Gabriel Efrain Humpire Mamani
Kidney abnormality segmentation in thorax-abdomen CT scans
Sep 06, 2023


Abstract:In this study, we introduce a deep learning approach for segmenting kidney parenchyma and kidney abnormalities to support clinicians in identifying and quantifying renal abnormalities such as cysts, lesions, masses, metastases, and primary tumors. Our end-to-end segmentation method was trained on 215 contrast-enhanced thoracic-abdominal CT scans, with half of these scans containing one or more abnormalities. We began by implementing our own version of the original 3D U-Net network and incorporated four additional components: an end-to-end multi-resolution approach, a set of task-specific data augmentations, a modified loss function using top-$k$, and spatial dropout. Furthermore, we devised a tailored post-processing strategy. Ablation studies demonstrated that each of the four modifications enhanced kidney abnormality segmentation performance, while three out of four improved kidney parenchyma segmentation. Subsequently, we trained the nnUNet framework on our dataset. By ensembling the optimized 3D U-Net and the nnUNet with our specialized post-processing, we achieved marginally superior results. Our best-performing model attained Dice scores of 0.965 and 0.947 for segmenting kidney parenchyma in two test sets (20 scans without abnormalities and 30 with abnormalities), outperforming an independent human observer who scored 0.944 and 0.925, respectively. In segmenting kidney abnormalities within the 30 test scans containing them, the top-performing method achieved a Dice score of 0.585, while an independent second human observer reached a score of 0.664, suggesting potential for further improvement in computerized methods. All training data is available to the research community under a CC-BY 4.0 license on https://doi.org/10.5281/zenodo.8014289
The Liver Tumor Segmentation Benchmark (LiTS)
Jan 13, 2019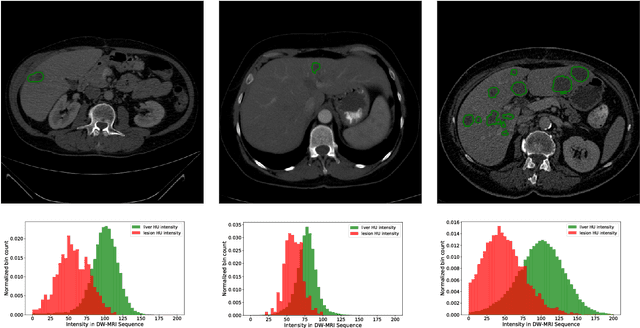
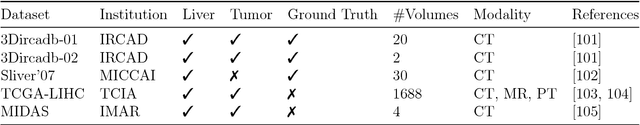

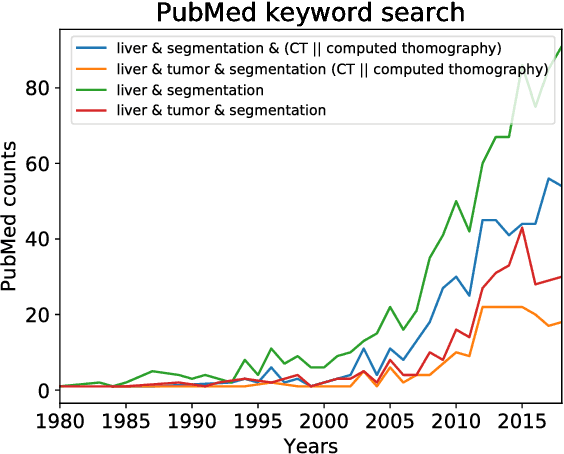
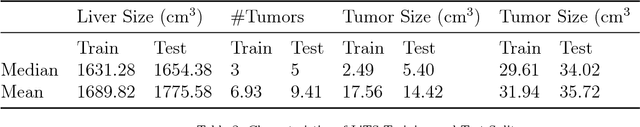
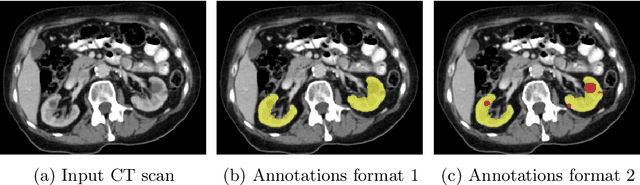
Abstract:In this work, we report the set-up and results of the Liver Tumor Segmentation Benchmark (LITS) organized in conjunction with the IEEE International Symposium on Biomedical Imaging (ISBI) 2016 and International Conference On Medical Image Computing Computer Assisted Intervention (MICCAI) 2017. Twenty four valid state-of-the-art liver and liver tumor segmentation algorithms were applied to a set of 131 computed tomography (CT) volumes with different types of tumor contrast levels (hyper-/hypo-intense), abnormalities in tissues (metastasectomie) size and varying amount of lesions. The submitted algorithms have been tested on 70 undisclosed volumes. The dataset is created in collaboration with seven hospitals and research institutions and manually reviewed by independent three radiologists. We found that not a single algorithm performed best for liver and tumors. The best liver segmentation algorithm achieved a Dice score of 0.96(MICCAI) whereas for tumor segmentation the best algorithm evaluated at 0.67(ISBI) and 0.70(MICCAI). The LITS image data and manual annotations continue to be publicly available through an online evaluation system as an ongoing benchmarking resource.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge