Xiahai Zhuang
Few-Shot Video Object Segmentation in X-Ray Angiography Using Local Matching and Spatio-Temporal Consistency Loss
Jan 02, 2026Abstract:We introduce a novel FSVOS model that employs a local matching strategy to restrict the search space to the most relevant neighboring pixels. Rather than relying on inefficient standard im2col-like implementations (e.g., spatial convolutions, depthwise convolutions and feature-shifting mechanisms) or hardware-specific CUDA kernels (e.g., deformable and neighborhood attention), which often suffer from limited portability across non-CUDA devices, we reorganize the local sampling process through a direction-based sampling perspective. Specifically, we implement a non-parametric sampling mechanism that enables dynamically varying sampling regions. This approach provides the flexibility to adapt to diverse spatial structures without the computational costs of parametric layers and the need for model retraining. To further enhance feature coherence across frames, we design a supervised spatio-temporal contrastive learning scheme that enforces consistency in feature representations. In addition, we introduce a publicly available benchmark dataset for multi-object segmentation in X-ray angiography videos (MOSXAV), featuring detailed, manually labeled segmentation ground truth. Extensive experiments on the CADICA, XACV, and MOSXAV datasets show that our proposed FSVOS method outperforms current state-of-the-art video segmentation methods in terms of segmentation accuracy and generalization capability (i.e., seen and unseen categories). This work offers enhanced flexibility and potential for a wide range of clinical applications.
Liver Fibrosis Quantification and Analysis: The LiQA Dataset and Baseline Method
Dec 22, 2025Abstract:Liver fibrosis represents a significant global health burden, necessitating accurate staging for effective clinical management. This report introduces the LiQA (Liver Fibrosis Quantification and Analysis) dataset, established as part of the CARE 2024 challenge. Comprising $440$ patients with multi-phase, multi-center MRI scans, the dataset is curated to benchmark algorithms for Liver Segmentation (LiSeg) and Liver Fibrosis Staging (LiFS) under complex real-world conditions, including domain shifts, missing modalities, and spatial misalignment. We further describe the challenge's top-performing methodology, which integrates a semi-supervised learning framework with external data for robust segmentation, and utilizes a multi-view consensus approach with Class Activation Map (CAM)-based regularization for staging. Evaluation of this baseline demonstrates that leveraging multi-source data and anatomical constraints significantly enhances model robustness in clinical settings.
CineMyoPS: Segmenting Myocardial Pathologies from Cine Cardiac MR
Jul 03, 2025
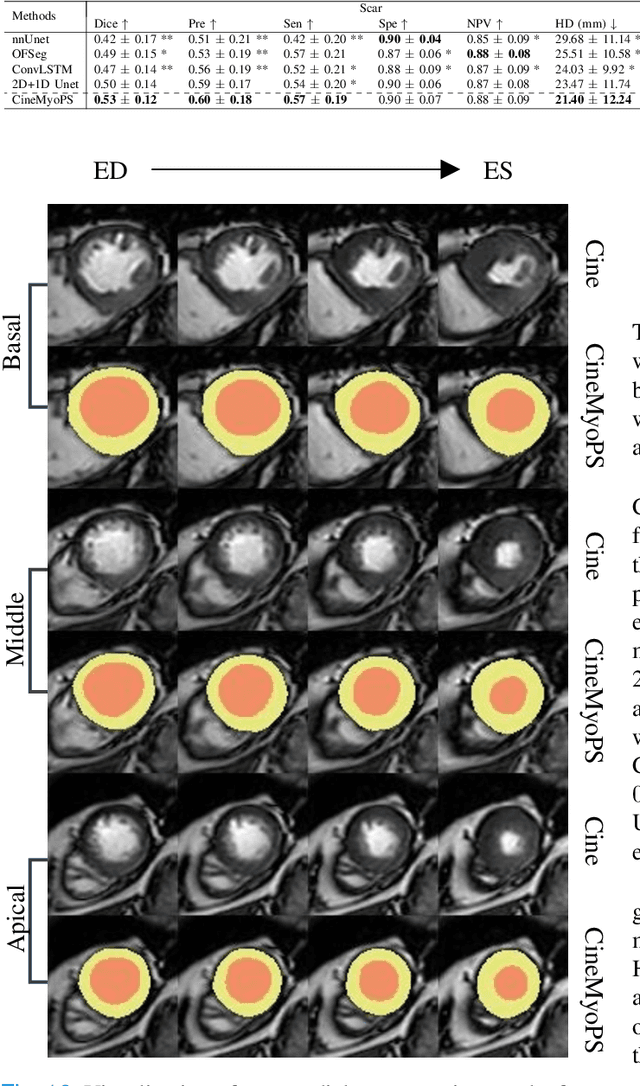


Abstract:Myocardial infarction (MI) is a leading cause of death worldwide. Late gadolinium enhancement (LGE) and T2-weighted cardiac magnetic resonance (CMR) imaging can respectively identify scarring and edema areas, both of which are essential for MI risk stratification and prognosis assessment. Although combining complementary information from multi-sequence CMR is useful, acquiring these sequences can be time-consuming and prohibitive, e.g., due to the administration of contrast agents. Cine CMR is a rapid and contrast-free imaging technique that can visualize both motion and structural abnormalities of the myocardium induced by acute MI. Therefore, we present a new end-to-end deep neural network, referred to as CineMyoPS, to segment myocardial pathologies, \ie scars and edema, solely from cine CMR images. Specifically, CineMyoPS extracts both motion and anatomy features associated with MI. Given the interdependence between these features, we design a consistency loss (resembling the co-training strategy) to facilitate their joint learning. Furthermore, we propose a time-series aggregation strategy to integrate MI-related features across the cardiac cycle, thereby enhancing segmentation accuracy for myocardial pathologies. Experimental results on a multi-center dataset demonstrate that CineMyoPS achieves promising performance in myocardial pathology segmentation, motion estimation, and anatomy segmentation.
Learning Concept-Driven Logical Rules for Interpretable and Generalizable Medical Image Classification
May 20, 2025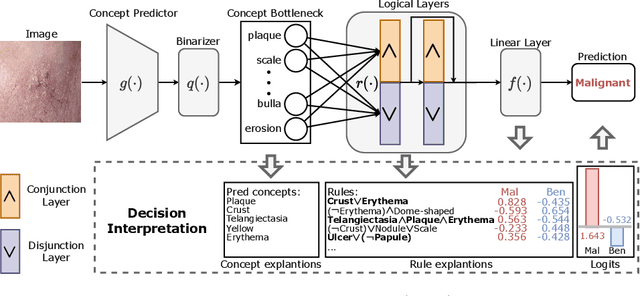
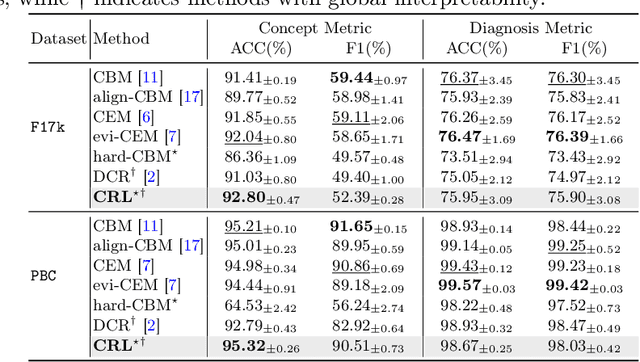
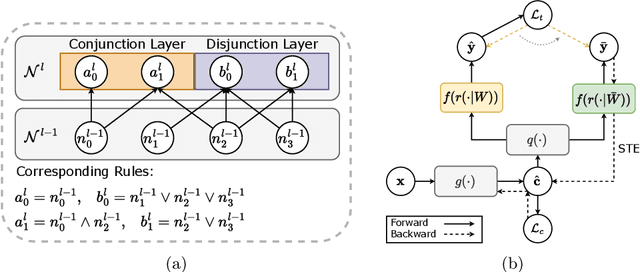
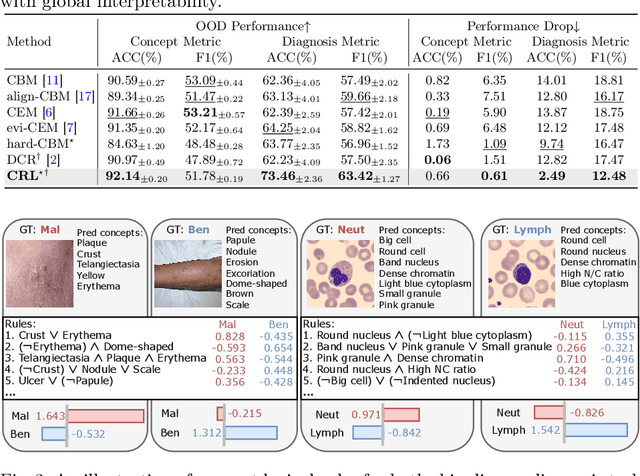
Abstract:The pursuit of decision safety in clinical applications highlights the potential of concept-based methods in medical imaging. While these models offer active interpretability, they often suffer from concept leakages, where unintended information within soft concept representations undermines both interpretability and generalizability. Moreover, most concept-based models focus solely on local explanations (instance-level), neglecting the global decision logic (dataset-level). To address these limitations, we propose Concept Rule Learner (CRL), a novel framework to learn Boolean logical rules from binarized visual concepts. CRL employs logical layers to capture concept correlations and extract clinically meaningful rules, thereby providing both local and global interpretability. Experiments on two medical image classification tasks show that CRL achieves competitive performance with existing methods while significantly improving generalizability to out-of-distribution data. The code of our work is available at https://github.com/obiyoag/crl.
Empowering Medical Multi-Agents with Clinical Consultation Flow for Dynamic Diagnosis
Mar 19, 2025



Abstract:Traditional AI-based healthcare systems often rely on single-modal data, limiting diagnostic accuracy due to incomplete information. However, recent advancements in foundation models show promising potential for enhancing diagnosis combining multi-modal information. While these models excel in static tasks, they struggle with dynamic diagnosis, failing to manage multi-turn interactions and often making premature diagnostic decisions due to insufficient persistence in information collection.To address this, we propose a multi-agent framework inspired by consultation flow and reinforcement learning (RL) to simulate the entire consultation process, integrating multiple clinical information for effective diagnosis. Our approach incorporates a hierarchical action set, structured from clinic consultation flow and medical textbook, to effectively guide the decision-making process. This strategy improves agent interactions, enabling them to adapt and optimize actions based on the dynamic state. We evaluated our framework on a public dynamic diagnosis benchmark. The proposed framework evidentially improves the baseline methods and achieves state-of-the-art performance compared to existing foundation model-based methods.
Towards Universal Learning-based Model for Cardiac Image Reconstruction: Summary of the CMRxRecon2024 Challenge
Mar 05, 2025Abstract:Cardiovascular magnetic resonance (CMR) offers diverse imaging contrasts for assessment of cardiac function and tissue characterization. However, acquiring each single CMR modality is often time-consuming, and comprehensive clinical protocols require multiple modalities with various sampling patterns, further extending the overall acquisition time and increasing susceptibility to motion artifacts. Existing deep learning-based reconstruction methods are often designed for specific acquisition parameters, which limits their ability to generalize across a variety of scan scenarios. As part of the CMRxRecon Series, the CMRxRecon2024 challenge provides diverse datasets encompassing multi-modality multi-view imaging with various sampling patterns, and a platform for the international community to develop and benchmark reconstruction solutions in two well-crafted tasks. Task 1 is a modality-universal setting, evaluating the out-of-distribution generalization of the reconstructed model, while Task 2 follows sampling-universal setting assessing the one-for-all adaptability of the universal model. Main contributions include providing the first and largest publicly available multi-modality, multi-view cardiac k-space dataset; developing a benchmarking platform that simulates clinical acceleration protocols, with a shared code library and tutorial for various k-t undersampling patterns and data processing; giving technical insights of enhanced data consistency based on physic-informed networks and adaptive prompt-learning embedding to be versatile to different clinical settings; additional finding on evaluation metrics to address the limitations of conventional ground-truth references in universal reconstruction tasks.
Contrast-Free Myocardial Scar Segmentation in Cine MRI using Motion and Texture Fusion
Jan 09, 2025Abstract:Late gadolinium enhancement MRI (LGE MRI) is the gold standard for the detection of myocardial scars for post myocardial infarction (MI). LGE MRI requires the injection of a contrast agent, which carries potential side effects and increases scanning time and patient discomfort. To address these issues, we propose a novel framework that combines cardiac motion observed in cine MRI with image texture information to segment the myocardium and scar tissue in the left ventricle. Cardiac motion tracking can be formulated as a full cardiac image cycle registration problem, which can be solved via deep neural networks. Experimental results prove that the proposed method can achieve scar segmentation based on non-contrasted cine images with comparable accuracy to LGE MRI. This demonstrates its potential as an alternative to contrast-enhanced techniques for scar detection.
InDeed: Interpretable image deep decomposition with guaranteed generalizability
Jan 02, 2025



Abstract:Image decomposition aims to analyze an image into elementary components, which is essential for numerous downstream tasks and also by nature provides certain interpretability to the analysis. Deep learning can be powerful for such tasks, but surprisingly their combination with a focus on interpretability and generalizability is rarely explored. In this work, we introduce a novel framework for interpretable deep image decomposition, combining hierarchical Bayesian modeling and deep learning to create an architecture-modularized and model-generalizable deep neural network (DNN). The proposed framework includes three steps: (1) hierarchical Bayesian modeling of image decomposition, (2) transforming the inference problem into optimization tasks, and (3) deep inference via a modularized Bayesian DNN. We further establish a theoretical connection between the loss function and the generalization error bound, which inspires a new test-time adaptation approach for out-of-distribution scenarios. We instantiated the application using two downstream tasks, \textit{i.e.}, image denoising, and unsupervised anomaly detection, and the results demonstrated improved generalizability as well as interpretability of our methods. The source code will be released upon the acceptance of this paper.
Trustworthy Contrast-enhanced Brain MRI Synthesis
Jul 10, 2024



Abstract:Contrast-enhanced brain MRI (CE-MRI) is a valuable diagnostic technique but may pose health risks and incur high costs. To create safer alternatives, multi-modality medical image translation aims to synthesize CE-MRI images from other available modalities. Although existing methods can generate promising predictions, they still face two challenges, i.e., exhibiting over-confidence and lacking interpretability on predictions. To address the above challenges, this paper introduces TrustI2I, a novel trustworthy method that reformulates multi-to-one medical image translation problem as a multimodal regression problem, aiming to build an uncertainty-aware and reliable system. Specifically, our method leverages deep evidential regression to estimate prediction uncertainties and employs an explicit intermediate and late fusion strategy based on the Mixture of Normal Inverse Gamma (MoNIG) distribution, enhancing both synthesis quality and interpretability. Additionally, we incorporate uncertainty calibration to improve the reliability of uncertainty. Validation on the BraTS2018 dataset demonstrates that our approach surpasses current methods, producing higher-quality images with rational uncertainty estimation.
Evidential Concept Embedding Models: Towards Reliable Concept Explanations for Skin Disease Diagnosis
Jun 27, 2024Abstract:Due to the high stakes in medical decision-making, there is a compelling demand for interpretable deep learning methods in medical image analysis. Concept Bottleneck Models (CBM) have emerged as an active interpretable framework incorporating human-interpretable concepts into decision-making. However, their concept predictions may lack reliability when applied to clinical diagnosis, impeding concept explanations' quality. To address this, we propose an evidential Concept Embedding Model (evi-CEM), which employs evidential learning to model the concept uncertainty. Additionally, we offer to leverage the concept uncertainty to rectify concept misalignments that arise when training CBMs using vision-language models without complete concept supervision. With the proposed methods, we can enhance concept explanations' reliability for both supervised and label-efficient settings. Furthermore, we introduce concept uncertainty for effective test-time intervention. Our evaluation demonstrates that evi-CEM achieves superior performance in terms of concept prediction, and the proposed concept rectification effectively mitigates concept misalignments for label-efficient training. Our code is available at https://github.com/obiyoag/evi-CEM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge