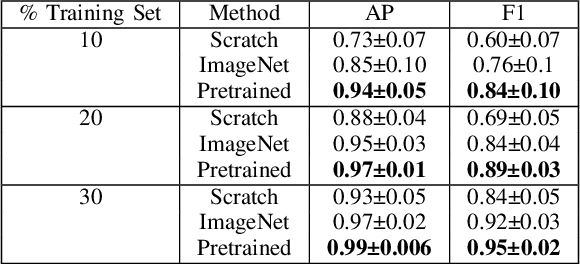
Abstract:Needle positioning is essential for various medical applications such as epidural anaesthesia. Physicians rely on their instincts while navigating the needle in epidural spaces. Thereby, identifying the tissue structures may be helpful to the physician as they can provide additional feedback in the needle insertion process. To this end, we propose a deep neural network that classifies the tissues from the phase and intensity data of complex OCT signals acquired at the needle tip. We investigate the performance of the deep neural network in a limited labelled dataset scenario and propose a novel contrastive pretraining strategy that learns invariant representation for phase and intensity data. We show that with 10% of the training set, our proposed pretraining strategy helps the model achieve an F1 score of 0.84 whereas the model achieves an F1 score of 0.60 without it. Further, we analyse the importance of phase and intensity individually towards tissue classification.
Multiple Instance Ensembling For Paranasal Anomaly Classification In The Maxillary Sinus
Mar 31, 2023Abstract:Paranasal anomalies are commonly discovered during routine radiological screenings and can present with a wide range of morphological features. This diversity can make it difficult for convolutional neural networks (CNNs) to accurately classify these anomalies, especially when working with limited datasets. Additionally, current approaches to paranasal anomaly classification are constrained to identifying a single anomaly at a time. These challenges necessitate the need for further research and development in this area. In this study, we investigate the feasibility of using a 3D convolutional neural network (CNN) to classify healthy maxillary sinuses (MS) and MS with polyps or cysts. The task of accurately identifying the relevant MS volume within larger head and neck Magnetic Resonance Imaging (MRI) scans can be difficult, but we develop a straightforward strategy to tackle this challenge. Our end-to-end solution includes the use of a novel sampling technique that not only effectively localizes the relevant MS volume, but also increases the size of the training dataset and improves classification results. Additionally, we employ a multiple instance ensemble prediction method to further boost classification performance. Finally, we identify the optimal size of MS volumes to achieve the highest possible classification performance on our dataset. With our multiple instance ensemble prediction strategy and sampling strategy, our 3D CNNs achieve an F1 of 0.85 whereas without it, they achieve an F1 of 0.70. We demonstrate the feasibility of classifying anomalies in the MS. We propose a data enlarging strategy alongside a novel ensembling strategy that proves to be beneficial for paranasal anomaly classification in the MS.
Patched Diffusion Models for Unsupervised Anomaly Detection in Brain MRI
Mar 07, 2023



Abstract:The use of supervised deep learning techniques to detect pathologies in brain MRI scans can be challenging due to the diversity of brain anatomy and the need for annotated data sets. An alternative approach is to use unsupervised anomaly detection, which only requires sample-level labels of healthy brains to create a reference representation. This reference representation can then be compared to unhealthy brain anatomy in a pixel-wise manner to identify abnormalities. To accomplish this, generative models are needed to create anatomically consistent MRI scans of healthy brains. While recent diffusion models have shown promise in this task, accurately generating the complex structure of the human brain remains a challenge. In this paper, we propose a method that reformulates the generation task of diffusion models as a patch-based estimation of healthy brain anatomy, using spatial context to guide and improve reconstruction. We evaluate our approach on data of tumors and multiple sclerosis lesions and demonstrate a relative improvement of 25.1% compared to existing baselines.
Unsupervised Anomaly Detection of Paranasal Anomalies in the Maxillary Sinus
Nov 01, 2022Abstract:Deep learning (DL) algorithms can be used to automate paranasal anomaly detection from Magnetic Resonance Imaging (MRI). However, previous works relied on supervised learning techniques to distinguish between normal and abnormal samples. This method limits the type of anomalies that can be classified as the anomalies need to be present in the training data. Further, many data points from normal and anomaly class are needed for the model to achieve satisfactory classification performance. However, experienced clinicians can segregate between normal samples (healthy maxillary sinus) and anomalous samples (anomalous maxillary sinus) after looking at a few normal samples. We mimic the clinicians ability by learning the distribution of healthy maxillary sinuses using a 3D convolutional auto-encoder (cAE) and its variant, a 3D variational autoencoder (VAE) architecture and evaluate cAE and VAE for this task. Concretely, we pose the paranasal anomaly detection as an unsupervised anomaly detection problem. Thereby, we are able to reduce the labelling effort of the clinicians as we only use healthy samples during training. Additionally, we can classify any type of anomaly that differs from the training distribution. We train our 3D cAE and VAE to learn a latent representation of healthy maxillary sinus volumes using L1 reconstruction loss. During inference, we use the reconstruction error to classify between normal and anomalous maxillary sinuses. We extract sub-volumes from larger head and neck MRIs and analyse the effect of different fields of view on the detection performance. Finally, we report which anomalies are easiest and hardest to classify using our approach. Our results demonstrate the feasibility of unsupervised detection of paranasal anomalies from MRIs with an AUPRC of 85% and 80% for cAE and VAE, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge