Yuanji Zhang
Medical-Knowledge Driven Multiple Instance Learning for Classifying Severe Abdominal Anomalies on Prenatal Ultrasound
Jul 02, 2025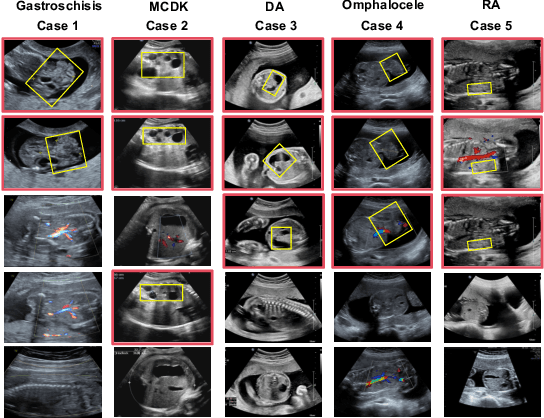
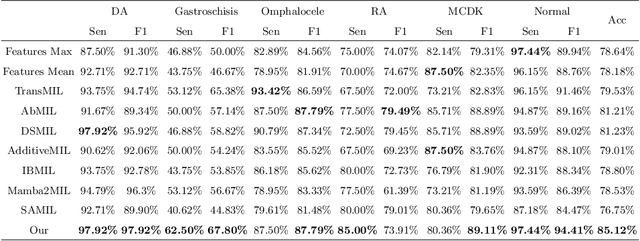
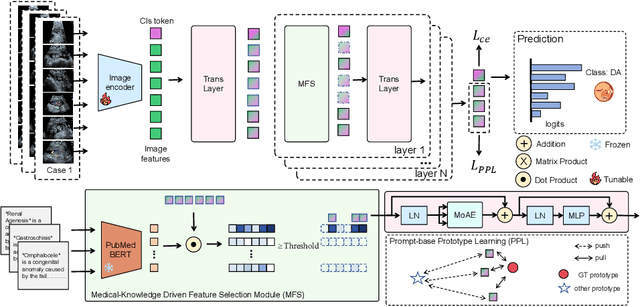
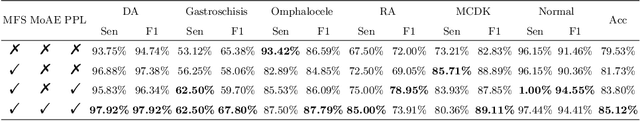
Abstract:Fetal abdominal malformations are serious congenital anomalies that require accurate diagnosis to guide pregnancy management and reduce mortality. Although AI has demonstrated significant potential in medical diagnosis, its application to prenatal abdominal anomalies remains limited. Most existing studies focus on image-level classification and rely on standard plane localization, placing less emphasis on case-level diagnosis. In this paper, we develop a case-level multiple instance learning (MIL)-based method, free of standard plane localization, for classifying fetal abdominal anomalies in prenatal ultrasound. Our contribution is three-fold. First, we adopt a mixture-of-attention-experts module (MoAE) to weight different attention heads for various planes. Secondly, we propose a medical-knowledge-driven feature selection module (MFS) to align image features with medical knowledge, performing self-supervised image token selection at the case-level. Finally, we propose a prompt-based prototype learning (PPL) to enhance the MFS. Extensively validated on a large prenatal abdominal ultrasound dataset containing 2,419 cases, with a total of 24,748 images and 6 categories, our proposed method outperforms the state-of-the-art competitors. Codes are available at:https://github.com/LL-AC/AAcls.
FetalFlex: Anatomy-Guided Diffusion Model for Flexible Control on Fetal Ultrasound Image Synthesis
Mar 19, 2025Abstract:Fetal ultrasound (US) examinations require the acquisition of multiple planes, each providing unique diagnostic information to evaluate fetal development and screening for congenital anomalies. However, obtaining a comprehensive, multi-plane annotated fetal US dataset remains challenging, particularly for rare or complex anomalies owing to their low incidence and numerous subtypes. This poses difficulties in training novice radiologists and developing robust AI models, especially for detecting abnormal fetuses. In this study, we introduce a Flexible Fetal US image generation framework (FetalFlex) to address these challenges, which leverages anatomical structures and multimodal information to enable controllable synthesis of fetal US images across diverse planes. Specifically, FetalFlex incorporates a pre-alignment module to enhance controllability and introduces a repaint strategy to ensure consistent texture and appearance. Moreover, a two-stage adaptive sampling strategy is developed to progressively refine image quality from coarse to fine levels. We believe that FetalFlex is the first method capable of generating both in-distribution normal and out-of-distribution abnormal fetal US images, without requiring any abnormal data. Experiments on multi-center datasets demonstrate that FetalFlex achieved state-of-the-art performance across multiple image quality metrics. A reader study further confirms the close alignment of the generated results with expert visual assessments. Furthermore, synthetic images by FetalFlex significantly improve the performance of six typical deep models in downstream classification and anomaly detection tasks. Lastly, FetalFlex's anatomy-level controllable generation offers a unique advantage for anomaly simulation and creating paired or counterfactual data at the pixel level. The demo is available at: https://dyf1023.github.io/FetalFlex/.
FetusMapV2: Enhanced Fetal Pose Estimation in 3D Ultrasound
Oct 30, 2023Abstract:Fetal pose estimation in 3D ultrasound (US) involves identifying a set of associated fetal anatomical landmarks. Its primary objective is to provide comprehensive information about the fetus through landmark connections, thus benefiting various critical applications, such as biometric measurements, plane localization, and fetal movement monitoring. However, accurately estimating the 3D fetal pose in US volume has several challenges, including poor image quality, limited GPU memory for tackling high dimensional data, symmetrical or ambiguous anatomical structures, and considerable variations in fetal poses. In this study, we propose a novel 3D fetal pose estimation framework (called FetusMapV2) to overcome the above challenges. Our contribution is three-fold. First, we propose a heuristic scheme that explores the complementary network structure-unconstrained and activation-unreserved GPU memory management approaches, which can enlarge the input image resolution for better results under limited GPU memory. Second, we design a novel Pair Loss to mitigate confusion caused by symmetrical and similar anatomical structures. It separates the hidden classification task from the landmark localization task and thus progressively eases model learning. Last, we propose a shape priors-based self-supervised learning by selecting the relatively stable landmarks to refine the pose online. Extensive experiments and diverse applications on a large-scale fetal US dataset including 1000 volumes with 22 landmarks per volume demonstrate that our method outperforms other strong competitors.
Multi-IMU with Online Self-Consistency for Freehand 3D Ultrasound Reconstruction
Jul 19, 2023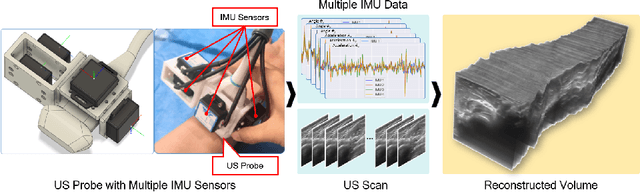
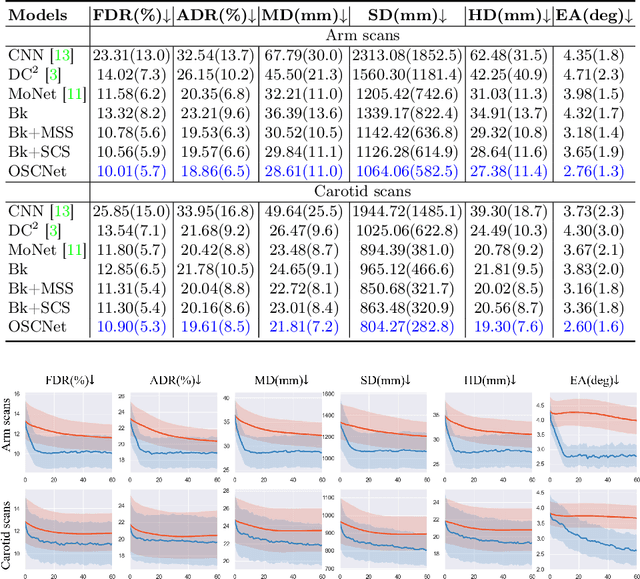
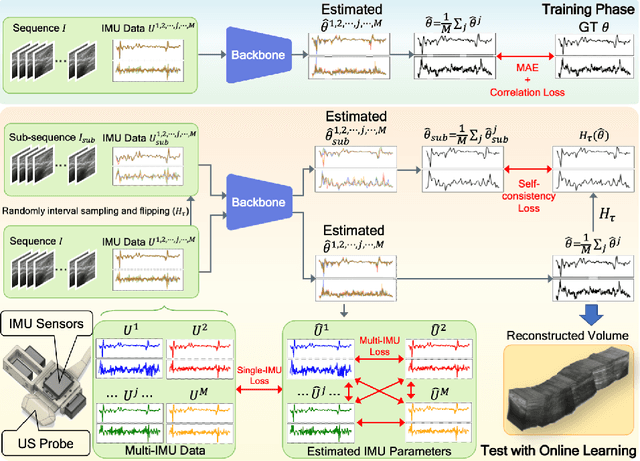
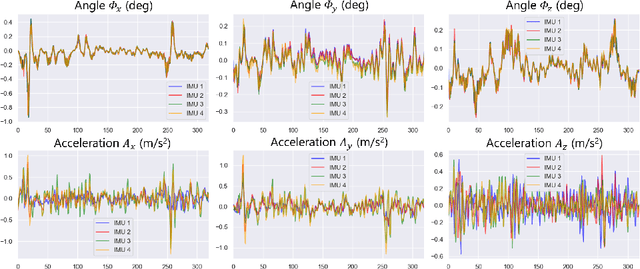
Abstract:Ultrasound (US) imaging is a popular tool in clinical diagnosis, offering safety, repeatability, and real-time capabilities. Freehand 3D US is a technique that provides a deeper understanding of scanned regions without increasing complexity. However, estimating elevation displacement and accumulation error remains challenging, making it difficult to infer the relative position using images alone. The addition of external lightweight sensors has been proposed to enhance reconstruction performance without adding complexity, which has been shown to be beneficial. We propose a novel online self-consistency network (OSCNet) using multiple inertial measurement units (IMUs) to improve reconstruction performance. OSCNet utilizes a modal-level self-supervised strategy to fuse multiple IMU information and reduce differences between reconstruction results obtained from each IMU data. Additionally, a sequence-level self-consistency strategy is proposed to improve the hierarchical consistency of prediction results among the scanning sequence and its sub-sequences. Experiments on large-scale arm and carotid datasets with multiple scanning tactics demonstrate that our OSCNet outperforms previous methods, achieving state-of-the-art reconstruction performance.
Inflated 3D Convolution-Transformer for Weakly-supervised Carotid Stenosis Grading with Ultrasound Videos
Jun 12, 2023



Abstract:Localization of the narrowest position of the vessel and corresponding vessel and remnant vessel delineation in carotid ultrasound (US) are essential for carotid stenosis grading (CSG) in clinical practice. However, the pipeline is time-consuming and tough due to the ambiguous boundaries of plaque and temporal variation. To automatize this procedure, a large number of manual delineations are usually required, which is not only laborious but also not reliable given the annotation difficulty. In this study, we present the first video classification framework for automatic CSG. Our contribution is three-fold. First, to avoid the requirement of laborious and unreliable annotation, we propose a novel and effective video classification network for weakly-supervised CSG. Second, to ease the model training, we adopt an inflation strategy for the network, where pre-trained 2D convolution weights can be adapted into the 3D counterpart in our network for an effective warm start. Third, to enhance the feature discrimination of the video, we propose a novel attention-guided multi-dimension fusion (AMDF) transformer encoder to model and integrate global dependencies within and across spatial and temporal dimensions, where two lightweight cross-dimensional attention mechanisms are designed. Our approach is extensively validated on a large clinically collected carotid US video dataset, demonstrating state-of-the-art performance compared with strong competitors.
Weakly-supervised High-fidelity Ultrasound Video Synthesis with Feature Decoupling
Jul 01, 2022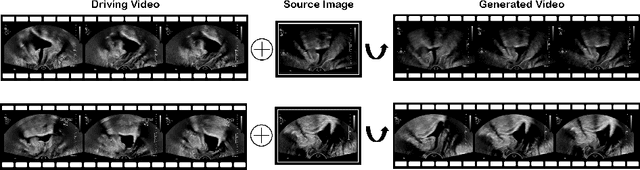
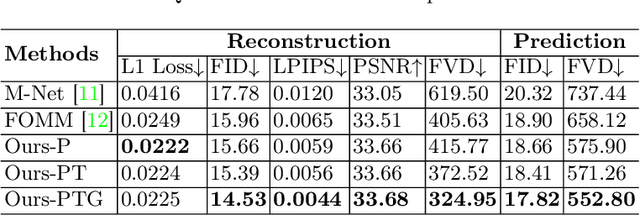
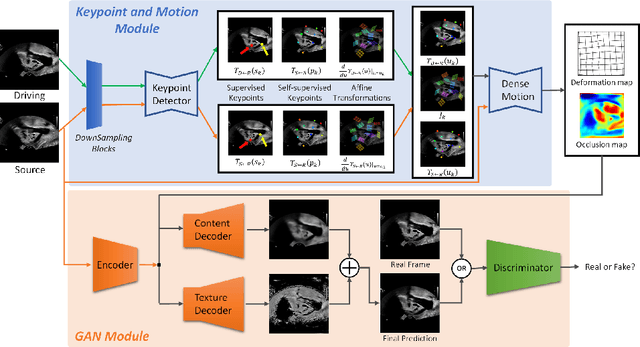
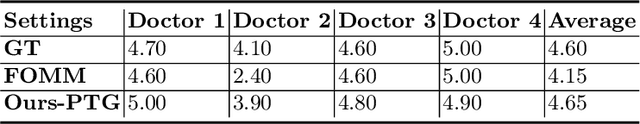
Abstract:Ultrasound (US) is widely used for its advantages of real-time imaging, radiation-free and portability. In clinical practice, analysis and diagnosis often rely on US sequences rather than a single image to obtain dynamic anatomical information. This is challenging for novices to learn because practicing with adequate videos from patients is clinically unpractical. In this paper, we propose a novel framework to synthesize high-fidelity US videos. Specifically, the synthesis videos are generated by animating source content images based on the motion of given driving videos. Our highlights are three-fold. First, leveraging the advantages of self- and fully-supervised learning, our proposed system is trained in weakly-supervised manner for keypoint detection. These keypoints then provide vital information for handling complex high dynamic motions in US videos. Second, we decouple content and texture learning using the dual decoders to effectively reduce the model learning difficulty. Last, we adopt the adversarial training strategy with GAN losses for further improving the sharpness of the generated videos, narrowing the gap between real and synthesis videos. We validate our method on a large in-house pelvic dataset with high dynamic motion. Extensive evaluation metrics and user study prove the effectiveness of our proposed method.
Statistical Dependency Guided Contrastive Learning for Multiple Labeling in Prenatal Ultrasound
Aug 11, 2021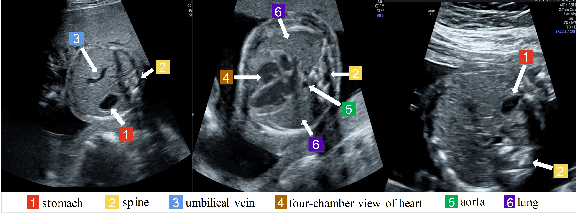
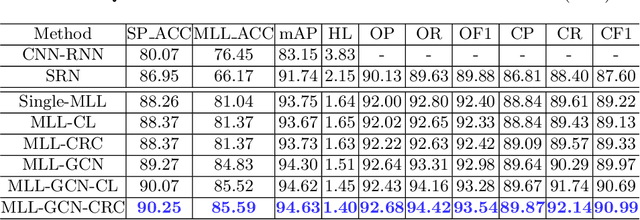
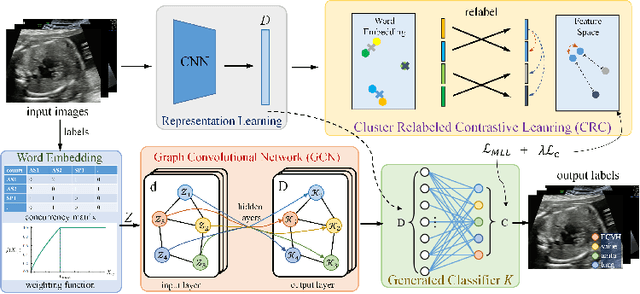
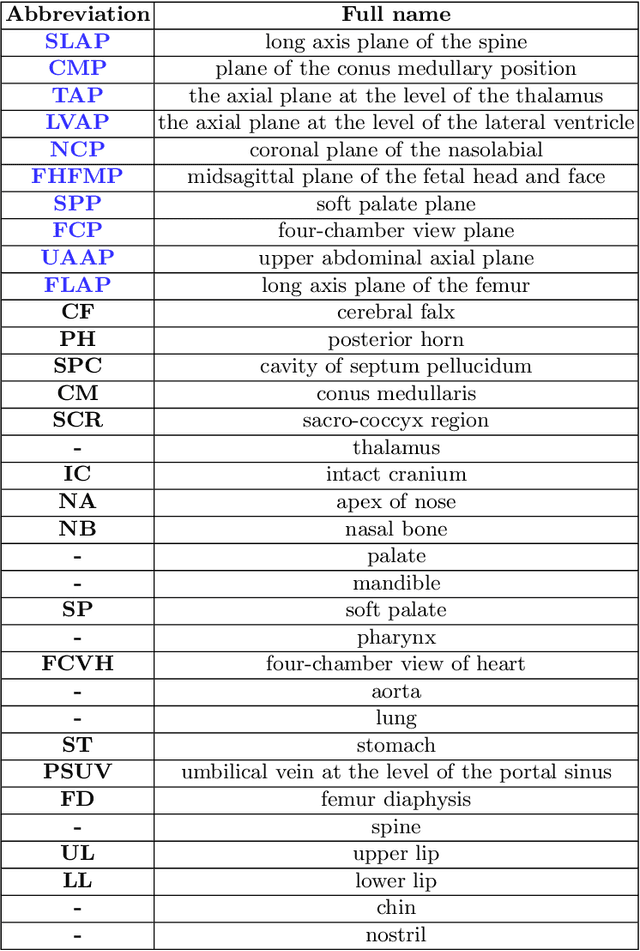
Abstract:Standard plane recognition plays an important role in prenatal ultrasound (US) screening. Automatically recognizing the standard plane along with the corresponding anatomical structures in US image can not only facilitate US image interpretation but also improve diagnostic efficiency. In this study, we build a novel multi-label learning (MLL) scheme to identify multiple standard planes and corresponding anatomical structures of fetus simultaneously. Our contribution is three-fold. First, we represent the class correlation by word embeddings to capture the fine-grained semantic and latent statistical concurrency. Second, we equip the MLL with a graph convolutional network to explore the inner and outer relationship among categories. Third, we propose a novel cluster relabel-based contrastive learning algorithm to encourage the divergence among ambiguous classes. Extensive validation was performed on our large in-house dataset. Our approach reports the highest accuracy as 90.25% for standard planes labeling, 85.59% for planes and structures labeling and mAP as 94.63%. The proposed MLL scheme provides a novel perspective for standard plane recognition and can be easily extended to other medical image classification tasks.
Searching Collaborative Agents for Multi-plane Localization in 3D Ultrasound
May 22, 2021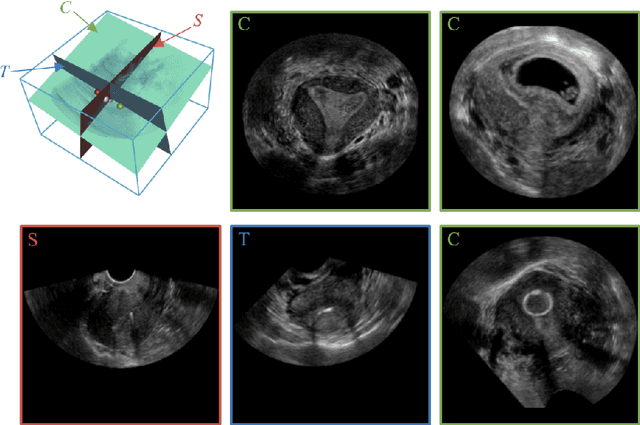
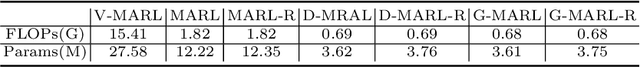
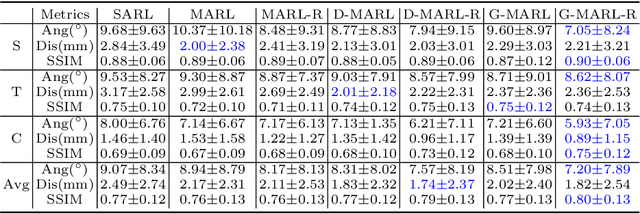
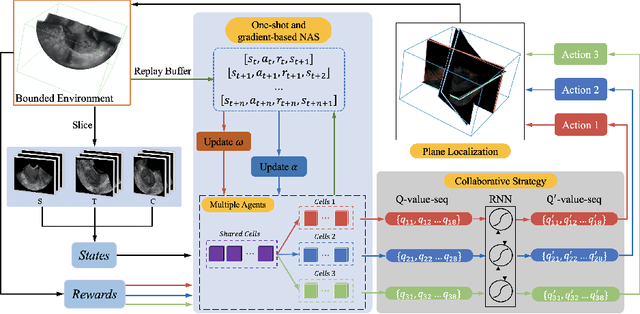

Abstract:3D ultrasound (US) has become prevalent due to its rich spatial and diagnostic information not contained in 2D US. Moreover, 3D US can contain multiple standard planes (SPs) in one shot. Thus, automatically localizing SPs in 3D US has the potential to improve user-independence and scanning-efficiency. However, manual SP localization in 3D US is challenging because of the low image quality, huge search space and large anatomical variability. In this work, we propose a novel multi-agent reinforcement learning (MARL) framework to simultaneously localize multiple SPs in 3D US. Our contribution is four-fold. First, our proposed method is general and it can accurately localize multiple SPs in different challenging US datasets. Second, we equip the MARL system with a recurrent neural network (RNN) based collaborative module, which can strengthen the communication among agents and learn the spatial relationship among planes effectively. Third, we explore to adopt the neural architecture search (NAS) to automatically design the network architecture of both the agents and the collaborative module. Last, we believe we are the first to realize automatic SP localization in pelvic US volumes, and note that our approach can handle both normal and abnormal uterus cases. Extensively validated on two challenging datasets of the uterus and fetal brain, our proposed method achieves the average localization accuracy of 7.03 degrees/1.59mm and 9.75 degrees/1.19mm. Experimental results show that our light-weight MARL model has higher accuracy than state-of-the-art methods.
Agent with Warm Start and Adaptive Dynamic Termination for Plane Localization in 3D Ultrasound
Mar 26, 2021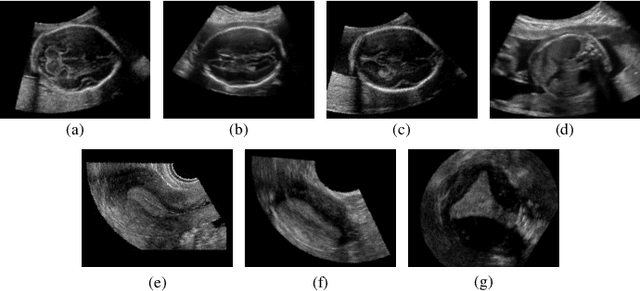
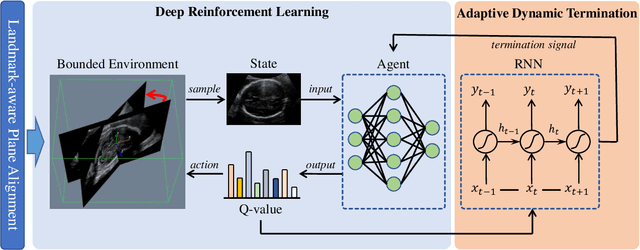
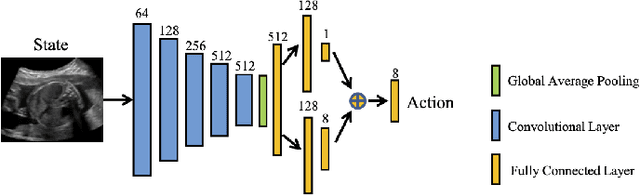
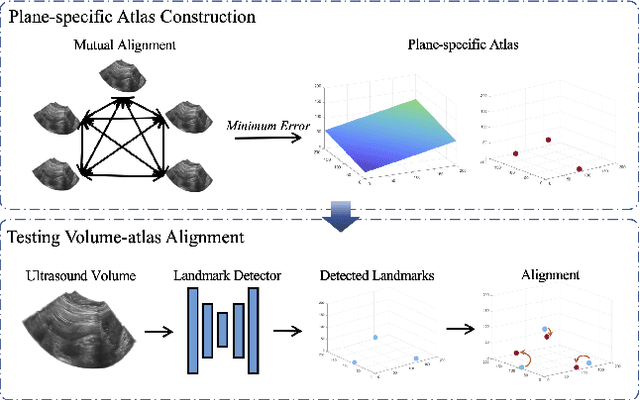
Abstract:Accurate standard plane (SP) localization is the fundamental step for prenatal ultrasound (US) diagnosis. Typically, dozens of US SPs are collected to determine the clinical diagnosis. 2D US has to perform scanning for each SP, which is time-consuming and operator-dependent. While 3D US containing multiple SPs in one shot has the inherent advantages of less user-dependency and more efficiency. Automatically locating SP in 3D US is very challenging due to the huge search space and large fetal posture variations. Our previous study proposed a deep reinforcement learning (RL) framework with an alignment module and active termination to localize SPs in 3D US automatically. However, termination of agent search in RL is important and affects the practical deployment. In this study, we enhance our previous RL framework with a newly designed adaptive dynamic termination to enable an early stop for the agent searching, saving at most 67% inference time, thus boosting the accuracy and efficiency of the RL framework at the same time. Besides, we validate the effectiveness and generalizability of our algorithm extensively on our in-house multi-organ datasets containing 433 fetal brain volumes, 519 fetal abdomen volumes, and 683 uterus volumes. Our approach achieves localization error of 2.52mm/10.26 degrees, 2.48mm/10.39 degrees, 2.02mm/10.48 degrees, 2.00mm/14.57 degrees, 2.61mm/9.71 degrees, 3.09mm/9.58 degrees, 1.49mm/7.54 degrees for the transcerebellar, transventricular, transthalamic planes in fetal brain, abdominal plane in fetal abdomen, and mid-sagittal, transverse and coronal planes in uterus, respectively. Experimental results show that our method is general and has the potential to improve the efficiency and standardization of US scanning.
Generalize Ultrasound Image Segmentation via Instant and Plug & Play Style Transfer
Jan 11, 2021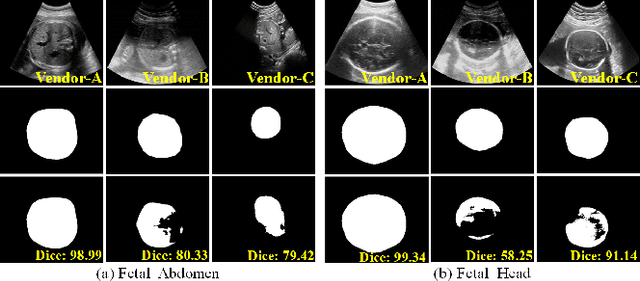

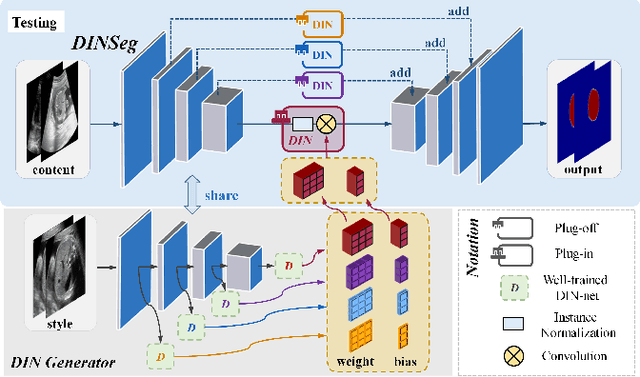

Abstract:Deep segmentation models that generalize to images with unknown appearance are important for real-world medical image analysis. Retraining models leads to high latency and complex pipelines, which are impractical in clinical settings. The situation becomes more severe for ultrasound image analysis because of their large appearance shifts. In this paper, we propose a novel method for robust segmentation under unknown appearance shifts. Our contribution is three-fold. First, we advance a one-stage plug-and-play solution by embedding hierarchical style transfer units into a segmentation architecture. Our solution can remove appearance shifts and perform segmentation simultaneously. Second, we adopt Dynamic Instance Normalization to conduct precise and dynamic style transfer in a learnable manner, rather than previously fixed style normalization. Third, our solution is fast and lightweight for routine clinical adoption. Given 400*400 image input, our solution only needs an additional 0.2ms and 1.92M FLOPs to handle appearance shifts compared to the baseline pipeline. Extensive experiments are conducted on a large dataset from three vendors demonstrate our proposed method enhances the robustness of deep segmentation models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge