Eugene Vorontsov
Mixed Magnification Aggregation for Generalizable Region-Level Representations in Computational Pathology
Feb 25, 2026Abstract:In recent years, a standard computational pathology workflow has emerged where whole slide images are cropped into tiles, these tiles are processed using a foundation model, and task-specific models are built using the resulting representations. At least 15 different foundation models have been proposed, and the vast majority are trained exclusively with tiles using the 20$\times$ magnification. However, it is well known that certain histologic features can only be discerned with larger context windows and requires a pathologist to zoom in and out when analyzing a whole slide image. Furthermore, creating 224$\times$224 pixel crops at 20$\times$ leads to a large number of tiles per slide, which can be gigapixel in size. To more accurately capture multi-resolution features and investigate the possibility of reducing the number of representations per slide, we propose a region-level mixing encoder. Our approach jointly fuses image tile representations of a mixed magnification foundation model using a masked embedding modeling pretraining step. We explore a design space for pretraining the proposed mixed-magnification region aggregators and evaluate our models on transfer to biomarker prediction tasks representing various cancer types. Results demonstrate cancer dependent improvements in predictive performance, highlighting the importance of spatial context and understanding.
PRISM2: Unlocking Multi-Modal General Pathology AI with Clinical Dialogue
Jun 16, 2025Abstract:Recent pathology foundation models can provide rich tile-level representations but fall short of delivering general-purpose clinical utility without further extensive model development. These models lack whole-slide image (WSI) understanding and are not trained with large-scale diagnostic data, limiting their performance on diverse downstream tasks. We introduce PRISM2, a multi-modal slide-level foundation model trained via clinical dialogue to enable scalable, generalizable pathology AI. PRISM2 is trained on nearly 700,000 specimens (2.3 million WSIs) paired with real-world clinical diagnostic reports in a two-stage process. In Stage 1, a vision-language model is trained using contrastive and captioning objectives to align whole slide embeddings with textual clinical diagnosis. In Stage 2, the language model is unfrozen to enable diagnostic conversation and extract more clinically meaningful representations from hidden states. PRISM2 achieves strong performance on diagnostic and biomarker prediction tasks, outperforming prior slide-level models including PRISM and TITAN. It also introduces a zero-shot yes/no classification approach that surpasses CLIP-style methods without prompt tuning or class enumeration. By aligning visual features with clinical reasoning, PRISM2 improves generalization on both data-rich and low-sample tasks, offering a scalable path forward for building general pathology AI agents capable of assisting diagnostic and prognostic decisions.
Virchow2: Scaling Self-Supervised Mixed Magnification Models in Pathology
Aug 14, 2024Abstract:Foundation models are rapidly being developed for computational pathology applications. However, it remains an open question which factors are most important for downstream performance with data scale and diversity, model size, and training algorithm all playing a role. In this work, we propose algorithmic modifications, tailored for pathology, and we present the result of scaling both data and model size, surpassing previous studies in both dimensions. We introduce two new models: Virchow2, a 632 million parameter vision transformer, and Virchow2G, a 1.9 billion parameter vision transformer, each trained with 3.1 million histopathology whole slide images, with diverse tissues, originating institutions, and stains. We achieve state of the art performance on 12 tile-level tasks, as compared to the top performing competing models. Our results suggest that data diversity and domain-specific methods can outperform models that only scale in the number of parameters, but, on average, performance benefits from the combination of domain-specific methods, data scale, and model scale.
Virchow 2: Scaling Self-Supervised Mixed Magnification Models in Pathology
Aug 01, 2024Abstract:Foundation models are rapidly being developed for computational pathology applications. However, it remains an open question which factors are most important for downstream performance with data scale and diversity, model size, and training algorithm all playing a role. In this work, we present the result of scaling both data and model size, surpassing previous studies in both dimensions, and introduce two new models: Virchow 2, a 632M parameter vision transformer, and Virchow 2G, a 1.85B parameter vision transformer, each trained with 3.1M histopathology whole slide images. To support this scale, we propose domain-inspired adaptations to the DINOv2 training algorithm, which is quickly becoming the default method in self-supervised learning for computational pathology. We achieve state of the art performance on twelve tile-level tasks, as compared to the top performing competing models. Our results suggest that data diversity and domain-specific training can outperform models that only scale in the number of parameters, but, on average, performance benefits from domain-tailoring, data scale, and model scale.
PRISM: A Multi-Modal Generative Foundation Model for Slide-Level Histopathology
May 16, 2024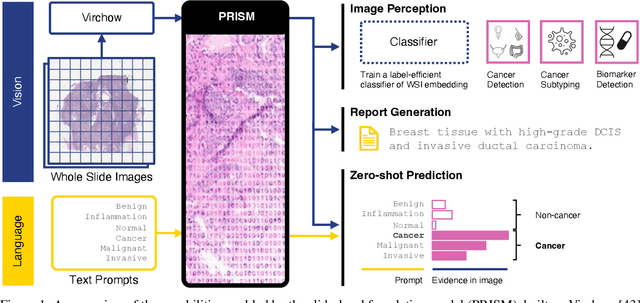
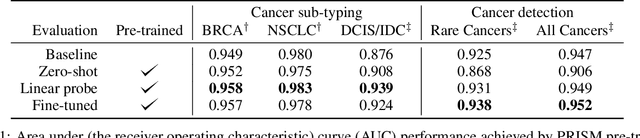
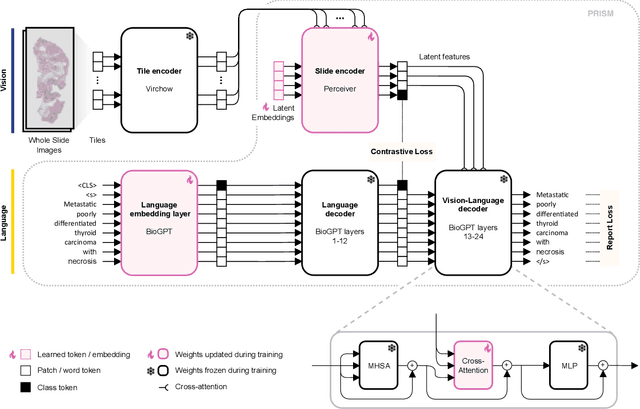
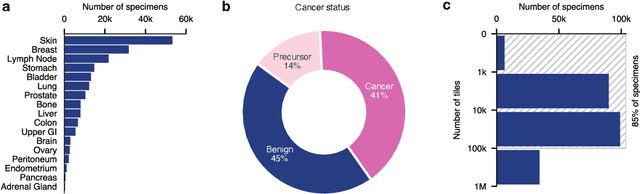
Abstract:Foundation models in computational pathology promise to unlock the development of new clinical decision support systems and models for precision medicine. However, there is a mismatch between most clinical analysis, which is defined at the level of one or more whole slide images, and foundation models to date, which process the thousands of image tiles contained in a whole slide image separately. The requirement to train a network to aggregate information across a large number of tiles in multiple whole slide images limits these models' impact. In this work, we present a slide-level foundation model for H&E-stained histopathology, PRISM, that builds on Virchow tile embeddings and leverages clinical report text for pre-training. Using the tile embeddings, PRISM produces slide-level embeddings with the ability to generate clinical reports, resulting in several modes of use. Using text prompts, PRISM achieves zero-shot cancer detection and sub-typing performance approaching and surpassing that of a supervised aggregator model. Using the slide embeddings with linear classifiers, PRISM surpasses supervised aggregator models. Furthermore, we demonstrate that fine-tuning of the PRISM slide encoder yields label-efficient training for biomarker prediction, a task that typically suffers from low availability of training data; an aggregator initialized with PRISM and trained on as little as 10% of the training data can outperform a supervised baseline that uses all of the data.
Adapting Self-Supervised Learning for Computational Pathology
May 02, 2024Abstract:Self-supervised learning (SSL) has emerged as a key technique for training networks that can generalize well to diverse tasks without task-specific supervision. This property makes SSL desirable for computational pathology, the study of digitized images of tissues, as there are many target applications and often limited labeled training samples. However, SSL algorithms and models have been primarily developed in the field of natural images and whether their performance can be improved by adaptation to particular domains remains an open question. In this work, we present an investigation of modifications to SSL for pathology data, specifically focusing on the DINOv2 algorithm. We propose alternative augmentations, regularization functions, and position encodings motivated by the characteristics of pathology images. We evaluate the impact of these changes on several benchmarks to demonstrate the value of tailored approaches.
Virchow: A Million-Slide Digital Pathology Foundation Model
Sep 21, 2023Abstract:Computational pathology uses artificial intelligence to enable precision medicine and decision support systems through the analysis of whole slide images. It has the potential to revolutionize the diagnosis and treatment of cancer. However, a major challenge to this objective is that for many specific computational pathology tasks the amount of data is inadequate for development. To address this challenge, we created Virchow, a 632 million parameter deep neural network foundation model for computational pathology. Using self-supervised learning, Virchow is trained on 1.5 million hematoxylin and eosin stained whole slide images from diverse tissue groups, which is orders of magnitude more data than previous works. When evaluated on downstream tasks including tile-level pan-cancer detection and subtyping and slide-level biomarker prediction, Virchow outperforms state-of-the-art systems both on internal datasets drawn from the same population as the pretraining data as well as external public datasets. Virchow achieves 93% balanced accuracy for pancancer tile classification, and AUCs of 0.983 for colon microsatellite instability status prediction and 0.967 for breast CDH1 status prediction. The gains in performance highlight the importance of pretraining on massive pathology image datasets, suggesting pretraining on even larger datasets could continue improving performance for many high-impact applications where limited amounts of training data are available, such as drug outcome prediction.
Image-level supervision and self-training for transformer-based cross-modality tumor segmentation
Sep 17, 2023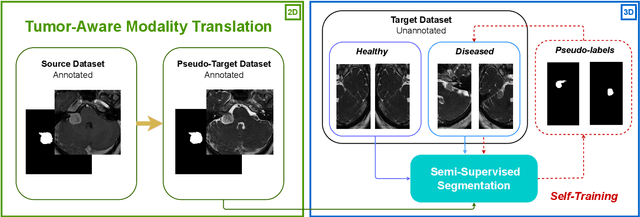
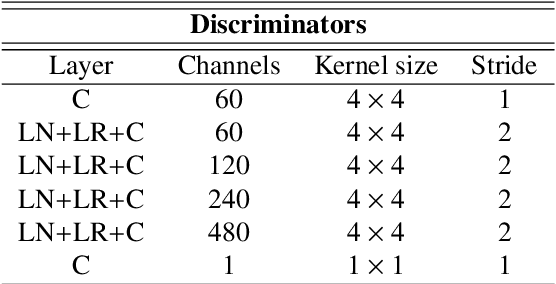
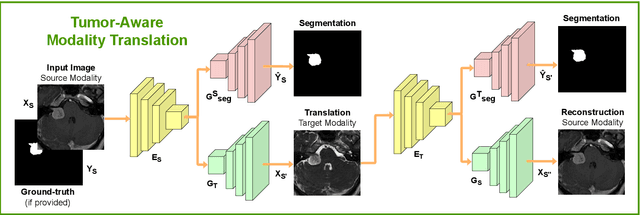
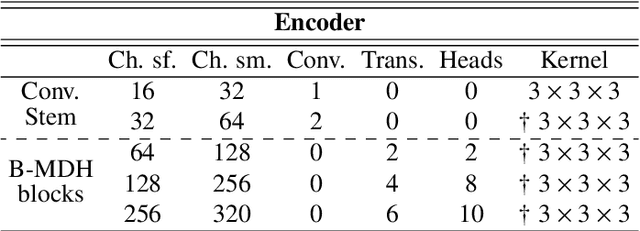
Abstract:Deep neural networks are commonly used for automated medical image segmentation, but models will frequently struggle to generalize well across different imaging modalities. This issue is particularly problematic due to the limited availability of annotated data, making it difficult to deploy these models on a larger scale. To overcome these challenges, we propose a new semi-supervised training strategy called MoDATTS. Our approach is designed for accurate cross-modality 3D tumor segmentation on unpaired bi-modal datasets. An image-to-image translation strategy between imaging modalities is used to produce annotated pseudo-target volumes and improve generalization to the unannotated target modality. We also use powerful vision transformer architectures and introduce an iterative self-training procedure to further close the domain gap between modalities. MoDATTS additionally allows the possibility to extend the training to unannotated target data by exploiting image-level labels with an unsupervised objective that encourages the model to perform 3D diseased-to-healthy translation by disentangling tumors from the background. The proposed model achieves superior performance compared to other methods from participating teams in the CrossMoDA 2022 challenge, as evidenced by its reported top Dice score of 0.87+/-0.04 for the VS segmentation. MoDATTS also yields consistent improvements in Dice scores over baselines on a cross-modality brain tumor segmentation task composed of four different contrasts from the BraTS 2020 challenge dataset, where 95% of a target supervised model performance is reached. We report that 99% and 100% of this maximum performance can be attained if 20% and 50% of the target data is additionally annotated, which further demonstrates that MoDATTS can be leveraged to reduce the annotation burden.
M-GenSeg: Domain Adaptation For Target Modality Tumor Segmentation With Annotation-Efficient Supervision
Dec 14, 2022

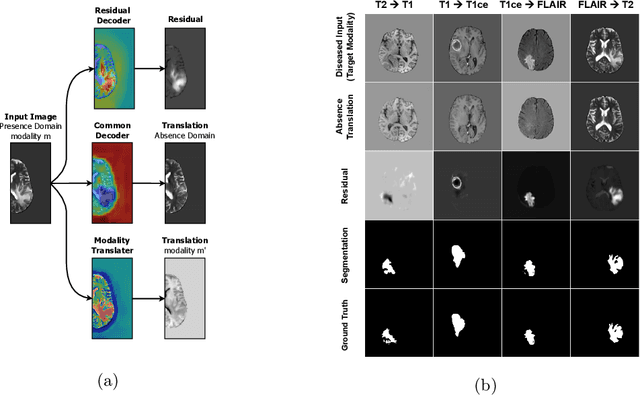
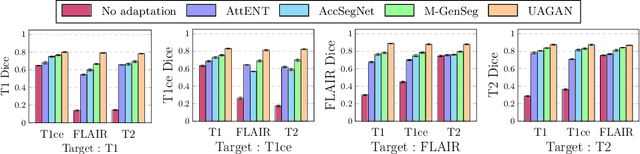
Abstract:Automated medical image segmentation using deep neural networks typically requires substantial supervised training. However, these models fail to generalize well across different imaging modalities. This shortcoming, amplified by the limited availability of annotated data, has been hampering the deployment of such methods at a larger scale across modalities. To address these issues, we propose M-GenSeg, a new semi-supervised training strategy for accurate cross-modality tumor segmentation on unpaired bi-modal datasets. Based on image-level labels, a first unsupervised objective encourages the model to perform diseased to healthy translation by disentangling tumors from the background, which encompasses the segmentation task. Then, teaching the model to translate between image modalities enables the synthesis of target images from a source modality, thus leveraging the pixel-level annotations from the source modality to enforce generalization to the target modality images. We evaluated the performance on a brain tumor segmentation datasets composed of four different contrast sequences from the public BraTS 2020 challenge dataset. We report consistent improvement in Dice scores on both source and unannotated target modalities. On all twelve distinct domain adaptation experiments, the proposed model shows a clear improvement over state-of-the-art domain-adaptive baselines, with absolute Dice gains on the target modality reaching 0.15.
Label noise in segmentation networks : mitigation must deal with bias
Jul 05, 2021
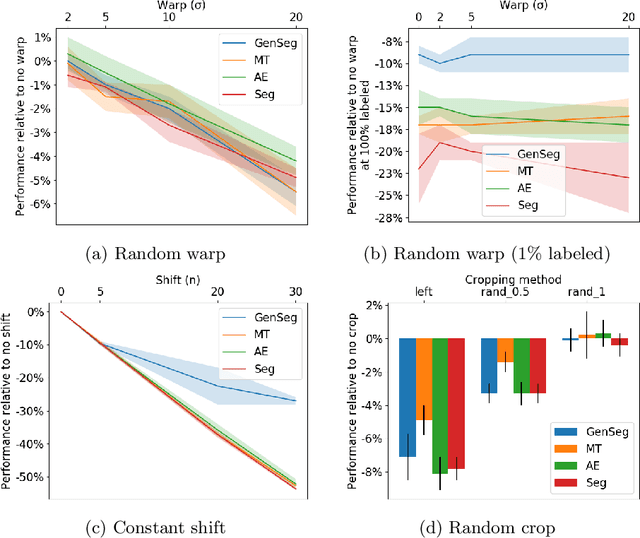

Abstract:Imperfect labels limit the quality of predictions learned by deep neural networks. This is particularly relevant in medical image segmentation, where reference annotations are difficult to collect and vary significantly even across expert annotators. Prior work on mitigating label noise focused on simple models of mostly uniform noise. In this work, we explore biased and unbiased errors artificially introduced to brain tumour annotations on MRI data. We found that supervised and semi-supervised segmentation methods are robust or fairly robust to unbiased errors but sensitive to biased errors. It is therefore important to identify the sorts of errors expected in medical image labels and especially mitigate the biased errors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge