Kenneth Philbrick
Health AI Developer Foundations
Nov 26, 2024



Abstract:Robust medical Machine Learning (ML) models have the potential to revolutionize healthcare by accelerating clinical research, improving workflows and outcomes, and producing novel insights or capabilities. Developing such ML models from scratch is cost prohibitive and requires substantial compute, data, and time (e.g., expert labeling). To address these challenges, we introduce Health AI Developer Foundations (HAI-DEF), a suite of pre-trained, domain-specific foundation models, tools, and recipes to accelerate building ML for health applications. The models cover various modalities and domains, including radiology (X-rays and computed tomography), histopathology, dermatological imaging, and audio. These models provide domain specific embeddings that facilitate AI development with less labeled data, shorter training times, and reduced computational costs compared to traditional approaches. In addition, we utilize a common interface and style across these models, and prioritize usability to enable developers to integrate HAI-DEF efficiently. We present model evaluations across various tasks and conclude with a discussion of their application and evaluation, covering the importance of ensuring efficacy, fairness, and equity. Finally, while HAI-DEF and specifically the foundation models lower the barrier to entry for ML in healthcare, we emphasize the importance of validation with problem- and population-specific data for each desired usage setting. This technical report will be updated over time as more modalities and features are added.
SOUP-GAN: Super-Resolution MRI Using Generative Adversarial Networks
Jun 04, 2021
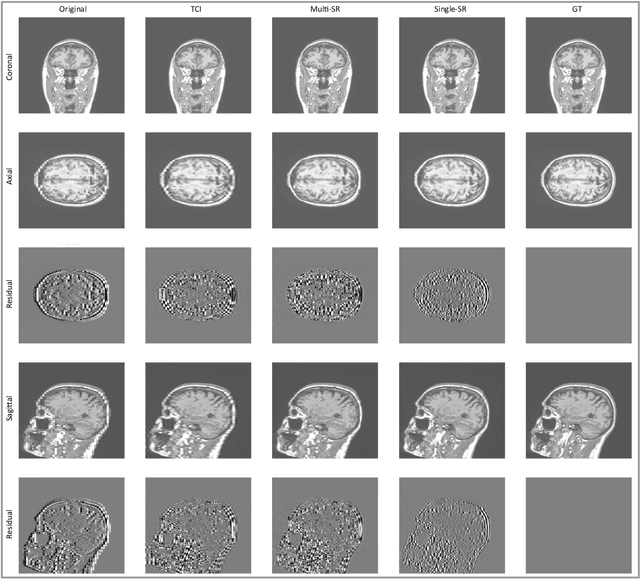
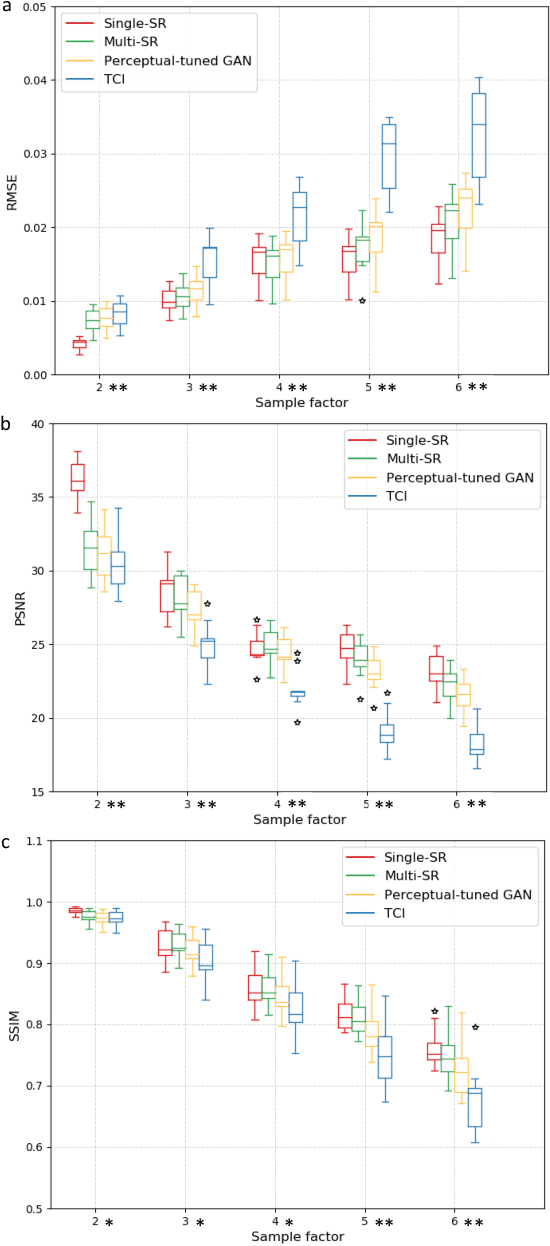
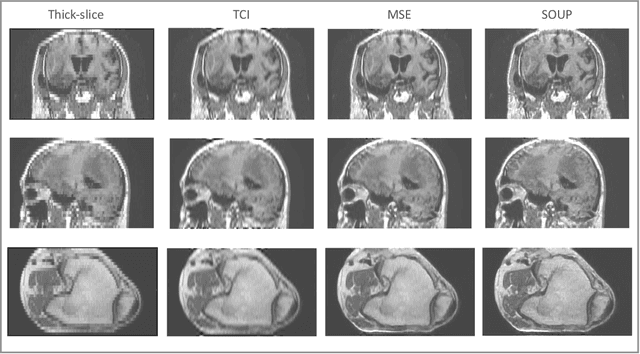

Abstract:There is a growing demand for high-resolution (HR) medical images in both the clinical and research applications. Image quality is inevitably traded off with the acquisition time for better patient comfort, lower examination costs, dose, and fewer motion-induced artifacts. For many image-based tasks, increasing the apparent resolution in the perpendicular plane to produce multi-planar reformats or 3D images is commonly used. Single image super-resolution (SR) is a promising technique to provide HR images based on unsupervised learning to increase resolution of a 2D image, but there are few reports on 3D SR. Further, perceptual loss is proposed in the literature to better capture the textual details and edges than using pixel-wise loss functions, by comparing the semantic distances in the high-dimensional feature space of a pre-trained 2D network (e.g., VGG). However, it is not clear how one should generalize it to 3D medical images, and the attendant implications are still unclear. In this paper, we propose a framework called SOUP-GAN: Super-resolution Optimized Using Perceptual-tuned Generative Adversarial Network (GAN), in order to produce thinner slice (e.g., high resolution in the 'Z' plane) medical images with anti-aliasing and deblurring. The proposed method outperforms other conventional resolution-enhancement methods and previous SR work on medical images upon both qualitative and quantitative comparisons. Specifically, we examine the model in terms of its generalization for various SR ratios and imaging modalities. By addressing those limitations, our model shows promise as a novel 3D SR interpolation technique, providing potential applications in both clinical and research settings.
Interactive segmentation of medical images through fully convolutional neural networks
Mar 19, 2019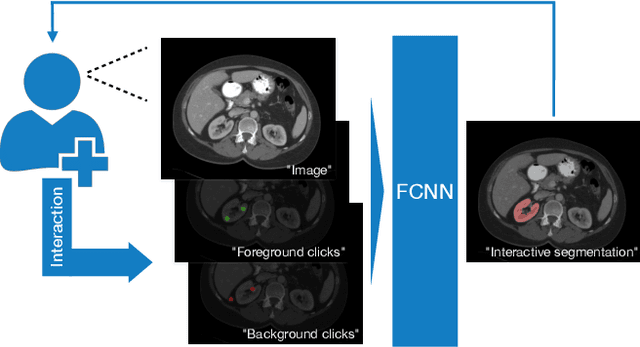
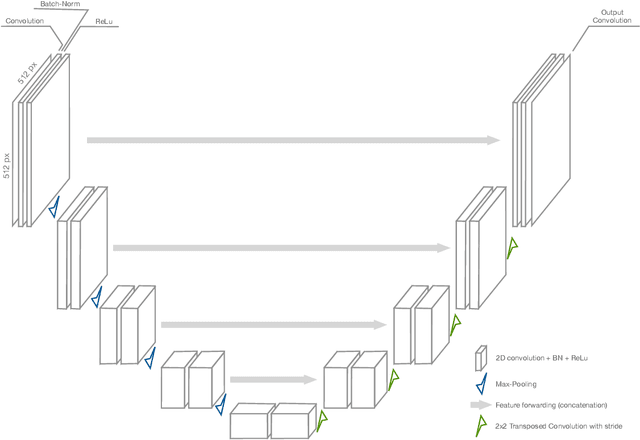
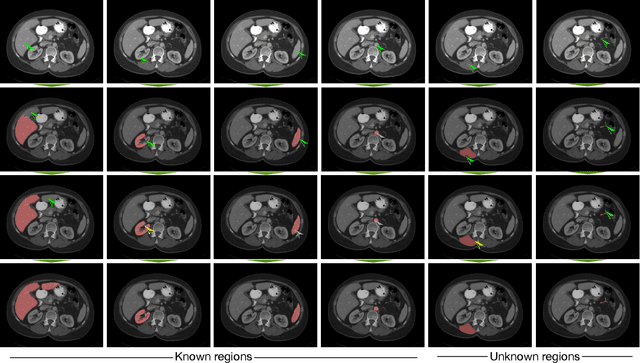
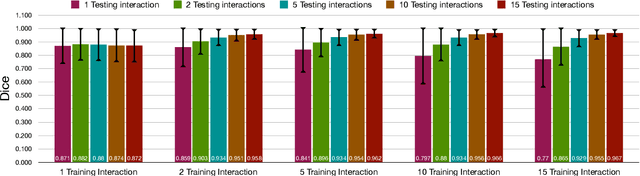
Abstract:Image segmentation plays an essential role in medicine for both diagnostic and interventional tasks. Segmentation approaches are either manual, semi-automated or fully-automated. Manual segmentation offers full control over the quality of the results, but is tedious, time consuming and prone to operator bias. Fully automated methods require no human effort, but often deliver sub-optimal results without providing users with the means to make corrections. Semi-automated approaches keep users in control of the results by providing means for interaction, but the main challenge is to offer a good trade-off between precision and required interaction. In this paper we present a deep learning (DL) based semi-automated segmentation approach that aims to be a "smart" interactive tool for region of interest delineation in medical images. We demonstrate its use for segmenting multiple organs on computed tomography (CT) of the abdomen. Our approach solves some of the most pressing clinical challenges: (i) it requires only one to a few user clicks to deliver excellent 2D segmentations in a fast and reliable fashion; (ii) it can generalize to previously unseen structures and "corner cases"; (iii) it delivers results that can be corrected quickly in a smart and intuitive way up to an arbitrary degree of precision chosen by the user and (iv) ensures high accuracy. We present our approach and compare it to other techniques and previous work to show the advantages brought by our method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge