Jikuan Qian
Multi-IMU with Online Self-Consistency for Freehand 3D Ultrasound Reconstruction
Jul 19, 2023
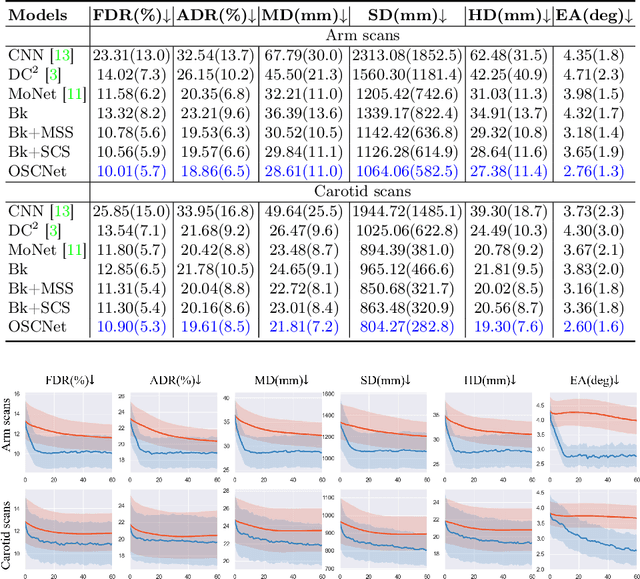
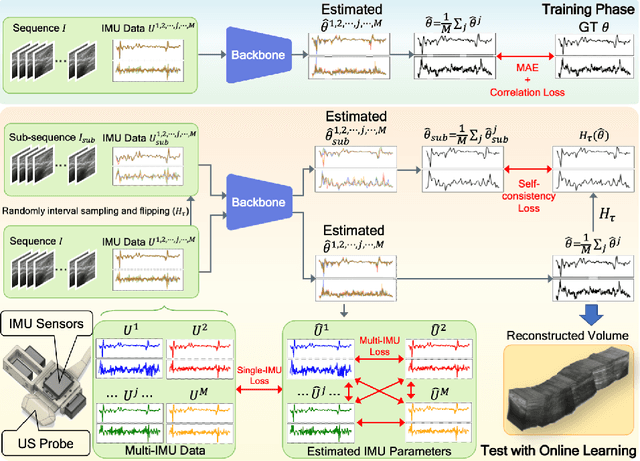
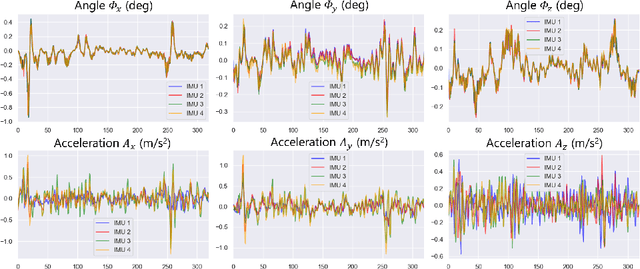
Abstract:Ultrasound (US) imaging is a popular tool in clinical diagnosis, offering safety, repeatability, and real-time capabilities. Freehand 3D US is a technique that provides a deeper understanding of scanned regions without increasing complexity. However, estimating elevation displacement and accumulation error remains challenging, making it difficult to infer the relative position using images alone. The addition of external lightweight sensors has been proposed to enhance reconstruction performance without adding complexity, which has been shown to be beneficial. We propose a novel online self-consistency network (OSCNet) using multiple inertial measurement units (IMUs) to improve reconstruction performance. OSCNet utilizes a modal-level self-supervised strategy to fuse multiple IMU information and reduce differences between reconstruction results obtained from each IMU data. Additionally, a sequence-level self-consistency strategy is proposed to improve the hierarchical consistency of prediction results among the scanning sequence and its sub-sequences. Experiments on large-scale arm and carotid datasets with multiple scanning tactics demonstrate that our OSCNet outperforms previous methods, achieving state-of-the-art reconstruction performance.
Agent with Tangent-based Formulation and Anatomical Perception for Standard Plane Localization in 3D Ultrasound
Jul 01, 2022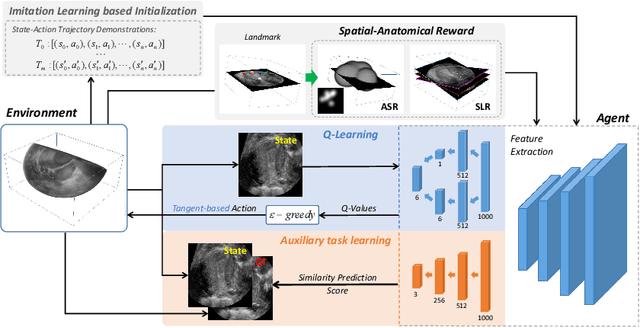
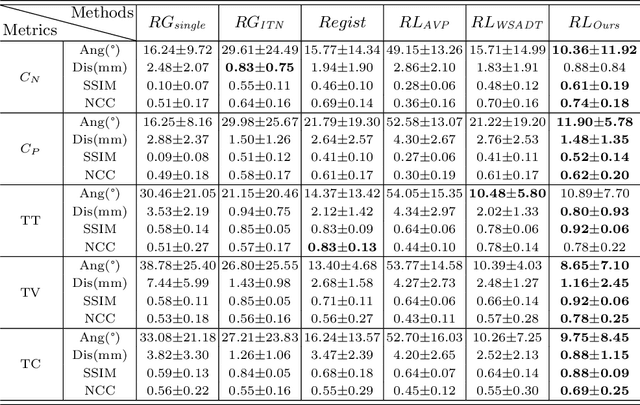
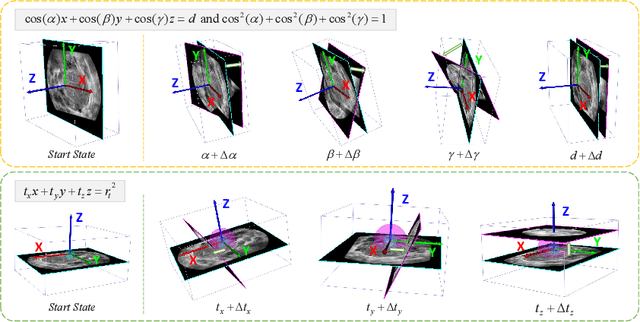
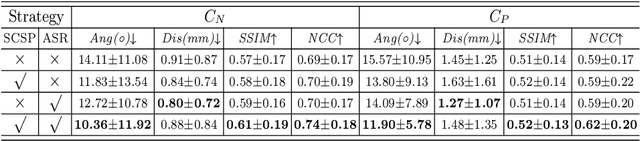
Abstract:Standard plane (SP) localization is essential in routine clinical ultrasound (US) diagnosis. Compared to 2D US, 3D US can acquire multiple view planes in one scan and provide complete anatomy with the addition of coronal plane. However, manually navigating SPs in 3D US is laborious and biased due to the orientation variability and huge search space. In this study, we introduce a novel reinforcement learning (RL) framework for automatic SP localization in 3D US. Our contribution is three-fold. First, we formulate SP localization in 3D US as a tangent-point-based problem in RL to restructure the action space and significantly reduce the search space. Second, we design an auxiliary task learning strategy to enhance the model's ability to recognize subtle differences crossing Non-SPs and SPs in plane search. Finally, we propose a spatial-anatomical reward to effectively guide learning trajectories by exploiting spatial and anatomical information simultaneously. We explore the efficacy of our approach on localizing four SPs on uterus and fetal brain datasets. The experiments indicate that our approach achieves a high localization accuracy as well as robust performance.
HASA: Hybrid Architecture Search with Aggregation Strategy for Echinococcosis Classification and Ovary Segmentation in Ultrasound Images
Apr 20, 2022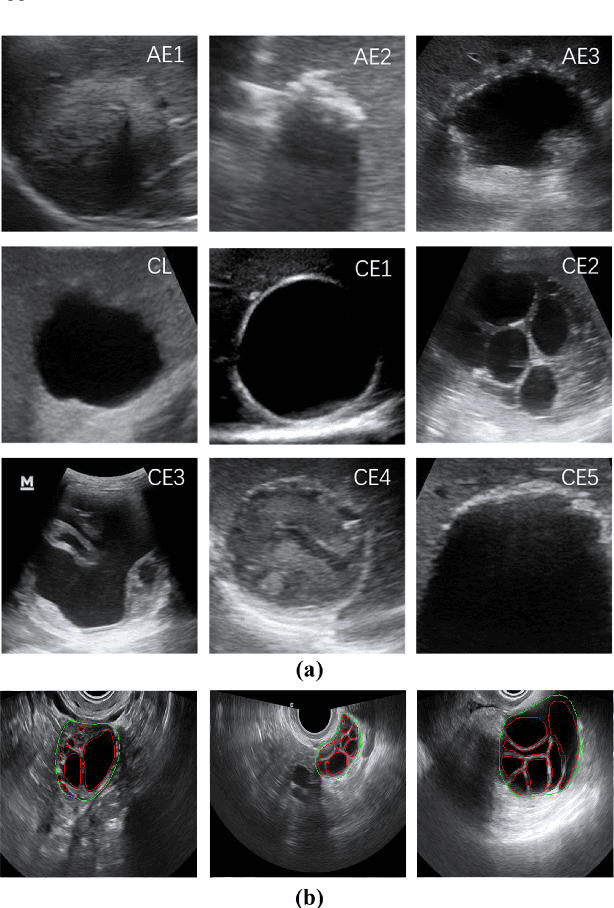
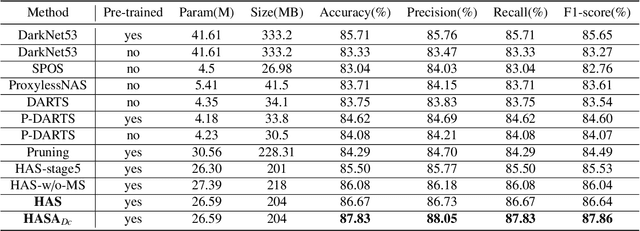
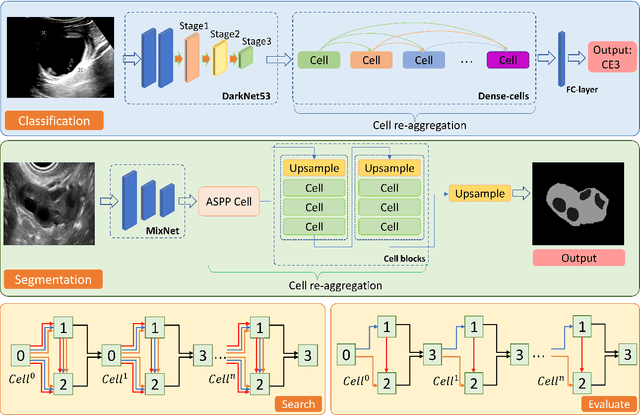
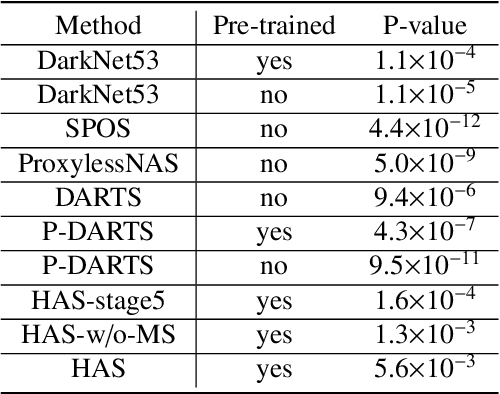
Abstract:Different from handcrafted features, deep neural networks can automatically learn task-specific features from data. Due to this data-driven nature, they have achieved remarkable success in various areas. However, manual design and selection of suitable network architectures are time-consuming and require substantial effort of human experts. To address this problem, researchers have proposed neural architecture search (NAS) algorithms which can automatically generate network architectures but suffer from heavy computational cost and instability if searching from scratch. In this paper, we propose a hybrid NAS framework for ultrasound (US) image classification and segmentation. The hybrid framework consists of a pre-trained backbone and several searched cells (i.e., network building blocks), which takes advantage of the strengths of both NAS and the expert knowledge from existing convolutional neural networks. Specifically, two effective and lightweight operations, a mixed depth-wise convolution operator and a squeeze-and-excitation block, are introduced into the candidate operations to enhance the variety and capacity of the searched cells. These two operations not only decrease model parameters but also boost network performance. Moreover, we propose a re-aggregation strategy for the searched cells, aiming to further improve the performance for different vision tasks. We tested our method on two large US image datasets, including a 9-class echinococcosis dataset containing 9566 images for classification and an ovary dataset containing 3204 images for segmentation. Ablation experiments and comparison with other handcrafted or automatically searched architectures demonstrate that our method can generate more powerful and lightweight models for the above US image classification and segmentation tasks.
Searching Collaborative Agents for Multi-plane Localization in 3D Ultrasound
May 22, 2021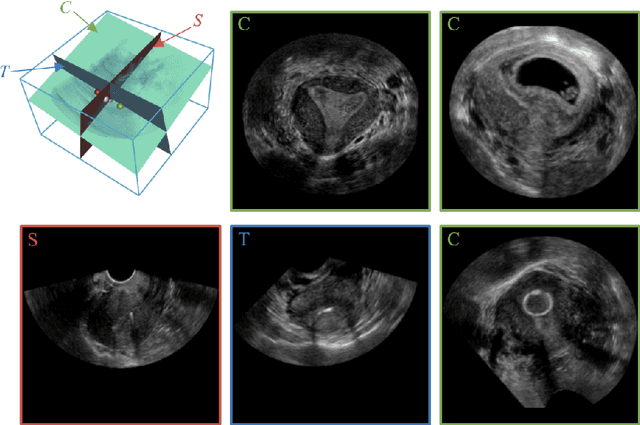
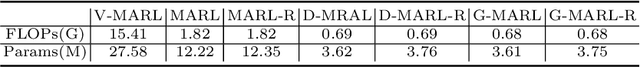
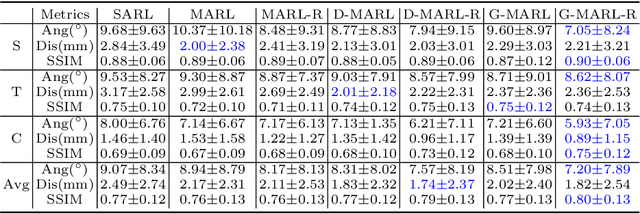
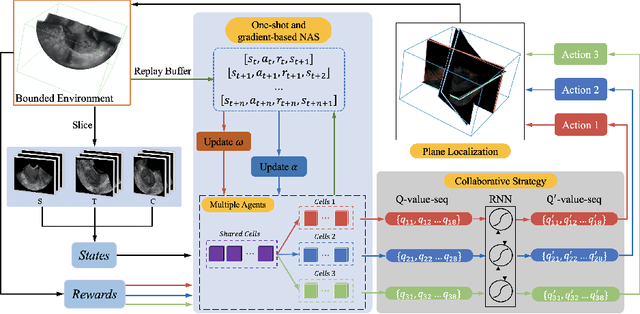

Abstract:3D ultrasound (US) has become prevalent due to its rich spatial and diagnostic information not contained in 2D US. Moreover, 3D US can contain multiple standard planes (SPs) in one shot. Thus, automatically localizing SPs in 3D US has the potential to improve user-independence and scanning-efficiency. However, manual SP localization in 3D US is challenging because of the low image quality, huge search space and large anatomical variability. In this work, we propose a novel multi-agent reinforcement learning (MARL) framework to simultaneously localize multiple SPs in 3D US. Our contribution is four-fold. First, our proposed method is general and it can accurately localize multiple SPs in different challenging US datasets. Second, we equip the MARL system with a recurrent neural network (RNN) based collaborative module, which can strengthen the communication among agents and learn the spatial relationship among planes effectively. Third, we explore to adopt the neural architecture search (NAS) to automatically design the network architecture of both the agents and the collaborative module. Last, we believe we are the first to realize automatic SP localization in pelvic US volumes, and note that our approach can handle both normal and abnormal uterus cases. Extensively validated on two challenging datasets of the uterus and fetal brain, our proposed method achieves the average localization accuracy of 7.03 degrees/1.59mm and 9.75 degrees/1.19mm. Experimental results show that our light-weight MARL model has higher accuracy than state-of-the-art methods.
Learn Fine-grained Adaptive Loss for Multiple Anatomical Landmark Detection in Medical Images
May 19, 2021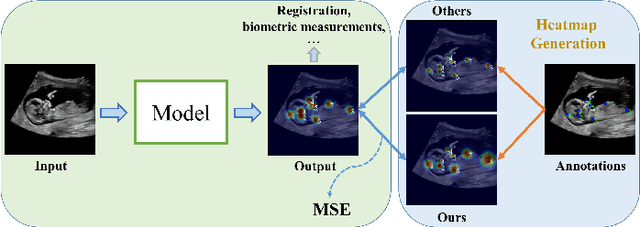
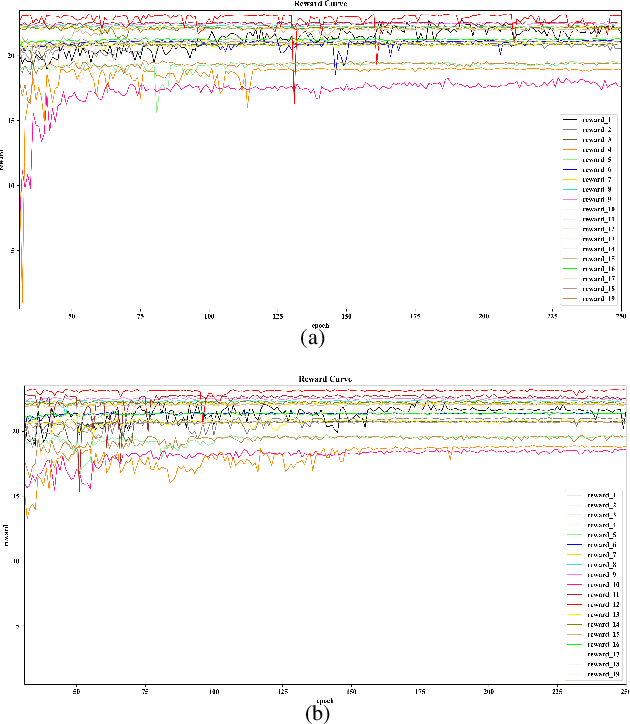
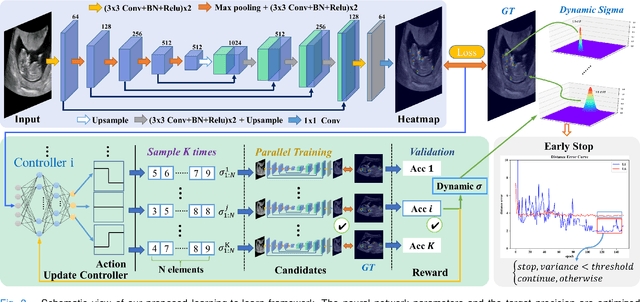
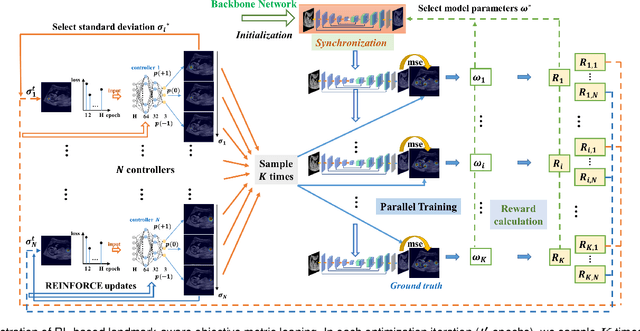
Abstract:Automatic and accurate detection of anatomical landmarks is an essential operation in medical image analysis with a multitude of applications. Recent deep learning methods have improved results by directly encoding the appearance of the captured anatomy with the likelihood maps (i.e., heatmaps). However, most current solutions overlook another essence of heatmap regression, the objective metric for regressing target heatmaps and rely on hand-crafted heuristics to set the target precision, thus being usually cumbersome and task-specific. In this paper, we propose a novel learning-to-learn framework for landmark detection to optimize the neural network and the target precision simultaneously. The pivot of this work is to leverage the reinforcement learning (RL) framework to search objective metrics for regressing multiple heatmaps dynamically during the training process, thus avoiding setting problem-specific target precision. We also introduce an early-stop strategy for active termination of the RL agent's interaction that adapts the optimal precision for separate targets considering exploration-exploitation tradeoffs. This approach shows better stability in training and improved localization accuracy in inference. Extensive experimental results on two different applications of landmark localization: 1) our in-house prenatal ultrasound (US) dataset and 2) the publicly available dataset of cephalometric X-Ray landmark detection, demonstrate the effectiveness of our proposed method. Our proposed framework is general and shows the potential to improve the efficiency of anatomical landmark detection.
Agent with Warm Start and Adaptive Dynamic Termination for Plane Localization in 3D Ultrasound
Mar 26, 2021
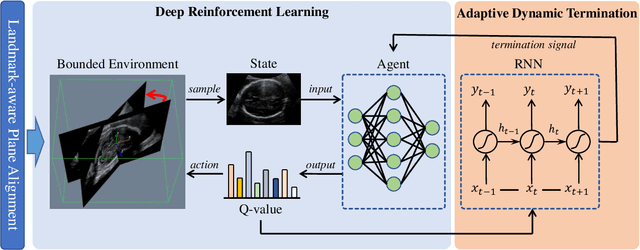
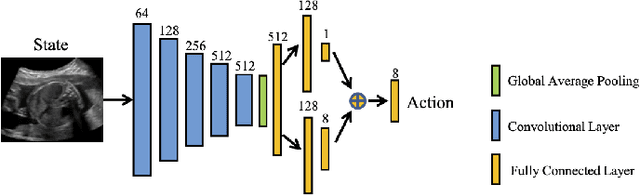
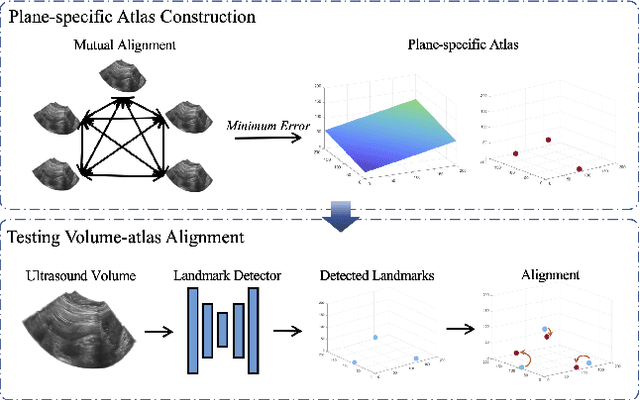
Abstract:Accurate standard plane (SP) localization is the fundamental step for prenatal ultrasound (US) diagnosis. Typically, dozens of US SPs are collected to determine the clinical diagnosis. 2D US has to perform scanning for each SP, which is time-consuming and operator-dependent. While 3D US containing multiple SPs in one shot has the inherent advantages of less user-dependency and more efficiency. Automatically locating SP in 3D US is very challenging due to the huge search space and large fetal posture variations. Our previous study proposed a deep reinforcement learning (RL) framework with an alignment module and active termination to localize SPs in 3D US automatically. However, termination of agent search in RL is important and affects the practical deployment. In this study, we enhance our previous RL framework with a newly designed adaptive dynamic termination to enable an early stop for the agent searching, saving at most 67% inference time, thus boosting the accuracy and efficiency of the RL framework at the same time. Besides, we validate the effectiveness and generalizability of our algorithm extensively on our in-house multi-organ datasets containing 433 fetal brain volumes, 519 fetal abdomen volumes, and 683 uterus volumes. Our approach achieves localization error of 2.52mm/10.26 degrees, 2.48mm/10.39 degrees, 2.02mm/10.48 degrees, 2.00mm/14.57 degrees, 2.61mm/9.71 degrees, 3.09mm/9.58 degrees, 1.49mm/7.54 degrees for the transcerebellar, transventricular, transthalamic planes in fetal brain, abdominal plane in fetal abdomen, and mid-sagittal, transverse and coronal planes in uterus, respectively. Experimental results show that our method is general and has the potential to improve the efficiency and standardization of US scanning.
FetusMap: Fetal Pose Estimation in 3D Ultrasound
Oct 11, 2019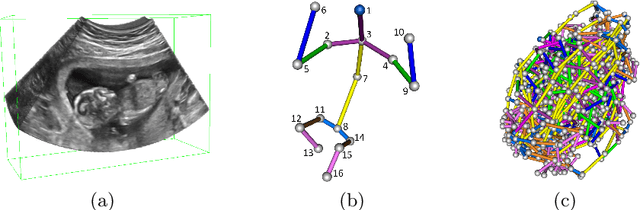
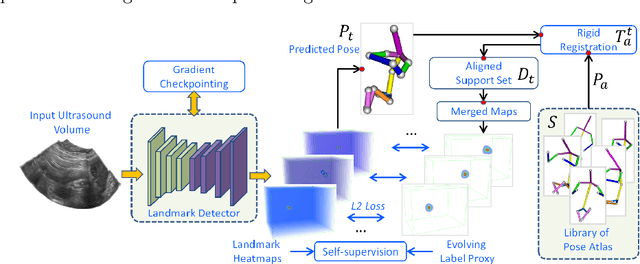
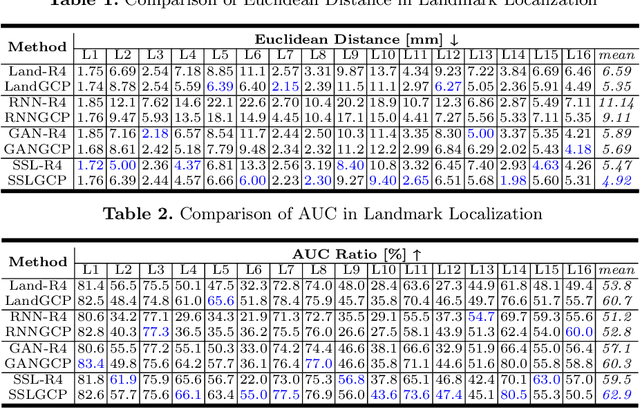
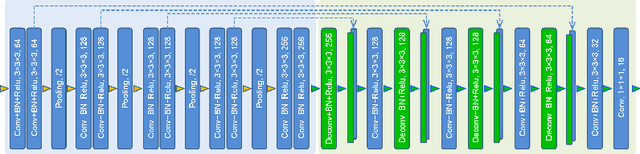
Abstract:The 3D ultrasound (US) entrance inspires a multitude of automated prenatal examinations. However, studies about the structuralized description of the whole fetus in 3D US are still rare. In this paper, we propose to estimate the 3D pose of fetus in US volumes to facilitate its quantitative analyses in global and local scales. Given the great challenges in 3D US, including the high volume dimension, poor image quality, symmetric ambiguity in anatomical structures and large variations of fetal pose, our contribution is three-fold. (i) This is the first work about 3D pose estimation of fetus in the literature. We aim to extract the skeleton of whole fetus and assign different segments/joints with correct torso/limb labels. (ii) We propose a self-supervised learning (SSL) framework to finetune the deep network to form visually plausible pose predictions. Specifically, we leverage the landmark-based registration to effectively encode case-adaptive anatomical priors and generate evolving label proxy for supervision. (iii) To enable our 3D network perceive better contextual cues with higher resolution input under limited computing resource, we further adopt the gradient check-pointing (GCP) strategy to save GPU memory and improve the prediction. Extensively validated on a large 3D US dataset, our method tackles varying fetal poses and achieves promising results. 3D pose estimation of fetus has potentials in serving as a map to provide navigation for many advanced studies.
Agent with Warm Start and Active Termination for Plane Localization in 3D Ultrasound
Oct 10, 2019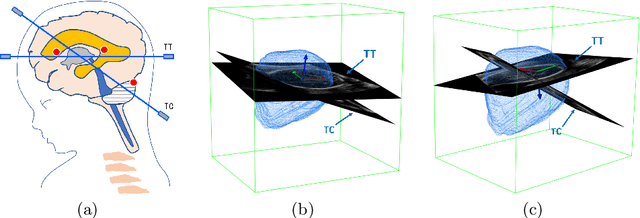
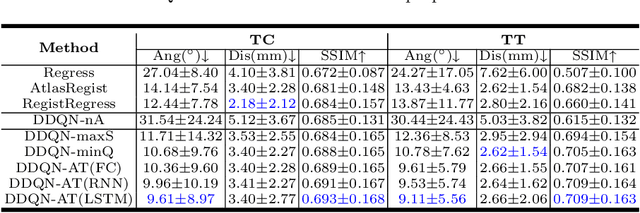
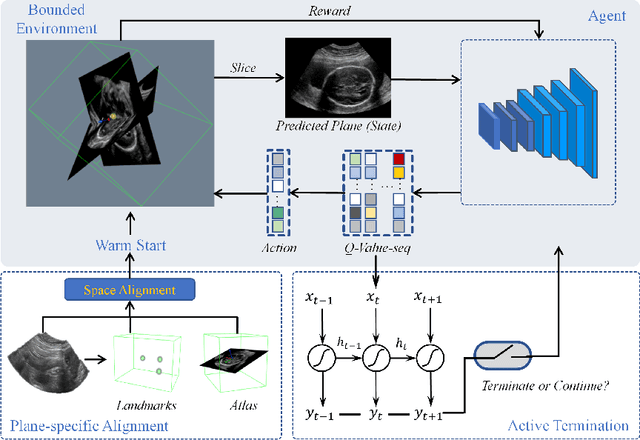
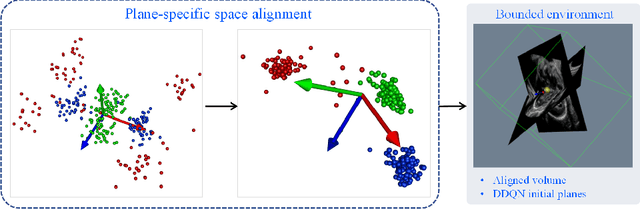
Abstract:Standard plane localization is crucial for ultrasound (US) diagnosis. In prenatal US, dozens of standard planes are manually acquired with a 2D probe. It is time-consuming and operator-dependent. In comparison, 3D US containing multiple standard planes in one shot has the inherent advantages of less user-dependency and more efficiency. However, manual plane localization in US volume is challenging due to the huge search space and large fetal posture variation. In this study, we propose a novel reinforcement learning (RL) framework to automatically localize fetal brain standard planes in 3D US. Our contribution is two-fold. First, we equip the RL framework with a landmark-aware alignment module to provide warm start and strong spatial bounds for the agent actions, thus ensuring its effectiveness. Second, instead of passively and empirically terminating the agent inference, we propose a recurrent neural network based strategy for active termination of the agent's interaction procedure. This improves both the accuracy and efficiency of the localization system. Extensively validated on our in-house large dataset, our approach achieves the accuracy of 3.4mm/9.6{\deg} and 2.7mm/9.1{\deg} for the transcerebellar and transthalamic plane localization, respectively. Ourproposed RL framework is general and has the potential to improve the efficiency and standardization of US scanning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge