Fausto Milletari
Adapting Self-Supervised Learning for Computational Pathology
May 02, 2024Abstract:Self-supervised learning (SSL) has emerged as a key technique for training networks that can generalize well to diverse tasks without task-specific supervision. This property makes SSL desirable for computational pathology, the study of digitized images of tissues, as there are many target applications and often limited labeled training samples. However, SSL algorithms and models have been primarily developed in the field of natural images and whether their performance can be improved by adaptation to particular domains remains an open question. In this work, we present an investigation of modifications to SSL for pathology data, specifically focusing on the DINOv2 algorithm. We propose alternative augmentations, regularization functions, and position encodings motivated by the characteristics of pathology images. We evaluate the impact of these changes on several benchmarks to demonstrate the value of tailored approaches.
Searching Learning Strategy with Reinforcement Learning for 3D Medical Image Segmentation
Jun 10, 2020
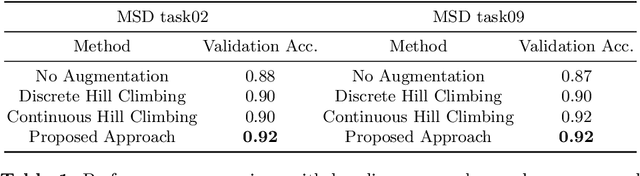

Abstract:Deep neural network (DNN) based approaches have been widely investigated and deployed in medical image analysis. For example, fully convolutional neural networks (FCN) achieve the state-of-the-art performance in several applications of 2D/3D medical image segmentation. Even the baseline neural network models (U-Net, V-Net, etc.) have been proven to be very effective and efficient when the training process is set up properly. Nevertheless, to fully exploit the potentials of neural networks, we propose an automated searching approach for the optimal training strategy with reinforcement learning. The proposed approach can be utilized for tuning hyper-parameters, and selecting necessary data augmentation with certain probabilities. The proposed approach is validated on several tasks of 3D medical image segmentation. The performance of the baseline model is boosted after searching, and it can achieve comparable accuracy to other manually-tuned state-of-the-art segmentation approaches.
* 9 pages, 1 figures
The Future of Digital Health with Federated Learning
Mar 18, 2020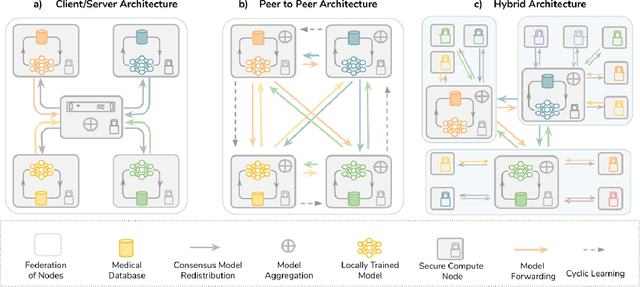
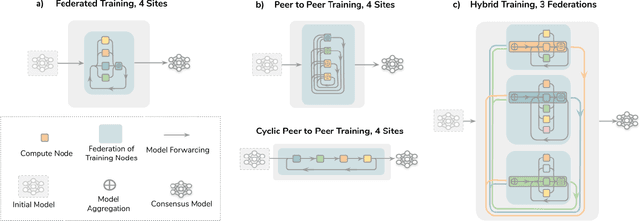
Abstract:Data-driven Machine Learning has emerged as a promising approach for building accurate and robust statistical models from medical data, which is collected in huge volumes by modern healthcare systems. Existing medical data is not fully exploited by ML primarily because it sits in data silos and privacy concerns restrict access to this data. However, without access to sufficient data, ML will be prevented from reaching its full potential and, ultimately, from making the transition from research to clinical practice. This paper considers key factors contributing to this issue, explores how Federated Learning (FL) may provide a solution for the future of digital health and highlights the challenges and considerations that need to be addressed.
NeurReg: Neural Registration and Its Application to Image Segmentation
Oct 04, 2019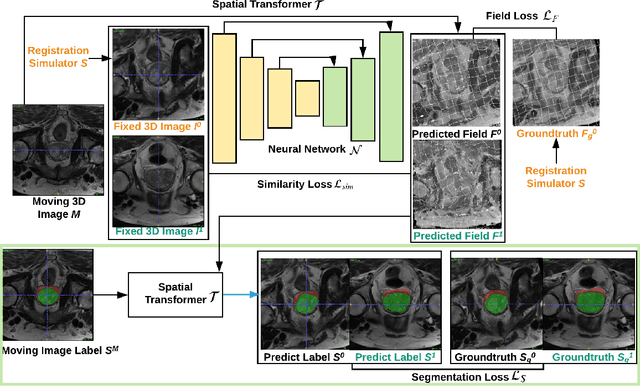

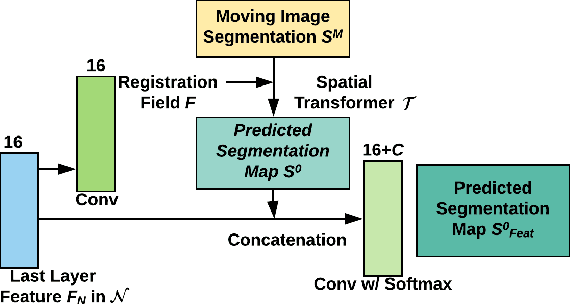
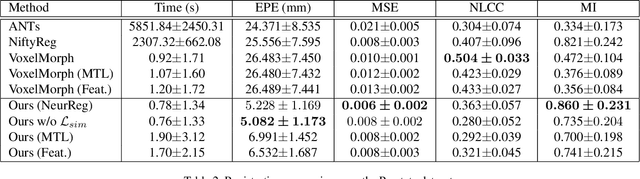
Abstract:Registration is a fundamental task in medical image analysis which can be applied to several tasks including image segmentation, intra-operative tracking, multi-modal image alignment, and motion analysis. Popular registration tools such as ANTs and NiftyReg optimize an objective function for each pair of images from scratch which is time-consuming for large images with complicated deformation. Facilitated by the rapid progress of deep learning, learning-based approaches such as VoxelMorph have been emerging for image registration. These approaches can achieve competitive performance in a fraction of a second on advanced GPUs. In this work, we construct a neural registration framework, called NeurReg, with a hybrid loss of displacement fields and data similarity, which substantially improves the current state-of-the-art of registrations. Within the framework, we simulate various transformations by a registration simulator which generates fixed image and displacement field ground truth for training. Furthermore, we design three segmentation frameworks based on the proposed registration framework: 1) atlas-based segmentation, 2) joint learning of both segmentation and registration tasks, and 3) multi-task learning with atlas-based segmentation as an intermediate feature. Extensive experimental results validate the effectiveness of the proposed NeurReg framework based on various metrics: the endpoint error (EPE) of the predicted displacement field, mean square error (MSE), normalized local cross-correlation (NLCC), mutual information (MI), Dice coefficient, uncertainty estimation, and the interpretability of the segmentation. The proposed NeurReg improves registration accuracy with fast inference speed, which can greatly accelerate related medical image analysis tasks.
Weakly supervised segmentation from extreme points
Oct 02, 2019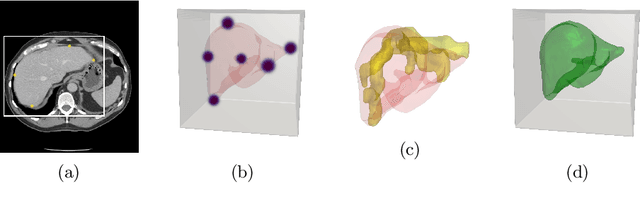
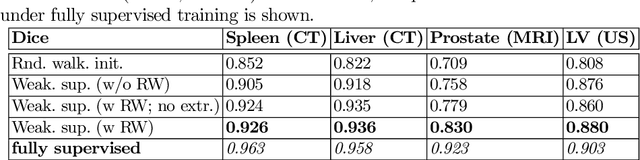

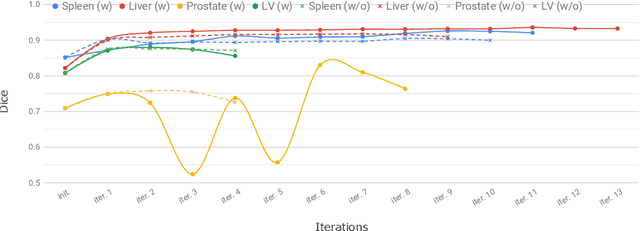
Abstract:Annotation of medical images has been a major bottleneck for the development of accurate and robust machine learning models. Annotation is costly and time-consuming and typically requires expert knowledge, especially in the medical domain. Here, we propose to use minimal user interaction in the form of extreme point clicks in order to train a segmentation model that can, in turn, be used to speed up the annotation of medical images. We use extreme points in each dimension of a 3D medical image to constrain an initial segmentation based on the random walker algorithm. This segmentation is then used as a weak supervisory signal to train a fully convolutional network that can segment the organ of interest based on the provided user clicks. We show that the network's predictions can be refined through several iterations of training and prediction using the same weakly annotated data. Ultimately, our method has the potential to speed up the generation process of new training datasets for the development of new machine learning and deep learning-based models for, but not exclusively, medical image analysis.
Correlation via synthesis: end-to-end nodule image generation and radiogenomic map learning based on generative adversarial network
Jul 08, 2019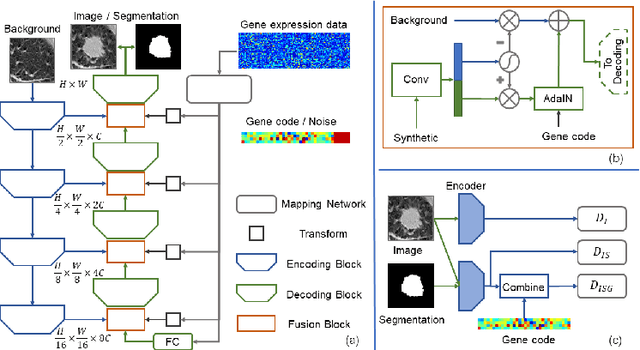

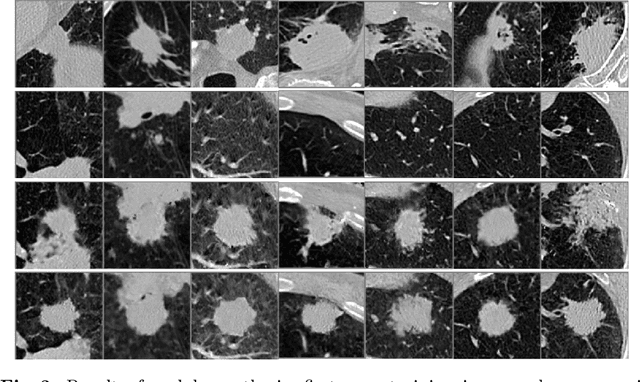
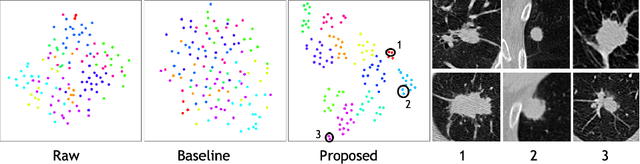
Abstract:Radiogenomic map linking image features and gene expression profiles is useful for noninvasively identifying molecular properties of a particular type of disease. Conventionally, such map is produced in three separate steps: 1) gene-clustering to "metagenes", 2) image feature extraction, and 3) statistical correlation between metagenes and image features. Each step is independently performed and relies on arbitrary measurements. In this work, we investigate the potential of an end-to-end method fusing gene data with image features to generate synthetic image and learn radiogenomic map simultaneously. To achieve this goal, we develop a generative adversarial network (GAN) conditioned on both background images and gene expression profiles, synthesizing the corresponding image. Image and gene features are fused at different scales to ensure the realism and quality of the synthesized image. We tested our method on non-small cell lung cancer (NSCLC) dataset. Results demonstrate that the proposed method produces realistic synthetic images, and provides a promising way to find gene-image relationship in a holistic end-to-end manner.
Interactive segmentation of medical images through fully convolutional neural networks
Mar 19, 2019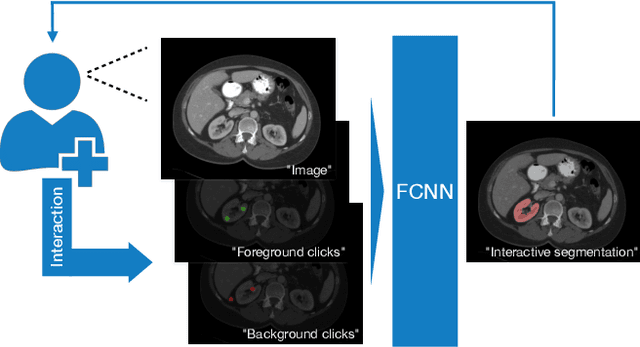
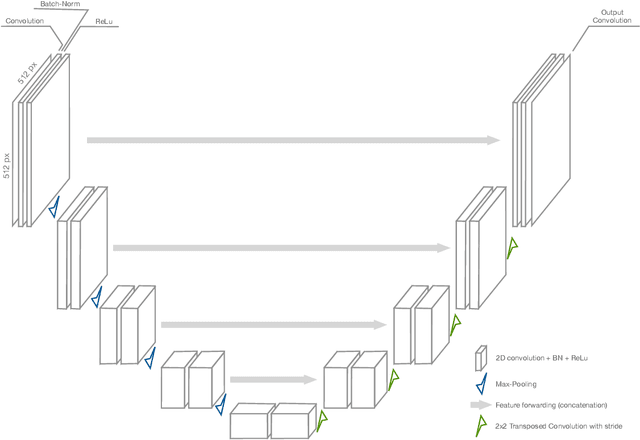
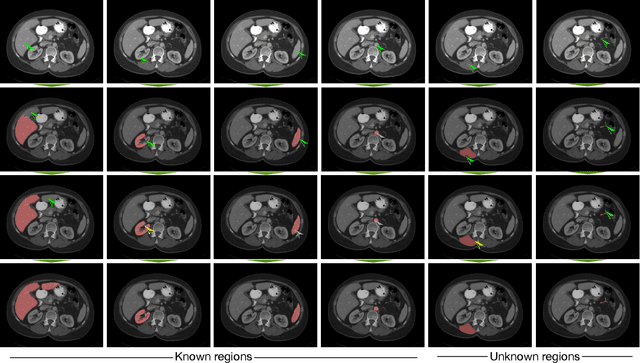
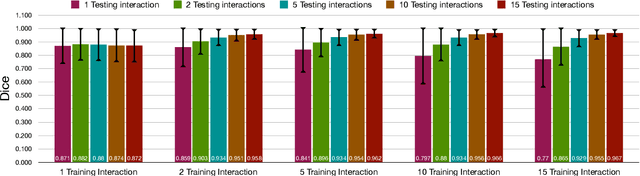
Abstract:Image segmentation plays an essential role in medicine for both diagnostic and interventional tasks. Segmentation approaches are either manual, semi-automated or fully-automated. Manual segmentation offers full control over the quality of the results, but is tedious, time consuming and prone to operator bias. Fully automated methods require no human effort, but often deliver sub-optimal results without providing users with the means to make corrections. Semi-automated approaches keep users in control of the results by providing means for interaction, but the main challenge is to offer a good trade-off between precision and required interaction. In this paper we present a deep learning (DL) based semi-automated segmentation approach that aims to be a "smart" interactive tool for region of interest delineation in medical images. We demonstrate its use for segmenting multiple organs on computed tomography (CT) of the abdomen. Our approach solves some of the most pressing clinical challenges: (i) it requires only one to a few user clicks to deliver excellent 2D segmentations in a fast and reliable fashion; (ii) it can generalize to previously unseen structures and "corner cases"; (iii) it delivers results that can be corrected quickly in a smart and intuitive way up to an arbitrary degree of precision chosen by the user and (iv) ensures high accuracy. We present our approach and compare it to other techniques and previous work to show the advantages brought by our method.
Straight to the point: reinforcement learning for user guidance in ultrasound
Mar 02, 2019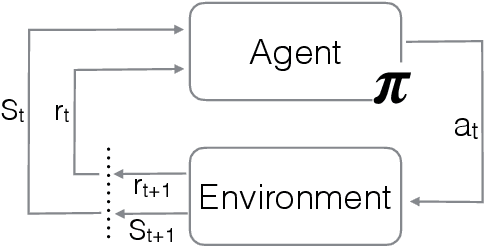
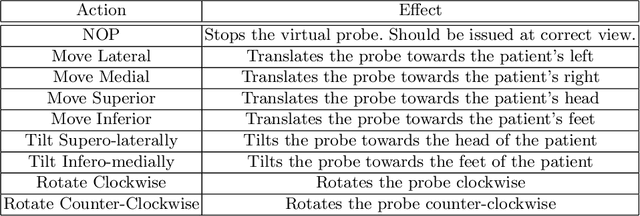
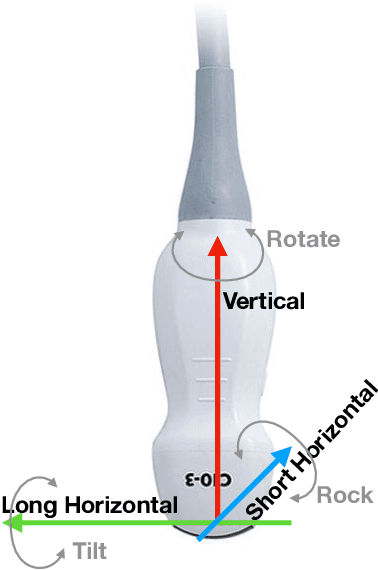
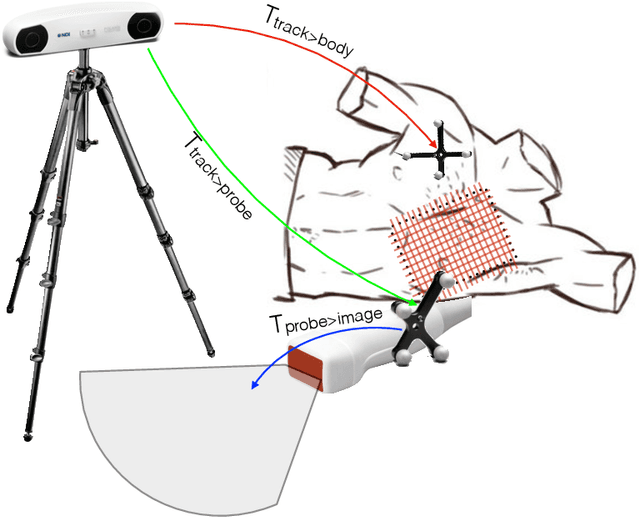
Abstract:Point of care ultrasound (POCUS) consists in the use of ultrasound imaging in critical or emergency situations to support clinical decisions by healthcare professionals and first responders. In this setting it is essential to be able to provide means to obtain diagnostic data to potentially inexperienced users who did not receive an extensive medical training. Interpretation and acquisition of ultrasound images is not trivial. First, the user needs to find a suitable sound window which can be used to get a clear image, and then he needs to correctly interpret it to perform a diagnosis. Although many recent approaches focus on developing smart ultrasound devices that add interpretation capabilities to existing systems, our goal in this paper is to present a reinforcement learning (RL) strategy which is capable to guide novice users to the correct sonic window and enable them to obtain clinically relevant pictures of the anatomy of interest. We apply our approach to cardiac images acquired from the parasternal long axis (PLAx) view of the left ventricle of the heart.
CFCM: Segmentation via Coarse to Fine Context Memory
Jun 04, 2018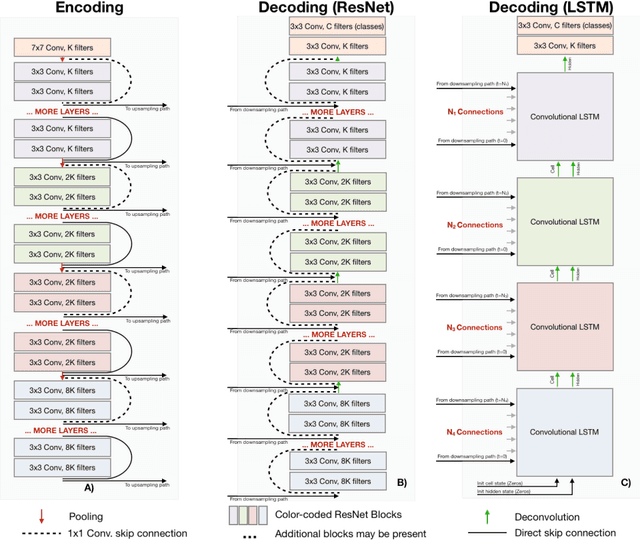
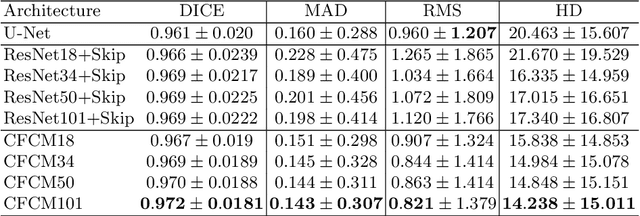
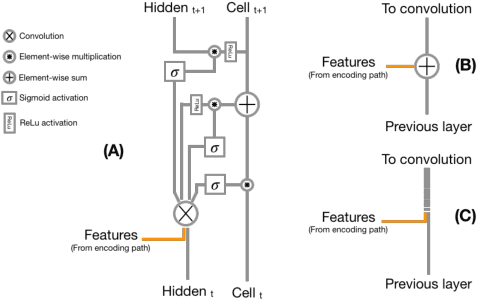
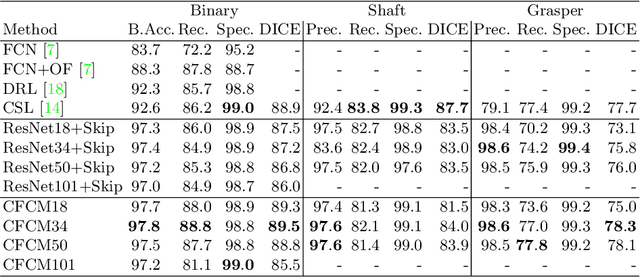
Abstract:Recent neural-network-based architectures for image segmentation make extensive usage of feature forwarding mechanisms to integrate information from multiple scales. Although yielding good results, even deeper architectures and alternative methods for feature fusion at different resolutions have been scarcely investigated for medical applications. In this work we propose to implement segmentation via an encoder-decoder architecture which differs from any other previously published method since (i) it employs a very deep architecture based on residual learning and (ii) combines features via a convolutional Long Short Term Memory (LSTM), instead of concatenation or summation. The intuition is that the memory mechanism implemented by LSTMs can better integrate features from different scales through a coarse-to-fine strategy; hence the name Coarse-to-Fine Context Memory (CFCM). We demonstrate the remarkable advantages of this approach on two datasets: the Montgomery county lung segmentation dataset, and the EndoVis 2015 challenge dataset for surgical instrument segmentation.
TOMAAT: volumetric medical image analysis as a cloud service
Apr 25, 2018
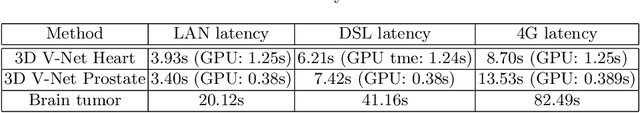
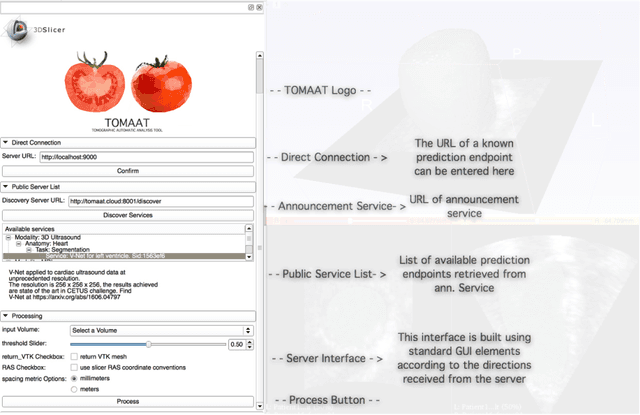
Abstract:Deep learning has been recently applied to a multitude of computer vision and medical image analysis problems. Although recent research efforts have improved the state of the art, most of the methods cannot be easily accessed, compared or used by either researchers or the general public. Researchers often publish their code and trained models on the internet, but this does not always enable these approaches to be easily used or integrated in stand-alone applications and existing workflows. In this paper we propose a framework which allows easy deployment and access of deep learning methods for segmentation through a cloud-based architecture. Our approach comprises three parts: a server, which wraps trained deep learning models and their pre- and post-processing data pipelines and makes them available on the cloud; a client which interfaces with the server to obtain predictions on user data; a service registry that informs clients about available prediction endpoints that are available in the cloud. These three parts constitute the open-source TOMAAT framework.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge