Vicente Grau
Cherise
EchoLVFM: One-Step Video Generation via Latent Flow Matching for Echocardiogram Synthesis
Mar 14, 2026Abstract:Echocardiography is widely used for assessing cardiac function, where clinically meaningful parameters such as left-ventricular ejection fraction (EF) play a central role in diagnosis and management. Generative models capable of synthesising realistic echocardiogram videos with explicit control over such parameters are valuable for data augmentation, counterfactual analysis, and specialist training. However, existing approaches typically rely on computationally expensive multi-step sampling and aggressive temporal normalisation, limiting efficiency and applicability to heterogeneous real-world data. We introduce EchoLVFM, a one-step latent video flow-matching framework for controllable echocardiogram generation. Operating in the latent space, EchoLVFM synthesises temporally coherent videos in a single inference step, achieving a $\mathbf{\sim 50\times}$ improvement in sampling efficiency compared to multi-step flow baselines while maintaining visual fidelity. The model supports global conditioning on clinical variables, demonstrated through precise control of EF, and enables reconstruction and counterfactual generation from partially observed sequences. A masked conditioning strategy further removes fixed-length constraints, allowing shorter sequences to be retained rather than discarded. We evaluate EchoLVFM on the CAMUS dataset under challenging single-frame conditioning. Quantitative and qualitative results demonstrate competitive video quality, strong EF adherence, and 57.9% discrimination accuracy by expert clinicians which is close to chance. These findings indicate that efficient, one-step flow matching can enable practical, controllable echocardiogram video synthesis without sacrificing fidelity. Code available at: https://github.com/EngEmmanuel/EchoLVFM
Seeing Beyond the Image: ECG and Anatomical Knowledge-Guided Myocardial Scar Segmentation from Late Gadolinium-Enhanced Images
Nov 18, 2025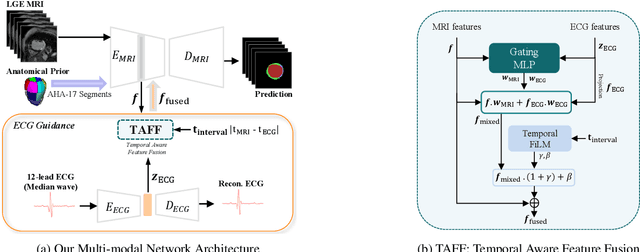
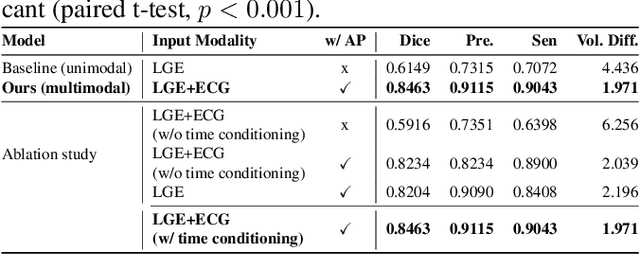
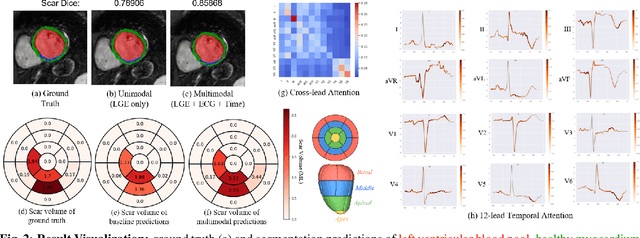
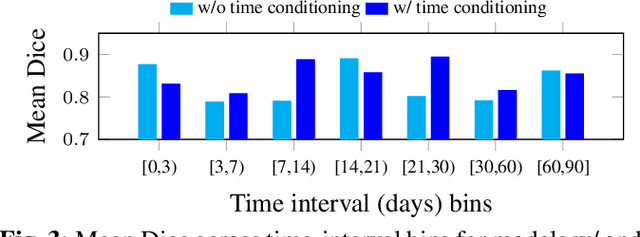
Abstract:Accurate segmentation of myocardial scar from late gadolinium enhanced (LGE) cardiac MRI is essential for evaluating tissue viability, yet remains challenging due to variable contrast and imaging artifacts. Electrocardiogram (ECG) signals provide complementary physiological information, as conduction abnormalities can help localize or suggest scarred myocardial regions. In this work, we propose a novel multimodal framework that integrates ECG-derived electrophysiological information with anatomical priors from the AHA-17 atlas for physiologically consistent LGE-based scar segmentation. As ECGs and LGE-MRIs are not acquired simultaneously, we introduce a Temporal Aware Feature Fusion (TAFF) mechanism that dynamically weights and fuses features based on their acquisition time difference. Our method was evaluated on a clinical dataset and achieved substantial gains over the state-of-the-art image-only baseline (nnU-Net), increasing the average Dice score for scars from 0.6149 to 0.8463 and achieving high performance in both precision (0.9115) and sensitivity (0.9043). These results show that integrating physiological and anatomical knowledge allows the model to "see beyond the image", setting a new direction for robust and physiologically grounded cardiac scar segmentation.
AortaDiff: A Unified Multitask Diffusion Framework For Contrast-Free AAA Imaging
Oct 01, 2025Abstract:While contrast-enhanced CT (CECT) is standard for assessing abdominal aortic aneurysms (AAA), the required iodinated contrast agents pose significant risks, including nephrotoxicity, patient allergies, and environmental harm. To reduce contrast agent use, recent deep learning methods have focused on generating synthetic CECT from non-contrast CT (NCCT) scans. However, most adopt a multi-stage pipeline that first generates images and then performs segmentation, which leads to error accumulation and fails to leverage shared semantic and anatomical structures. To address this, we propose a unified deep learning framework that generates synthetic CECT images from NCCT scans while simultaneously segmenting the aortic lumen and thrombus. Our approach integrates conditional diffusion models (CDM) with multi-task learning, enabling end-to-end joint optimization of image synthesis and anatomical segmentation. Unlike previous multitask diffusion models, our approach requires no initial predictions (e.g., a coarse segmentation mask), shares both encoder and decoder parameters across tasks, and employs a semi-supervised training strategy to learn from scans with missing segmentation labels, a common constraint in real-world clinical data. We evaluated our method on a cohort of 264 patients, where it consistently outperformed state-of-the-art single-task and multi-stage models. For image synthesis, our model achieved a PSNR of 25.61 dB, compared to 23.80 dB from a single-task CDM. For anatomical segmentation, it improved the lumen Dice score to 0.89 from 0.87 and the challenging thrombus Dice score to 0.53 from 0.48 (nnU-Net). These segmentation enhancements led to more accurate clinical measurements, reducing the lumen diameter MAE to 4.19 mm from 5.78 mm and the thrombus area error to 33.85% from 41.45% when compared to nnU-Net. Code is available at https://github.com/yuxuanou623/AortaDiff.git.
From Gaze to Insight: Bridging Human Visual Attention and Vision Language Model Explanation for Weakly-Supervised Medical Image Segmentation
Apr 15, 2025Abstract:Medical image segmentation remains challenging due to the high cost of pixel-level annotations for training. In the context of weak supervision, clinician gaze data captures regions of diagnostic interest; however, its sparsity limits its use for segmentation. In contrast, vision-language models (VLMs) provide semantic context through textual descriptions but lack the explanation precision required. Recognizing that neither source alone suffices, we propose a teacher-student framework that integrates both gaze and language supervision, leveraging their complementary strengths. Our key insight is that gaze data indicates where clinicians focus during diagnosis, while VLMs explain why those regions are significant. To implement this, the teacher model first learns from gaze points enhanced by VLM-generated descriptions of lesion morphology, establishing a foundation for guiding the student model. The teacher then directs the student through three strategies: (1) Multi-scale feature alignment to fuse visual cues with textual semantics; (2) Confidence-weighted consistency constraints to focus on reliable predictions; (3) Adaptive masking to limit error propagation in uncertain areas. Experiments on the Kvasir-SEG, NCI-ISBI, and ISIC datasets show that our method achieves Dice scores of 80.78%, 80.53%, and 84.22%, respectively-improving 3-5% over gaze baselines without increasing the annotation burden. By preserving correlations among predictions, gaze data, and lesion descriptions, our framework also maintains clinical interpretability. This work illustrates how integrating human visual attention with AI-generated semantic context can effectively overcome the limitations of individual weak supervision signals, thereby advancing the development of deployable, annotation-efficient medical AI systems. Code is available at: https://github.com/jingkunchen/FGI.git.
Unsupervised Patch-GAN with Targeted Patch Ranking for Fine-Grained Novelty Detection in Medical Imaging
Jan 29, 2025



Abstract:Detecting novel anomalies in medical imaging is challenging due to the limited availability of labeled data for rare abnormalities, which often display high variability and subtlety. This challenge is further compounded when small abnormal regions are embedded within larger normal areas, as whole-image predictions frequently overlook these subtle deviations. To address these issues, we propose an unsupervised Patch-GAN framework designed to detect and localize anomalies by capturing both local detail and global structure. Our framework first reconstructs masked images to learn fine-grained, normal-specific features, allowing for enhanced sensitivity to minor deviations from normality. By dividing these reconstructed images into patches and assessing the authenticity of each patch, our approach identifies anomalies at a more granular level, overcoming the limitations of whole-image evaluation. Additionally, a patch-ranking mechanism prioritizes regions with higher abnormal scores, reinforcing the alignment between local patch discrepancies and the global image context. Experimental results on the ISIC 2016 skin lesion and BraTS 2019 brain tumor datasets validate our framework's effectiveness, achieving AUCs of 95.79% and 96.05%, respectively, and outperforming three state-of-the-art baselines.
Contrast-Free Myocardial Scar Segmentation in Cine MRI using Motion and Texture Fusion
Jan 09, 2025Abstract:Late gadolinium enhancement MRI (LGE MRI) is the gold standard for the detection of myocardial scars for post myocardial infarction (MI). LGE MRI requires the injection of a contrast agent, which carries potential side effects and increases scanning time and patient discomfort. To address these issues, we propose a novel framework that combines cardiac motion observed in cine MRI with image texture information to segment the myocardium and scar tissue in the left ventricle. Cardiac motion tracking can be formulated as a full cardiac image cycle registration problem, which can be solved via deep neural networks. Experimental results prove that the proposed method can achieve scar segmentation based on non-contrasted cine images with comparable accuracy to LGE MRI. This demonstrates its potential as an alternative to contrast-enhanced techniques for scar detection.
Transesophageal Echocardiography Generation using Anatomical Models
Oct 09, 2024Abstract:Through automation, deep learning (DL) can enhance the analysis of transesophageal echocardiography (TEE) images. However, DL methods require large amounts of high-quality data to produce accurate results, which is difficult to satisfy. Data augmentation is commonly used to tackle this issue. In this work, we develop a pipeline to generate synthetic TEE images and corresponding semantic labels. The proposed data generation pipeline expands on an existing pipeline that generates synthetic transthoracic echocardiography images by transforming slices from anatomical models into synthetic images. We also demonstrate that such images can improve DL network performance through a left-ventricle semantic segmentation task. For the pipeline's unpaired image-to-image (I2I) translation section, we explore two generative methods: CycleGAN and contrastive unpaired translation. Next, we evaluate the synthetic images quantitatively using the Fr\'echet Inception Distance (FID) Score and qualitatively through a human perception quiz involving expert cardiologists and the average researcher. In this study, we achieve a dice score improvement of up to 10% when we augment datasets with our synthetic images. Furthermore, we compare established methods of assessing unpaired I2I translation and observe a disagreement when evaluating the synthetic images. Finally, we see which metric better predicts the generated data's efficacy when used for data augmentation.
NeCA: 3D Coronary Artery Tree Reconstruction from Two 2D Projections by Neural Implicit Representation
Sep 06, 2024
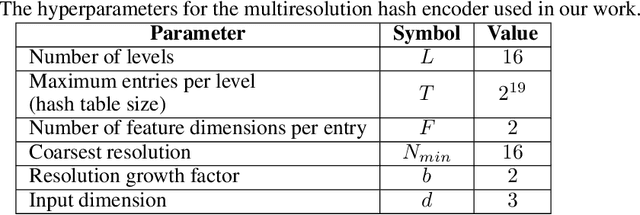


Abstract:Cardiovascular diseases (CVDs) are the most common health threats worldwide. 2D x-ray invasive coronary angiography (ICA) remains as the most widely adopted imaging modality for CVDs diagnosis. However, in current clinical practice, it is often difficult for the cardiologists to interpret the 3D geometry of coronary vessels based on 2D planes. Moreover, due to the radiation limit, in general only two angiographic projections are acquired, providing limited information of the vessel geometry and necessitating 3D coronary tree reconstruction based only on two ICA projections. In this paper, we propose a self-supervised deep learning method called NeCA, which is based on implicit neural representation using the multiresolution hash encoder and differentiable cone-beam forward projector layer in order to achieve 3D coronary artery tree reconstruction from two projections. We validate our method using six different metrics on coronary computed tomography angiography data in terms of right coronary artery and left anterior descending respectively. The evaluation results demonstrate that our NeCA method, without 3D ground truth for supervision and large datasets for training, achieves promising performance in both vessel topology preservation and branch-connectivity maintaining compared to the supervised deep learning model.
Personalized Topology-Informed 12-Lead ECG Electrode Localization from Incomplete Cardiac MRIs for Efficient Cardiac Digital Twins
Aug 25, 2024Abstract:Cardiac digital twins (CDTs) offer personalized \textit{in-silico} cardiac representations for the inference of multi-scale properties tied to cardiac mechanisms. The creation of CDTs requires precise information about the electrode position on the torso, especially for the personalized electrocardiogram (ECG) calibration. However, current studies commonly rely on additional acquisition of torso imaging and manual/semi-automatic methods for ECG electrode localization. In this study, we propose a novel and efficient topology-informed model to fully automatically extract personalized ECG electrode locations from 2D clinically standard cardiac MRIs. Specifically, we obtain the sparse torso contours from the cardiac MRIs and then localize the electrodes from the contours. Cardiac MRIs aim at imaging of the heart instead of the torso, leading to incomplete torso geometry within the imaging. To tackle the missing topology, we incorporate the electrodes as a subset of the keypoints, which can be explicitly aligned with the 3D torso topology. The experimental results demonstrate that the proposed model outperforms the time-consuming conventional method in terms of accuracy (Euclidean distance: $1.24 \pm 0.293$ cm vs. $1.48 \pm 0.362$ cm) and efficiency ($2$~s vs. $30$-$35$~min). We further demonstrate the effectiveness of using the detected electrodes for \textit{in-silico} ECG simulation, highlighting their potential for creating accurate and efficient CDT models. The code will be released publicly after the manuscript is accepted for publication.
Deep Learning-based 3D Coronary Tree Reconstruction from Two 2D Non-simultaneous X-ray Angiography Projections
Jul 19, 2024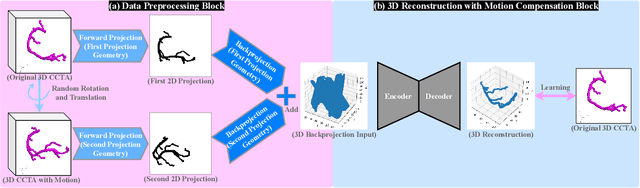

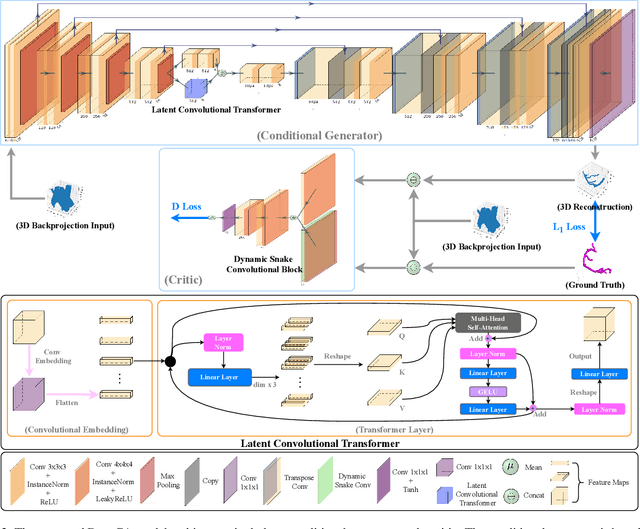

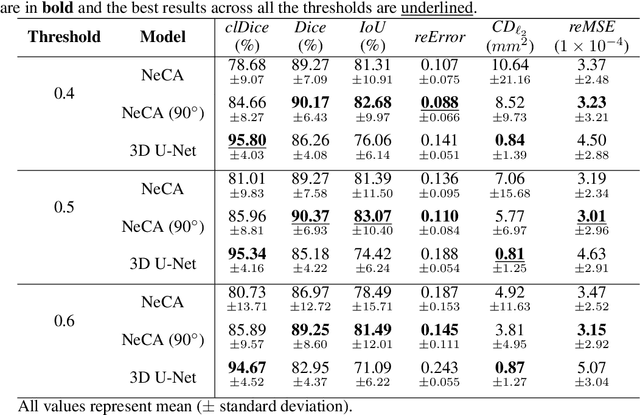
Abstract:Cardiovascular diseases (CVDs) are the most common cause of death worldwide. Invasive x-ray coronary angiography (ICA) is one of the most important imaging modalities for the diagnosis of CVDs. ICA typically acquires only two 2D projections, which makes the 3D geometry of coronary vessels difficult to interpret, thus requiring 3D coronary tree reconstruction from two projections. State-of-the-art approaches require significant manual interactions and cannot correct the non-rigid cardiac and respiratory motions between non-simultaneous projections. In this study, we propose a novel deep learning pipeline. We leverage the Wasserstein conditional generative adversarial network with gradient penalty, latent convolutional transformer layers, and a dynamic snake convolutional critic to implicitly compensate for the non-rigid motion and provide 3D coronary tree reconstruction. Through simulating projections from coronary computed tomography angiography (CCTA), we achieve the generalisation of 3D coronary tree reconstruction on real non-simultaneous ICA projections. We incorporate an application-specific evaluation metric to validate our proposed model on both a CCTA dataset and a real ICA dataset, together with Chamfer L1 distance. The results demonstrate the good performance of our model in vessel topology preservation, recovery of missing features, and generalisation ability to real ICA data. To the best of our knowledge, this is the first study that leverages deep learning to achieve 3D coronary tree reconstruction from two real non-simultaneous x-ray angiography projections.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge