Juzheng Miao
Perceive and Calibrate: Analyzing and Enhancing Robustness of Medical Multi-Modal Large Language Models
Dec 26, 2025Abstract:Medical Multi-modal Large Language Models (MLLMs) have shown promising clinical performance. However, their sensitivity to real-world input perturbations, such as imaging artifacts and textual errors, critically undermines their clinical applicability. Systematic analysis of such noise impact on medical MLLMs remains largely unexplored. Furthermore, while several works have investigated the MLLMs' robustness in general domains, they primarily focus on text modality and rely on costly fine-tuning. They are inadequate to address the complex noise patterns and fulfill the strict safety standards in medicine. To bridge this gap, this work systematically analyzes the impact of various perturbations on medical MLLMs across both visual and textual modalities. Building on our findings, we introduce a training-free Inherent-enhanced Multi-modal Calibration (IMC) framework that leverages MLLMs' inherent denoising capabilities following the perceive-and-calibrate principle for cross-modal robustness enhancement. For the visual modality, we propose a Perturbation-aware Denoising Calibration (PDC) which leverages MLLMs' own vision encoder to identify noise patterns and perform prototype-guided feature calibration. For text denoising, we design a Self-instantiated Multi-agent System (SMS) that exploits the MLLMs' self-assessment capabilities to refine noisy text through a cooperative hierarchy of agents. We construct a benchmark containing 11 types of noise across both image and text modalities on 2 datasets. Experimental results demonstrate our method achieves the state-of-the-art performance across multiple modalities, showing potential to enhance MLLMs' robustness in real clinical scenarios.
Medical Large Vision Language Models with Multi-Image Visual Ability
May 25, 2025Abstract:Medical large vision-language models (LVLMs) have demonstrated promising performance across various single-image question answering (QA) benchmarks, yet their capability in processing multi-image clinical scenarios remains underexplored. Unlike single image based tasks, medical tasks involving multiple images often demand sophisticated visual understanding capabilities, such as temporal reasoning and cross-modal analysis, which are poorly supported by current medical LVLMs. To bridge this critical gap, we present the Med-MIM instruction dataset, comprising 83.2K medical multi-image QA pairs that span four types of multi-image visual abilities (temporal understanding, reasoning, comparison, co-reference). Using this dataset, we fine-tune Mantis and LLaVA-Med, resulting in two specialized medical VLMs: MIM-LLaVA-Med and Med-Mantis, both optimized for multi-image analysis. Additionally, we develop the Med-MIM benchmark to comprehensively evaluate the medical multi-image understanding capabilities of LVLMs. We assess eight popular LVLMs, including our two models, on the Med-MIM benchmark. Experimental results show that both Med-Mantis and MIM-LLaVA-Med achieve superior performance on the held-in and held-out subsets of the Med-MIM benchmark, demonstrating that the Med-MIM instruction dataset effectively enhances LVLMs' multi-image understanding capabilities in the medical domain.
FM-OSD: Foundation Model-Enabled One-Shot Detection of Anatomical Landmarks
Jul 07, 2024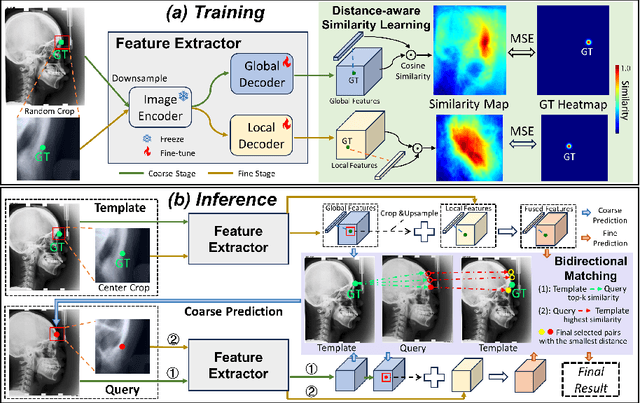
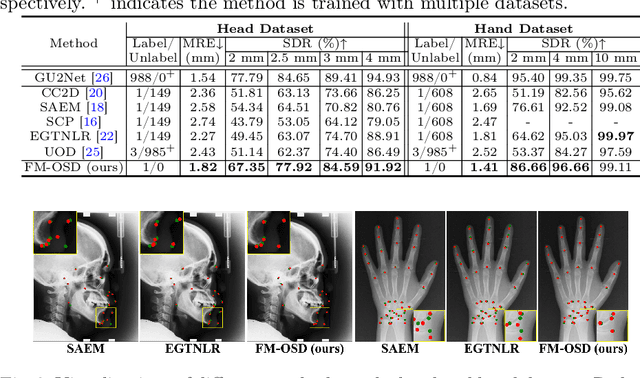
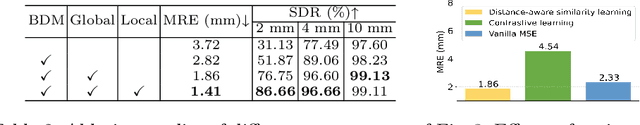
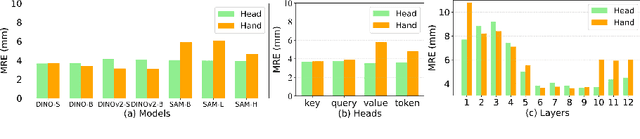
Abstract:One-shot detection of anatomical landmarks is gaining significant attention for its efficiency in using minimal labeled data to produce promising results. However, the success of current methods heavily relies on the employment of extensive unlabeled data to pre-train an effective feature extractor, which limits their applicability in scenarios where a substantial amount of unlabeled data is unavailable. In this paper, we propose the first foundation model-enabled one-shot landmark detection (FM-OSD) framework for accurate landmark detection in medical images by utilizing solely a single template image without any additional unlabeled data. Specifically, we use the frozen image encoder of visual foundation models as the feature extractor, and introduce dual-branch global and local feature decoders to increase the resolution of extracted features in a coarse to fine manner. The introduced feature decoders are efficiently trained with a distance-aware similarity learning loss to incorporate domain knowledge from the single template image. Moreover, a novel bidirectional matching strategy is developed to improve both robustness and accuracy of landmark detection in the case of scattered similarity map obtained by foundation models. We validate our method on two public anatomical landmark detection datasets. By using solely a single template image, our method demonstrates significant superiority over strong state-of-the-art one-shot landmark detection methods.
Cross Prompting Consistency with Segment Anything Model for Semi-supervised Medical Image Segmentation
Jul 07, 2024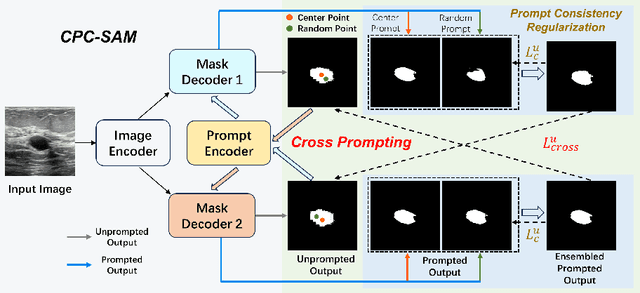
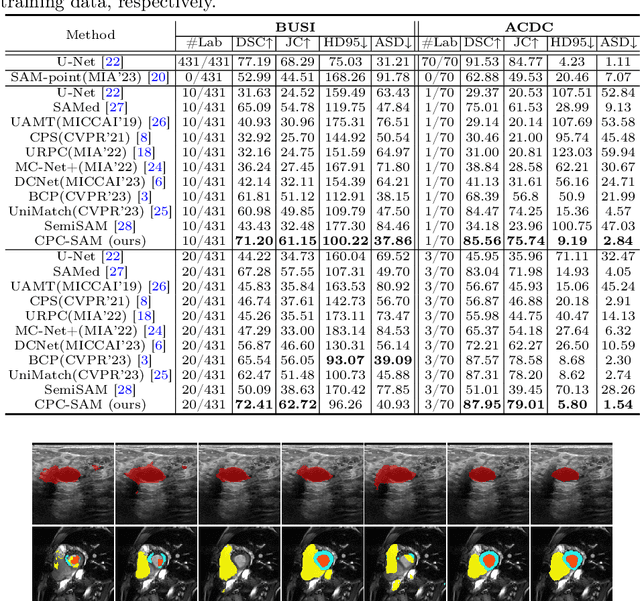
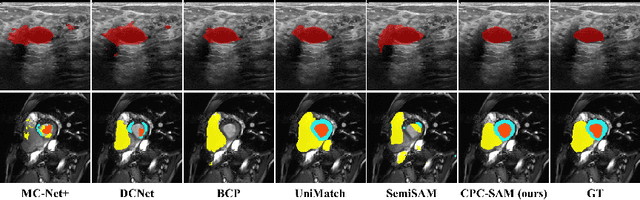
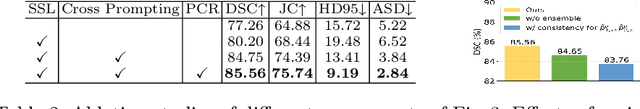
Abstract:Semi-supervised learning (SSL) has achieved notable progress in medical image segmentation. To achieve effective SSL, a model needs to be able to efficiently learn from limited labeled data and effectively exploiting knowledge from abundant unlabeled data. Recent developments in visual foundation models, such as the Segment Anything Model (SAM), have demonstrated remarkable adaptability with improved sample efficiency. To harness the power of foundation models for application in SSL, we propose a cross prompting consistency method with segment anything model (CPC-SAM) for semi-supervised medical image segmentation. Our method employs SAM's unique prompt design and innovates a cross-prompting strategy within a dual-branch framework to automatically generate prompts and supervisions across two decoder branches, enabling effectively learning from both scarce labeled and valuable unlabeled data. We further design a novel prompt consistency regularization, to reduce the prompt position sensitivity and to enhance the output invariance under different prompts. We validate our method on two medical image segmentation tasks. The extensive experiments with different labeled-data ratios and modalities demonstrate the superiority of our proposed method over the state-of-the-art SSL methods, with more than 9% Dice improvement on the breast cancer segmentation task.
Emulating Full Client Participation: A Long-Term Client Selection Strategy for Federated Learning
May 22, 2024



Abstract:Client selection significantly affects the system convergence efficiency and is a crucial problem in federated learning. Existing methods often select clients by evaluating each round individually and overlook the necessity for long-term optimization, resulting in suboptimal performance and potential fairness issues. In this study, we propose a novel client selection strategy designed to emulate the performance achieved with full client participation. In a single round, we select clients by minimizing the gradient-space estimation error between the client subset and the full client set. In multi-round selection, we introduce a novel individual fairness constraint, which ensures that clients with similar data distributions have similar frequencies of being selected. This constraint guides the client selection process from a long-term perspective. We employ Lyapunov optimization and submodular functions to efficiently identify the optimal subset of clients, and provide a theoretical analysis of the convergence ability. Experiments demonstrate that the proposed strategy significantly improves both accuracy and fairness compared to previous methods while also exhibiting efficiency by incurring minimal time overhead.
CauDR: A Causality-inspired Domain Generalization Framework for Fundus-based Diabetic Retinopathy Grading
Sep 27, 2023Abstract:Diabetic retinopathy (DR) is the most common diabetic complication, which usually leads to retinal damage, vision loss, and even blindness. A computer-aided DR grading system has a significant impact on helping ophthalmologists with rapid screening and diagnosis. Recent advances in fundus photography have precipitated the development of novel retinal imaging cameras and their subsequent implementation in clinical practice. However, most deep learning-based algorithms for DR grading demonstrate limited generalization across domains. This inferior performance stems from variance in imaging protocols and devices inducing domain shifts. We posit that declining model performance between domains arises from learning spurious correlations in the data. Incorporating do-operations from causality analysis into model architectures may mitigate this issue and improve generalizability. Specifically, a novel universal structural causal model (SCM) was proposed to analyze spurious correlations in fundus imaging. Building on this, a causality-inspired diabetic retinopathy grading framework named CauDR was developed to eliminate spurious correlations and achieve more generalizable DR diagnostics. Furthermore, existing datasets were reorganized into 4DR benchmark for DG scenario. Results demonstrate the effectiveness and the state-of-the-art (SOTA) performance of CauDR.
MA-SAM: Modality-agnostic SAM Adaptation for 3D Medical Image Segmentation
Sep 16, 2023



Abstract:The Segment Anything Model (SAM), a foundation model for general image segmentation, has demonstrated impressive zero-shot performance across numerous natural image segmentation tasks. However, SAM's performance significantly declines when applied to medical images, primarily due to the substantial disparity between natural and medical image domains. To effectively adapt SAM to medical images, it is important to incorporate critical third-dimensional information, i.e., volumetric or temporal knowledge, during fine-tuning. Simultaneously, we aim to harness SAM's pre-trained weights within its original 2D backbone to the fullest extent. In this paper, we introduce a modality-agnostic SAM adaptation framework, named as MA-SAM, that is applicable to various volumetric and video medical data. Our method roots in the parameter-efficient fine-tuning strategy to update only a small portion of weight increments while preserving the majority of SAM's pre-trained weights. By injecting a series of 3D adapters into the transformer blocks of the image encoder, our method enables the pre-trained 2D backbone to extract third-dimensional information from input data. The effectiveness of our method has been comprehensively evaluated on four medical image segmentation tasks, by using 10 public datasets across CT, MRI, and surgical video data. Remarkably, without using any prompt, our method consistently outperforms various state-of-the-art 3D approaches, surpassing nnU-Net by 0.9%, 2.6%, and 9.9% in Dice for CT multi-organ segmentation, MRI prostate segmentation, and surgical scene segmentation respectively. Our model also demonstrates strong generalization, and excels in challenging tumor segmentation when prompts are used. Our code is available at: https://github.com/cchen-cc/MA-SAM.
FFCNet: Fourier Transform-Based Frequency Learning and Complex Convolutional Network for Colon Disease Classification
Jul 04, 2022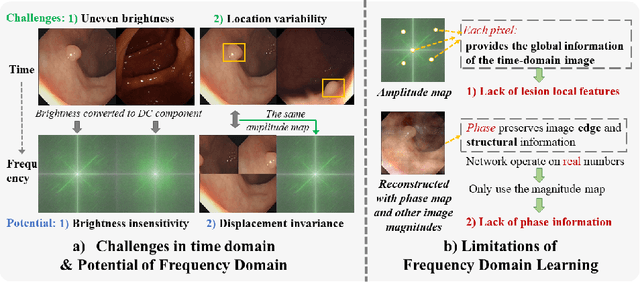
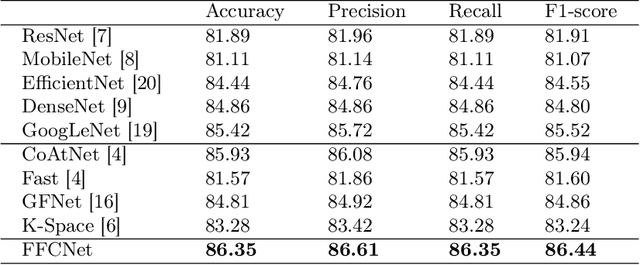
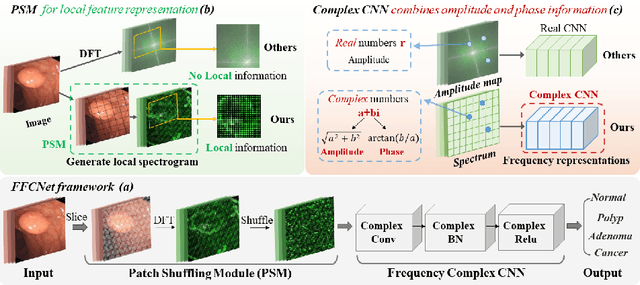
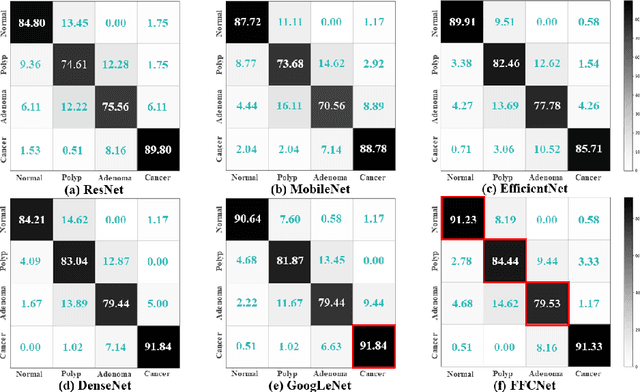
Abstract:Reliable automatic classification of colonoscopy images is of great significance in assessing the stage of colonic lesions and formulating appropriate treatment plans. However, it is challenging due to uneven brightness, location variability, inter-class similarity, and intra-class dissimilarity, affecting the classification accuracy. To address the above issues, we propose a Fourier-based Frequency Complex Network (FFCNet) for colon disease classification in this study. Specifically, FFCNet is a novel complex network that enables the combination of complex convolutional networks with frequency learning to overcome the loss of phase information caused by real convolution operations. Also, our Fourier transform transfers the average brightness of an image to a point in the spectrum (the DC component), alleviating the effects of uneven brightness by decoupling image content and brightness. Moreover, the image patch scrambling module in FFCNet generates random local spectral blocks, empowering the network to learn long-range and local diseasespecific features and improving the discriminative ability of hard samples. We evaluated the proposed FFCNet on an in-house dataset with 2568 colonoscopy images, showing our method achieves high performance outperforming previous state-of-the art methods with an accuracy of 86:35% and an accuracy of 4.46% higher than the backbone. The project page with code is available at https://github.com/soleilssss/FFCNet.
Fine-grained Correlation Loss for Regression
Jul 01, 2022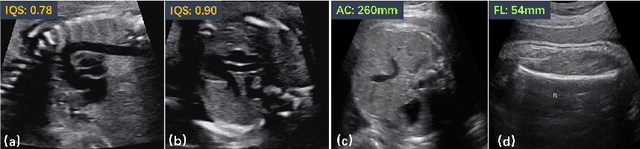
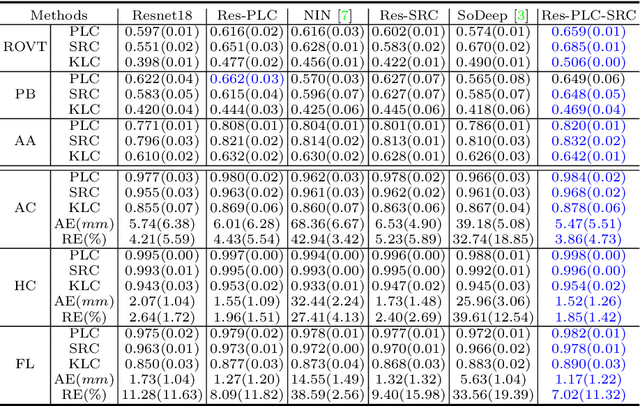
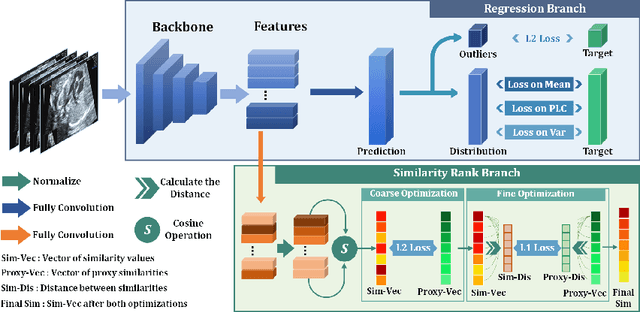
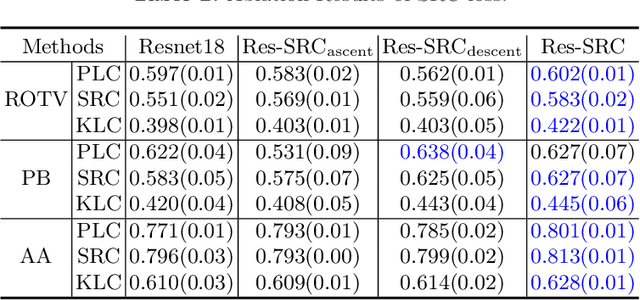
Abstract:Regression learning is classic and fundamental for medical image analysis. It provides the continuous mapping for many critical applications, like the attribute estimation, object detection, segmentation and non-rigid registration. However, previous studies mainly took the case-wise criteria, like the mean square errors, as the optimization objectives. They ignored the very important population-wise correlation criterion, which is exactly the final evaluation metric in many tasks. In this work, we propose to revisit the classic regression tasks with novel investigations on directly optimizing the fine-grained correlation losses. We mainly explore two complementary correlation indexes as learnable losses: Pearson linear correlation (PLC) and Spearman rank correlation (SRC). The contributions of this paper are two folds. First, for the PLC on global level, we propose a strategy to make it robust against the outliers and regularize the key distribution factors. These efforts significantly stabilize the learning and magnify the efficacy of PLC. Second, for the SRC on local level, we propose a coarse-to-fine scheme to ease the learning of the exact ranking order among samples. Specifically, we convert the learning for the ranking of samples into the learning of similarity relationships among samples. We extensively validate our method on two typical ultrasound image regression tasks, including the image quality assessment and bio-metric measurement. Experiments prove that, with the fine-grained guidance in directly optimizing the correlation, the regression performances are significantly improved. Our proposed correlation losses are general and can be extended to more important applications.
AWSnet: An Auto-weighted Supervision Attention Network for Myocardial Scar and Edema Segmentation in Multi-sequence Cardiac Magnetic Resonance Images
Jan 14, 2022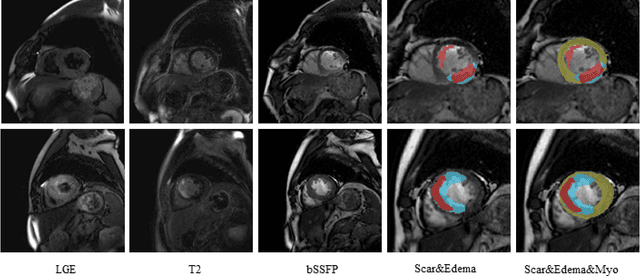
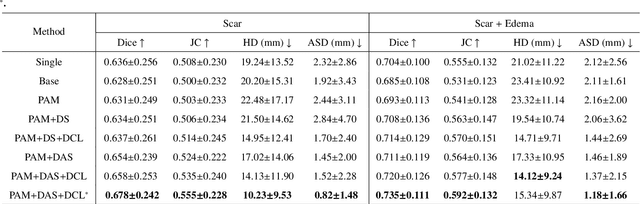
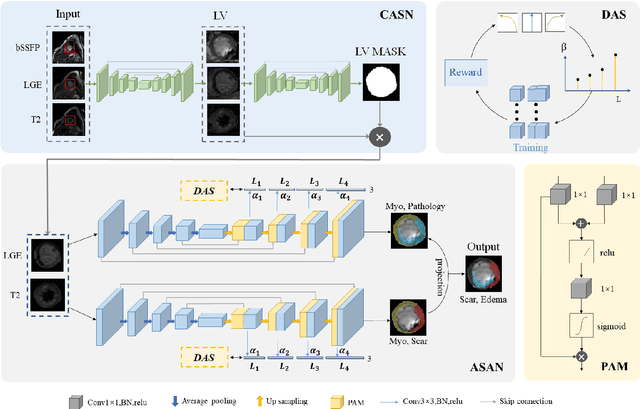
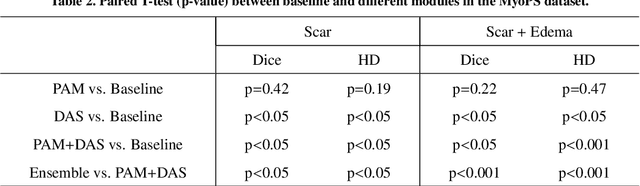
Abstract:Multi-sequence cardiac magnetic resonance (CMR) provides essential pathology information (scar and edema) to diagnose myocardial infarction. However, automatic pathology segmentation can be challenging due to the difficulty of effectively exploring the underlying information from the multi-sequence CMR data. This paper aims to tackle the scar and edema segmentation from multi-sequence CMR with a novel auto-weighted supervision framework, where the interactions among different supervised layers are explored under a task-specific objective using reinforcement learning. Furthermore, we design a coarse-to-fine framework to boost the small myocardial pathology region segmentation with shape prior knowledge. The coarse segmentation model identifies the left ventricle myocardial structure as a shape prior, while the fine segmentation model integrates a pixel-wise attention strategy with an auto-weighted supervision model to learn and extract salient pathological structures from the multi-sequence CMR data. Extensive experimental results on a publicly available dataset from Myocardial pathology segmentation combining multi-sequence CMR (MyoPS 2020) demonstrate our method can achieve promising performance compared with other state-of-the-art methods. Our method is promising in advancing the myocardial pathology assessment on multi-sequence CMR data. To motivate the community, we have made our code publicly available via https://github.com/soleilssss/AWSnet/tree/master.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge