Yan Cao
Enhancing WiFi CSI Fingerprinting: A Deep Auxiliary Learning Approach
Oct 26, 2025Abstract:Radio frequency (RF) fingerprinting techniques provide a promising supplement to cryptography-based approaches but rely on dedicated equipment to capture in-phase and quadrature (IQ) samples, hindering their wide adoption. Recent advances advocate easily obtainable channel state information (CSI) by commercial WiFi devices for lightweight RF fingerprinting, while falling short in addressing the challenges of coarse granularity of CSI measurements in an open-world setting. In this paper, we propose CSI2Q, a novel CSI fingerprinting system that achieves comparable performance to IQ-based approaches. Instead of extracting fingerprints directly from raw CSI measurements, CSI2Q first transforms frequency-domain CSI measurements into time-domain signals that share the same feature space with IQ samples. Then, we employ a deep auxiliary learning strategy to transfer useful knowledge from an IQ fingerprinting model to the CSI counterpart. Finally, the trained CSI model is combined with an OpenMax function to estimate the likelihood of unknown ones. We evaluate CSI2Q on one synthetic CSI dataset involving 85 devices and two real CSI datasets, including 10 and 25 WiFi routers, respectively. Our system achieves accuracy increases of at least 16% on the synthetic CSI dataset, 20% on the in-lab CSI dataset, and 17% on the in-the-wild CSI dataset.
Flip Learning: Weakly Supervised Erase to Segment Nodules in Breast Ultrasound
Mar 27, 2025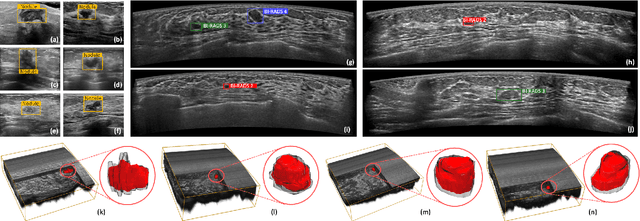
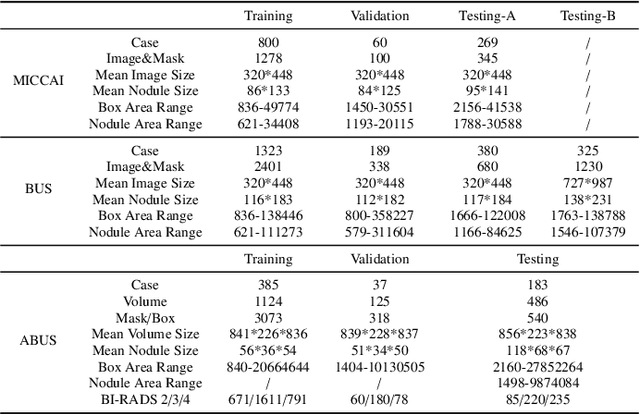
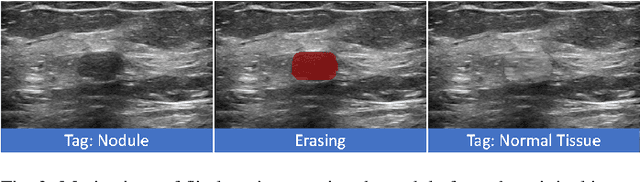
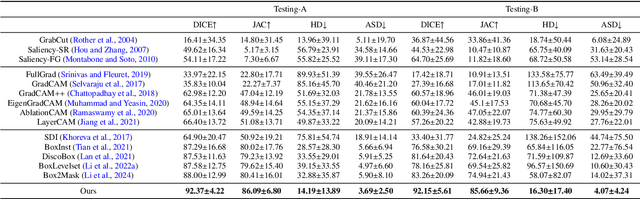
Abstract:Accurate segmentation of nodules in both 2D breast ultrasound (BUS) and 3D automated breast ultrasound (ABUS) is crucial for clinical diagnosis and treatment planning. Therefore, developing an automated system for nodule segmentation can enhance user independence and expedite clinical analysis. Unlike fully-supervised learning, weakly-supervised segmentation (WSS) can streamline the laborious and intricate annotation process. However, current WSS methods face challenges in achieving precise nodule segmentation, as many of them depend on inaccurate activation maps or inefficient pseudo-mask generation algorithms. In this study, we introduce a novel multi-agent reinforcement learning-based WSS framework called Flip Learning, which relies solely on 2D/3D boxes for accurate segmentation. Specifically, multiple agents are employed to erase the target from the box to facilitate classification tag flipping, with the erased region serving as the predicted segmentation mask. The key contributions of this research are as follows: (1) Adoption of a superpixel/supervoxel-based approach to encode the standardized environment, capturing boundary priors and expediting the learning process. (2) Introduction of three meticulously designed rewards, comprising a classification score reward and two intensity distribution rewards, to steer the agents' erasing process precisely, thereby avoiding both under- and over-segmentation. (3) Implementation of a progressive curriculum learning strategy to enable agents to interact with the environment in a progressively challenging manner, thereby enhancing learning efficiency. Extensively validated on the large in-house BUS and ABUS datasets, our Flip Learning method outperforms state-of-the-art WSS methods and foundation models, and achieves comparable performance as fully-supervised learning algorithms.
Robust Box Prompt based SAM for Medical Image Segmentation
Jul 31, 2024Abstract:The Segment Anything Model (SAM) can achieve satisfactory segmentation performance under high-quality box prompts. However, SAM's robustness is compromised by the decline in box quality, limiting its practicality in clinical reality. In this study, we propose a novel Robust Box prompt based SAM (\textbf{RoBox-SAM}) to ensure SAM's segmentation performance under prompts with different qualities. Our contribution is three-fold. First, we propose a prompt refinement module to implicitly perceive the potential targets, and output the offsets to directly transform the low-quality box prompt into a high-quality one. We then provide an online iterative strategy for further prompt refinement. Second, we introduce a prompt enhancement module to automatically generate point prompts to assist the box-promptable segmentation effectively. Last, we build a self-information extractor to encode the prior information from the input image. These features can optimize the image embeddings and attention calculation, thus, the robustness of SAM can be further enhanced. Extensive experiments on the large medical segmentation dataset including 99,299 images, 5 modalities, and 25 organs/targets validated the efficacy of our proposed RoBox-SAM.
Thyroid ultrasound diagnosis improvement via multi-view self-supervised learning and two-stage pre-training
Feb 18, 2024Abstract:Thyroid nodule classification and segmentation in ultrasound images are crucial for computer-aided diagnosis; however, they face limitations owing to insufficient labeled data. In this study, we proposed a multi-view contrastive self-supervised method to improve thyroid nodule classification and segmentation performance with limited manual labels. Our method aligns the transverse and longitudinal views of the same nodule, thereby enabling the model to focus more on the nodule area. We designed an adaptive loss function that eliminates the limitations of the paired data. Additionally, we adopted a two-stage pre-training to exploit the pre-training on ImageNet and thyroid ultrasound images. Extensive experiments were conducted on a large-scale dataset collected from multiple centers. The results showed that the proposed method significantly improves nodule classification and segmentation performance with limited manual labels and outperforms state-of-the-art self-supervised methods. The two-stage pre-training also significantly exceeded ImageNet pre-training.
FetusMapV2: Enhanced Fetal Pose Estimation in 3D Ultrasound
Oct 30, 2023Abstract:Fetal pose estimation in 3D ultrasound (US) involves identifying a set of associated fetal anatomical landmarks. Its primary objective is to provide comprehensive information about the fetus through landmark connections, thus benefiting various critical applications, such as biometric measurements, plane localization, and fetal movement monitoring. However, accurately estimating the 3D fetal pose in US volume has several challenges, including poor image quality, limited GPU memory for tackling high dimensional data, symmetrical or ambiguous anatomical structures, and considerable variations in fetal poses. In this study, we propose a novel 3D fetal pose estimation framework (called FetusMapV2) to overcome the above challenges. Our contribution is three-fold. First, we propose a heuristic scheme that explores the complementary network structure-unconstrained and activation-unreserved GPU memory management approaches, which can enlarge the input image resolution for better results under limited GPU memory. Second, we design a novel Pair Loss to mitigate confusion caused by symmetrical and similar anatomical structures. It separates the hidden classification task from the landmark localization task and thus progressively eases model learning. Last, we propose a shape priors-based self-supervised learning by selecting the relatively stable landmarks to refine the pose online. Extensive experiments and diverse applications on a large-scale fetal US dataset including 1000 volumes with 22 landmarks per volume demonstrate that our method outperforms other strong competitors.
A Foundation Model for General Moving Object Segmentation in Medical Images
Oct 04, 2023



Abstract:Medical image segmentation aims to delineate the anatomical or pathological structures of interest, playing a crucial role in clinical diagnosis. A substantial amount of high-quality annotated data is crucial for constructing high-precision deep segmentation models. However, medical annotation is highly cumbersome and time-consuming, especially for medical videos or 3D volumes, due to the huge labeling space and poor inter-frame consistency. Recently, a fundamental task named Moving Object Segmentation (MOS) has made significant advancements in natural images. Its objective is to delineate moving objects from the background within image sequences, requiring only minimal annotations. In this paper, we propose the first foundation model, named iMOS, for MOS in medical images. Extensive experiments on a large multi-modal medical dataset validate the effectiveness of the proposed iMOS. Specifically, with the annotation of only a small number of images in the sequence, iMOS can achieve satisfactory tracking and segmentation performance of moving objects throughout the entire sequence in bi-directions. We hope that the proposed iMOS can help accelerate the annotation speed of experts, and boost the development of medical foundation models.
FFPN: Fourier Feature Pyramid Network for Ultrasound Image Segmentation
Aug 26, 2023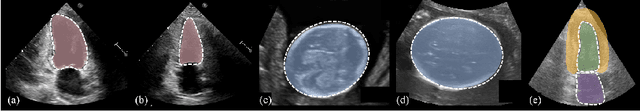
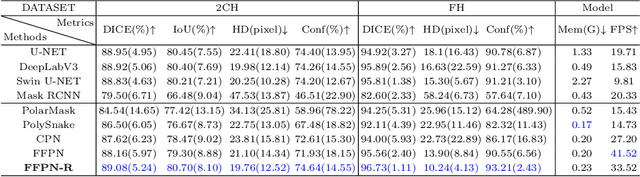
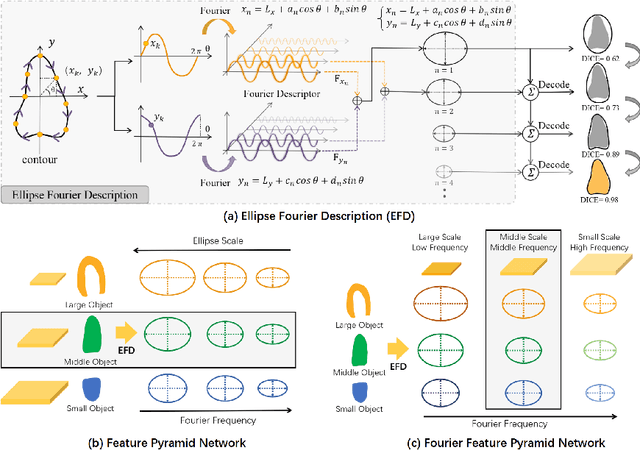
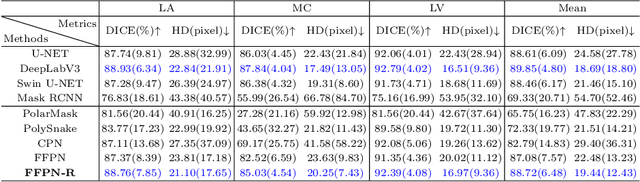
Abstract:Ultrasound (US) image segmentation is an active research area that requires real-time and highly accurate analysis in many scenarios. The detect-to-segment (DTS) frameworks have been recently proposed to balance accuracy and efficiency. However, existing approaches may suffer from inadequate contour encoding or fail to effectively leverage the encoded results. In this paper, we introduce a novel Fourier-anchor-based DTS framework called Fourier Feature Pyramid Network (FFPN) to address the aforementioned issues. The contributions of this paper are two fold. First, the FFPN utilizes Fourier Descriptors to adequately encode contours. Specifically, it maps Fourier series with similar amplitudes and frequencies into the same layer of the feature map, thereby effectively utilizing the encoded Fourier information. Second, we propose a Contour Sampling Refinement (CSR) module based on the contour proposals and refined features produced by the FFPN. This module extracts rich features around the predicted contours to further capture detailed information and refine the contours. Extensive experimental results on three large and challenging datasets demonstrate that our method outperforms other DTS methods in terms of accuracy and efficiency. Furthermore, our framework can generalize well to other detection or segmentation tasks.
Online Reflective Learning for Robust Medical Image Segmentation
Jul 01, 2022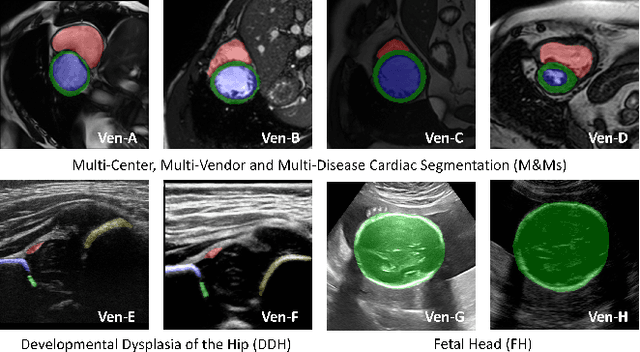
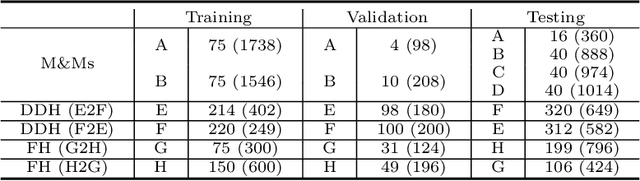
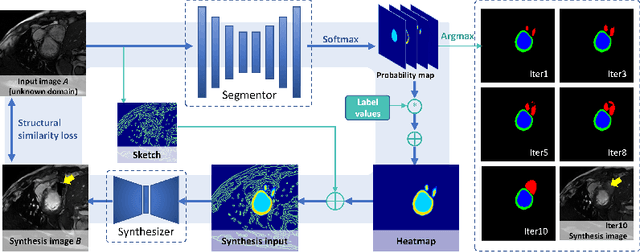
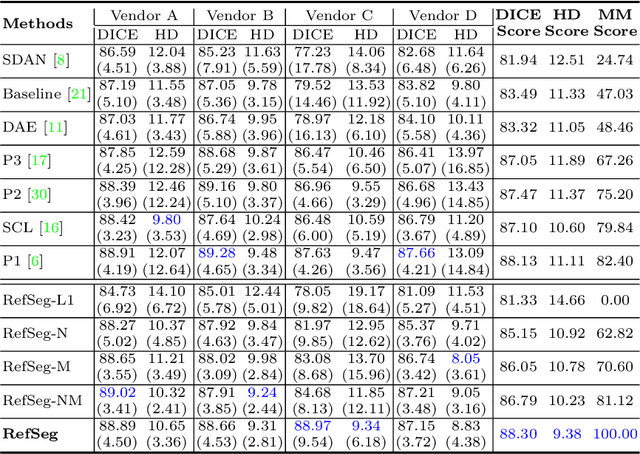
Abstract:Deep segmentation models often face the failure risks when the testing image presents unseen distributions. Improving model robustness against these risks is crucial for the large-scale clinical application of deep models. In this study, inspired by human learning cycle, we propose a novel online reflective learning framework (RefSeg) to improve segmentation robustness. Based on the reflection-on-action conception, our RefSeg firstly drives the deep model to take action to obtain semantic segmentation. Then, RefSeg triggers the model to reflect itself. Because making deep models realize their segmentation failures during testing is challenging, RefSeg synthesizes a realistic proxy image from the semantic mask to help deep models build intuitive and effective reflections. This proxy translates and emphasizes the segmentation flaws. By maximizing the structural similarity between the raw input and the proxy, the reflection-on-action loop is closed with segmentation robustness improved. RefSeg runs in the testing phase and is general for segmentation models. Extensive validation on three medical image segmentation tasks with a public cardiac MR dataset and two in-house large ultrasound datasets show that our RefSeg remarkably improves model robustness and reports state-of-the-art performance over strong competitors.
Statistical Dependency Guided Contrastive Learning for Multiple Labeling in Prenatal Ultrasound
Aug 11, 2021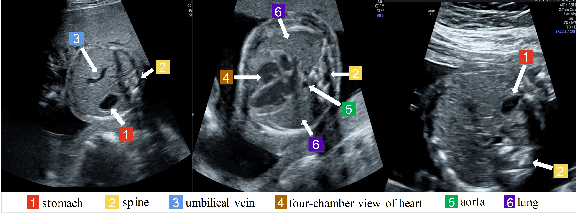
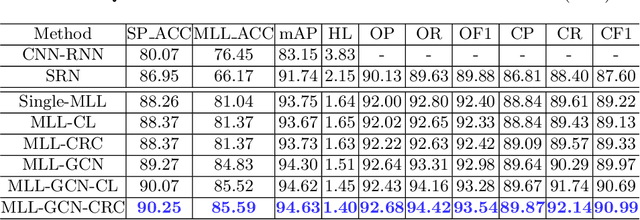
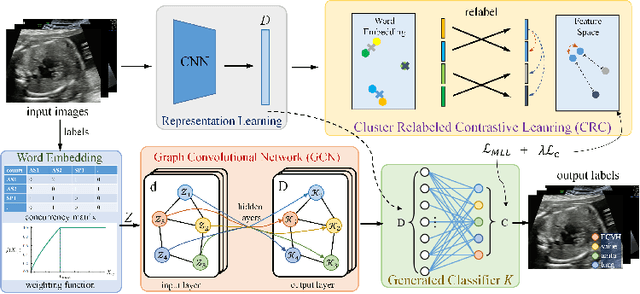
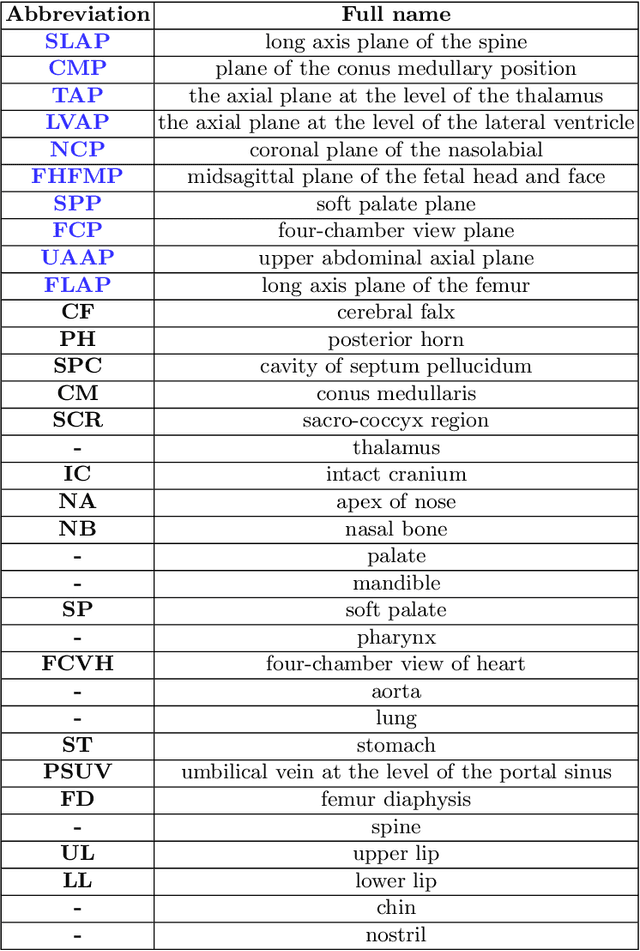
Abstract:Standard plane recognition plays an important role in prenatal ultrasound (US) screening. Automatically recognizing the standard plane along with the corresponding anatomical structures in US image can not only facilitate US image interpretation but also improve diagnostic efficiency. In this study, we build a novel multi-label learning (MLL) scheme to identify multiple standard planes and corresponding anatomical structures of fetus simultaneously. Our contribution is three-fold. First, we represent the class correlation by word embeddings to capture the fine-grained semantic and latent statistical concurrency. Second, we equip the MLL with a graph convolutional network to explore the inner and outer relationship among categories. Third, we propose a novel cluster relabel-based contrastive learning algorithm to encourage the divergence among ambiguous classes. Extensive validation was performed on our large in-house dataset. Our approach reports the highest accuracy as 90.25% for standard planes labeling, 85.59% for planes and structures labeling and mAP as 94.63%. The proposed MLL scheme provides a novel perspective for standard plane recognition and can be easily extended to other medical image classification tasks.
Joint Landmark and Structure Learning for Automatic Evaluation of Developmental Dysplasia of the Hip
Jun 10, 2021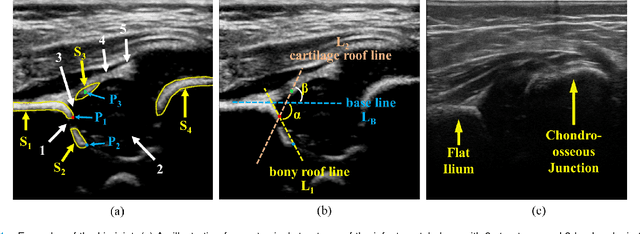
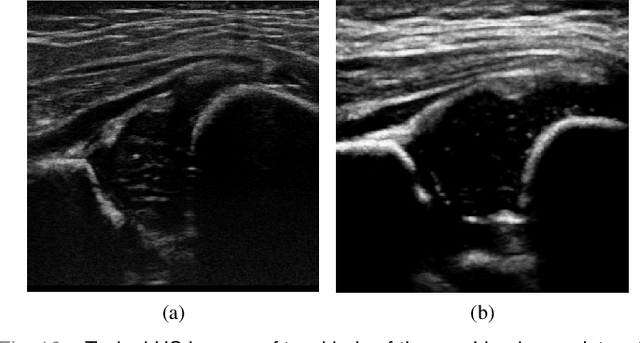
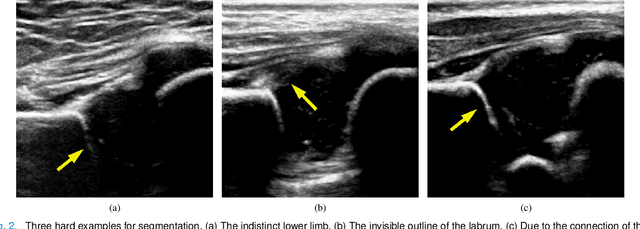
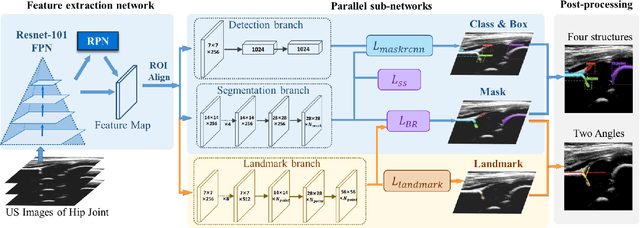
Abstract:The ultrasound (US) screening of the infant hip is vital for the early diagnosis of developmental dysplasia of the hip (DDH). The US diagnosis of DDH refers to measuring alpha and beta angles that quantify hip joint development. These two angles are calculated from key anatomical landmarks and structures of the hip. However, this measurement process is not trivial for sonographers and usually requires a thorough understanding of complex anatomical structures. In this study, we propose a multi-task framework to learn the relationships among landmarks and structures jointly and automatically evaluate DDH. Our multi-task networks are equipped with three novel modules. Firstly, we adopt Mask R-CNN as the basic framework to detect and segment key anatomical structures and add one landmark detection branch to form a new multi-task framework. Secondly, we propose a novel shape similarity loss to refine the incomplete anatomical structure prediction robustly and accurately. Thirdly, we further incorporate the landmark-structure consistent prior to ensure the consistency of the bony rim estimated from the segmented structure and the detected landmark. In our experiments, 1,231 US images of the infant hip from 632 patients are collected, of which 247 images from 126 patients are tested. The average errors in alpha and beta angles are 2.221 degrees and 2.899 degrees. About 93% and 85% estimates of alpha and beta angles have errors less than 5 degrees, respectively. Experimental results demonstrate that the proposed method can accurately and robustly realize the automatic evaluation of DDH, showing great potential for clinical application.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge