Jun Lyu
Enabling Ultra-Fast Cardiovascular Imaging Across Heterogeneous Clinical Environments with a Generalist Foundation Model and Multimodal Database
Dec 25, 2025Abstract:Multimodal cardiovascular magnetic resonance (CMR) imaging provides comprehensive and non-invasive insights into cardiovascular disease (CVD) diagnosis and underlying mechanisms. Despite decades of advancements, its widespread clinical adoption remains constrained by prolonged scan times and heterogeneity across medical environments. This underscores the urgent need for a generalist reconstruction foundation model for ultra-fast CMR imaging, one capable of adapting across diverse imaging scenarios and serving as the essential substrate for all downstream analyses. To enable this goal, we curate MMCMR-427K, the largest and most comprehensive multimodal CMR k-space database to date, comprising 427,465 multi-coil k-space data paired with structured metadata across 13 international centers, 12 CMR modalities, 15 scanners, and 17 CVD categories in populations across three continents. Building on this unprecedented resource, we introduce CardioMM, a generalist reconstruction foundation model capable of dynamically adapting to heterogeneous fast CMR imaging scenarios. CardioMM unifies semantic contextual understanding with physics-informed data consistency to deliver robust reconstructions across varied scanners, protocols, and patient presentations. Comprehensive evaluations demonstrate that CardioMM achieves state-of-the-art performance in the internal centers and exhibits strong zero-shot generalization to unseen external settings. Even at imaging acceleration up to 24x, CardioMM reliably preserves key cardiac phenotypes, quantitative myocardial biomarkers, and diagnostic image quality, enabling a substantial increase in CMR examination throughput without compromising clinical integrity. Together, our open-access MMCMR-427K database and CardioMM framework establish a scalable pathway toward high-throughput, high-quality, and clinically accessible cardiovascular imaging.
Towards Universal Learning-based Model for Cardiac Image Reconstruction: Summary of the CMRxRecon2024 Challenge
Mar 05, 2025Abstract:Cardiovascular magnetic resonance (CMR) offers diverse imaging contrasts for assessment of cardiac function and tissue characterization. However, acquiring each single CMR modality is often time-consuming, and comprehensive clinical protocols require multiple modalities with various sampling patterns, further extending the overall acquisition time and increasing susceptibility to motion artifacts. Existing deep learning-based reconstruction methods are often designed for specific acquisition parameters, which limits their ability to generalize across a variety of scan scenarios. As part of the CMRxRecon Series, the CMRxRecon2024 challenge provides diverse datasets encompassing multi-modality multi-view imaging with various sampling patterns, and a platform for the international community to develop and benchmark reconstruction solutions in two well-crafted tasks. Task 1 is a modality-universal setting, evaluating the out-of-distribution generalization of the reconstructed model, while Task 2 follows sampling-universal setting assessing the one-for-all adaptability of the universal model. Main contributions include providing the first and largest publicly available multi-modality, multi-view cardiac k-space dataset; developing a benchmarking platform that simulates clinical acceleration protocols, with a shared code library and tutorial for various k-t undersampling patterns and data processing; giving technical insights of enhanced data consistency based on physic-informed networks and adaptive prompt-learning embedding to be versatile to different clinical settings; additional finding on evaluation metrics to address the limitations of conventional ground-truth references in universal reconstruction tasks.
Rapid Whole Brain Mesoscale In-vivo MR Imaging using Multi-scale Implicit Neural Representation
Feb 12, 2025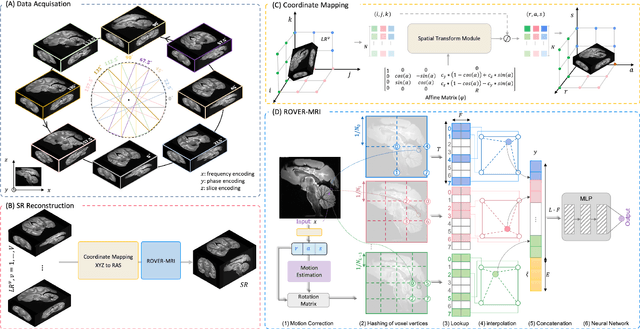
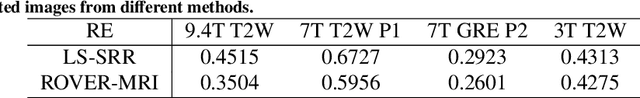
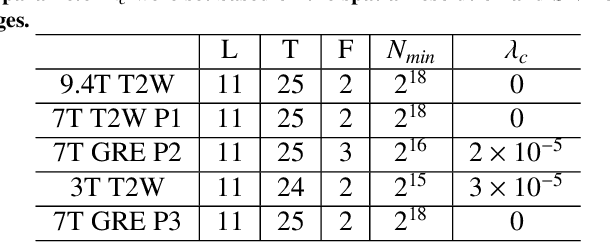
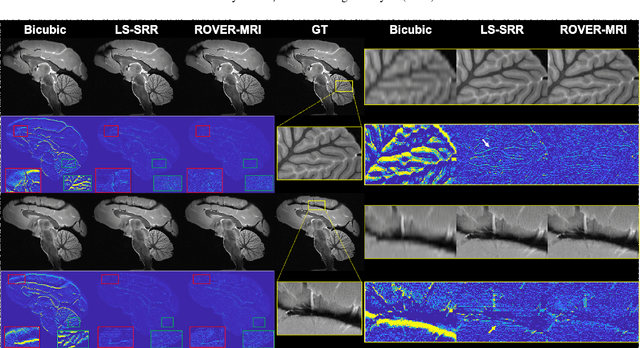

Abstract:Purpose: To develop and validate a novel image reconstruction technique using implicit neural representations (INR) for multi-view thick-slice acquisitions while reducing the scan time but maintaining high signal-to-noise ratio (SNR). Methods: We propose Rotating-view super-resolution (ROVER)-MRI, an unsupervised neural network-based algorithm designed to reconstruct MRI data from multi-view thick slices, effectively reducing scan time by 2-fold while maintaining fine anatomical details. We compare our method to both bicubic interpolation and the current state-of-the-art regularized least-squares super-resolution reconstruction (LS-SRR) technique. Validation is performed using ground-truth ex-vivo monkey brain data, and we demonstrate superior reconstruction quality across several in-vivo human datasets. Notably, we achieve the reconstruction of a whole human brain in-vivo T2-weighted image with an unprecedented 180{\mu}m isotropic spatial resolution, accomplished in just 17 minutes of scan time on a 7T MRI scanner. Results: ROVER-MRI outperformed LS-SRR method in terms of reconstruction quality with 22.4% lower relative error (RE) and 7.5% lower full-width half maximum (FWHM) indicating better preservation of fine structural details in nearly half the scan time. Conclusion: ROVER-MRI offers an efficient and robust approach for mesoscale MR imaging, enabling rapid, high-resolution whole-brain scans. Its versatility holds great promise for research applications requiring anatomical details and time-efficient imaging.
MSV-Mamba: A Multiscale Vision Mamba Network for Echocardiography Segmentation
Jan 13, 2025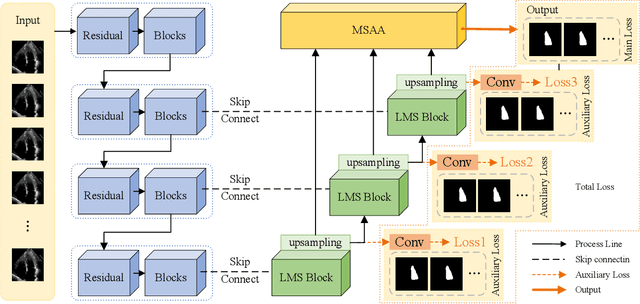
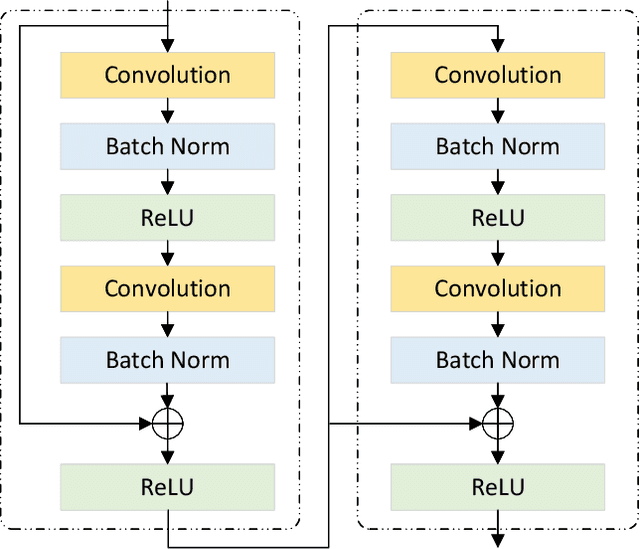
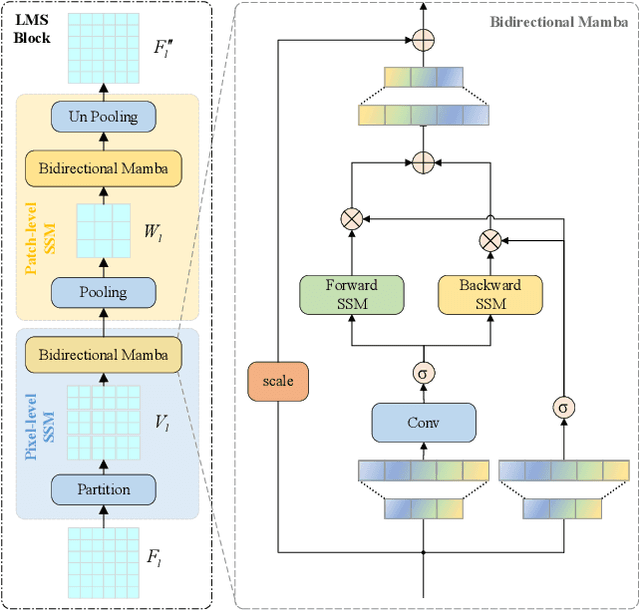
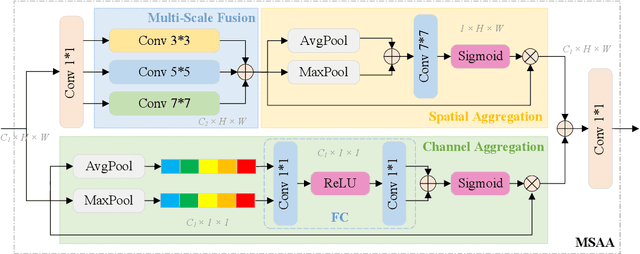
Abstract:Ultrasound imaging frequently encounters challenges, such as those related to elevated noise levels, diminished spatiotemporal resolution, and the complexity of anatomical structures. These factors significantly hinder the model's ability to accurately capture and analyze structural relationships and dynamic patterns across various regions of the heart. Mamba, an emerging model, is one of the most cutting-edge approaches that is widely applied to diverse vision and language tasks. To this end, this paper introduces a U-shaped deep learning model incorporating a large-window Mamba scale (LMS) module and a hierarchical feature fusion approach for echocardiographic segmentation. First, a cascaded residual block serves as an encoder and is employed to incrementally extract multiscale detailed features. Second, a large-window multiscale mamba module is integrated into the decoder to capture global dependencies across regions and enhance the segmentation capability for complex anatomical structures. Furthermore, our model introduces auxiliary losses at each decoder layer and employs a dual attention mechanism to fuse multilayer features both spatially and across channels. This approach enhances segmentation performance and accuracy in delineating complex anatomical structures. Finally, the experimental results using the EchoNet-Dynamic and CAMUS datasets demonstrate that the model outperforms other methods in terms of both accuracy and robustness. For the segmentation of the left ventricular endocardium (${LV}_{endo}$), the model achieved optimal values of 95.01 and 93.36, respectively, while for the left ventricular epicardium (${LV}_{epi}$), values of 87.35 and 87.80, respectively, were achieved. This represents an improvement ranging between 0.54 and 1.11 compared with the best-performing model.
CMRxRecon2024: A Multi-Modality, Multi-View K-Space Dataset Boosting Universal Machine Learning for Accelerated Cardiac MRI
Jun 27, 2024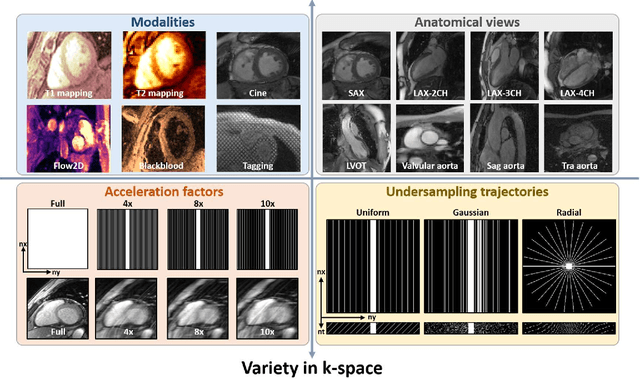
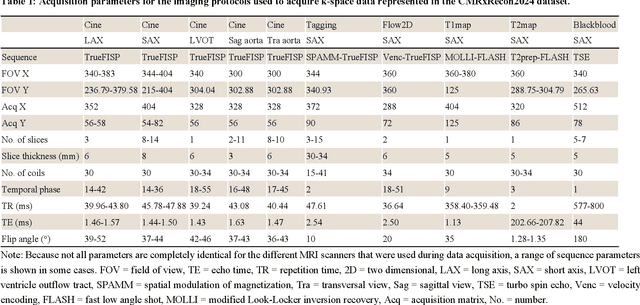
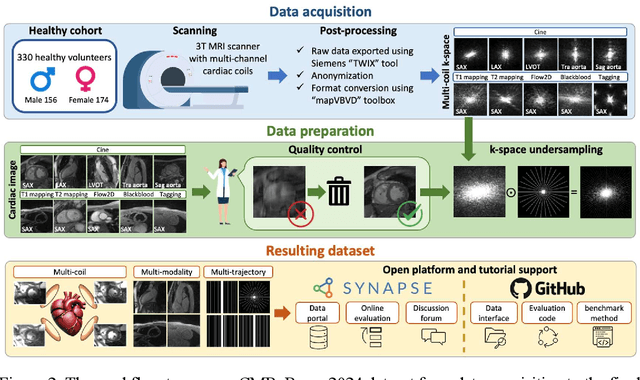
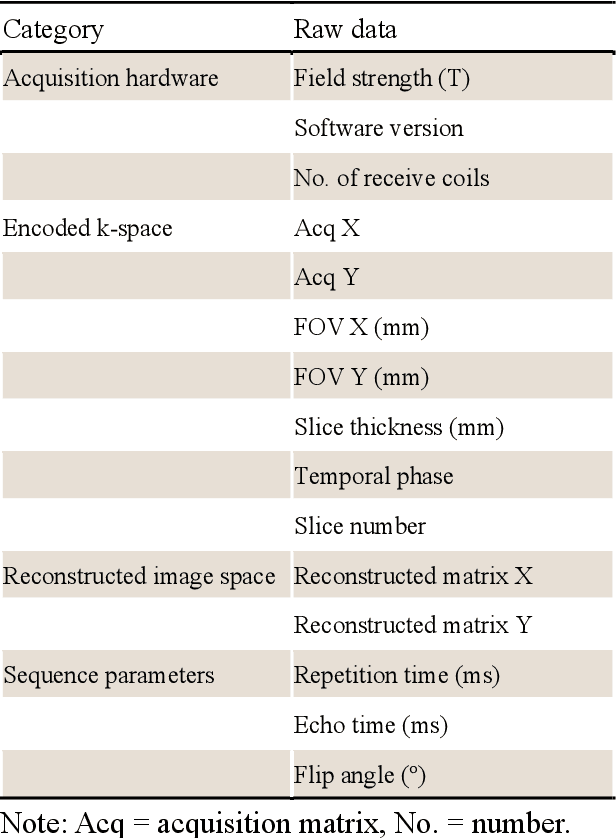
Abstract:Cardiac magnetic resonance imaging (MRI) has emerged as a clinically gold-standard technique for diagnosing cardiac diseases, thanks to its ability to provide diverse information with multiple modalities and anatomical views. Accelerated cardiac MRI is highly expected to achieve time-efficient and patient-friendly imaging, and then advanced image reconstruction approaches are required to recover high-quality, clinically interpretable images from undersampled measurements. However, the lack of publicly available cardiac MRI k-space dataset in terms of both quantity and diversity has severely hindered substantial technological progress, particularly for data-driven artificial intelligence. Here, we provide a standardized, diverse, and high-quality CMRxRecon2024 dataset to facilitate the technical development, fair evaluation, and clinical transfer of cardiac MRI reconstruction approaches, towards promoting the universal frameworks that enable fast and robust reconstructions across different cardiac MRI protocols in clinical practice. To the best of our knowledge, the CMRxRecon2024 dataset is the largest and most diverse publicly available cardiac k-space dataset. It is acquired from 330 healthy volunteers, covering commonly used modalities, anatomical views, and acquisition trajectories in clinical cardiac MRI workflows. Besides, an open platform with tutorials, benchmarks, and data processing tools is provided to facilitate data usage, advanced method development, and fair performance evaluation.
An Empirical Study on the Fairness of Foundation Models for Multi-Organ Image Segmentation
Jun 18, 2024



Abstract:The segmentation foundation model, e.g., Segment Anything Model (SAM), has attracted increasing interest in the medical image community. Early pioneering studies primarily concentrated on assessing and improving SAM's performance from the perspectives of overall accuracy and efficiency, yet little attention was given to the fairness considerations. This oversight raises questions about the potential for performance biases that could mirror those found in task-specific deep learning models like nnU-Net. In this paper, we explored the fairness dilemma concerning large segmentation foundation models. We prospectively curate a benchmark dataset of 3D MRI and CT scans of the organs including liver, kidney, spleen, lung and aorta from a total of 1056 healthy subjects with expert segmentations. Crucially, we document demographic details such as gender, age, and body mass index (BMI) for each subject to facilitate a nuanced fairness analysis. We test state-of-the-art foundation models for medical image segmentation, including the original SAM, medical SAM and SAT models, to evaluate segmentation efficacy across different demographic groups and identify disparities. Our comprehensive analysis, which accounts for various confounding factors, reveals significant fairness concerns within these foundational models. Moreover, our findings highlight not only disparities in overall segmentation metrics, such as the Dice Similarity Coefficient but also significant variations in the spatial distribution of segmentation errors, offering empirical evidence of the nuanced challenges in ensuring fairness in medical image segmentation.
The state-of-the-art in Cardiac MRI Reconstruction: Results of the CMRxRecon Challenge in MICCAI 2023
Apr 01, 2024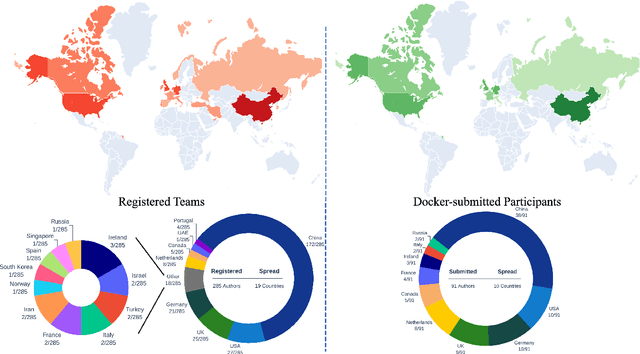
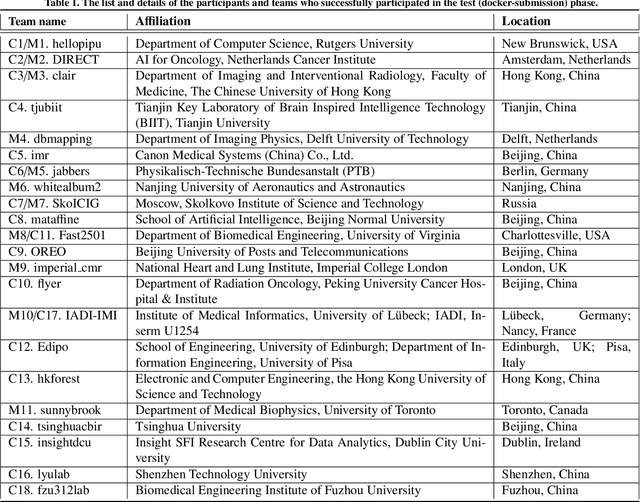
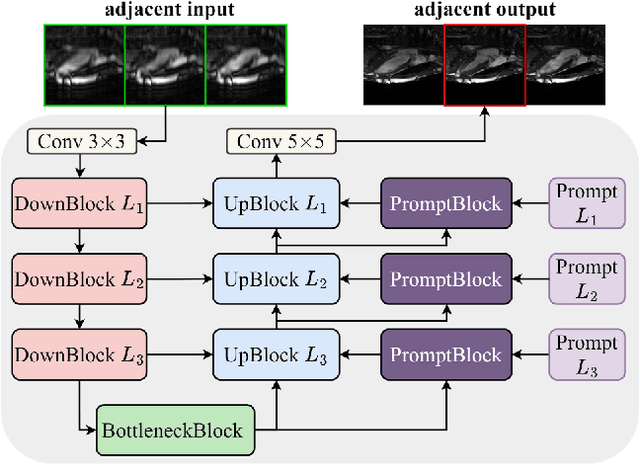
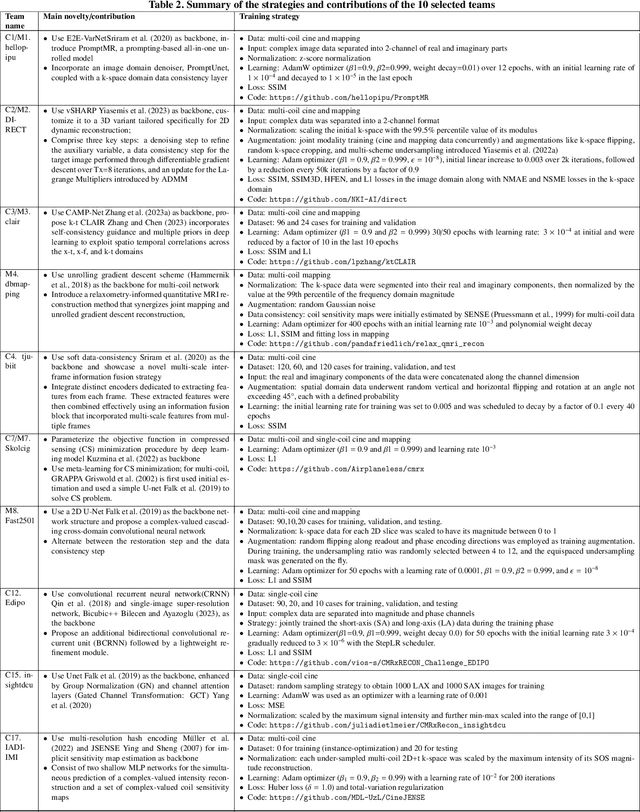
Abstract:Cardiac MRI, crucial for evaluating heart structure and function, faces limitations like slow imaging and motion artifacts. Undersampling reconstruction, especially data-driven algorithms, has emerged as a promising solution to accelerate scans and enhance imaging performance using highly under-sampled data. Nevertheless, the scarcity of publicly available cardiac k-space datasets and evaluation platform hinder the development of data-driven reconstruction algorithms. To address this issue, we organized the Cardiac MRI Reconstruction Challenge (CMRxRecon) in 2023, in collaboration with the 26th International Conference on MICCAI. CMRxRecon presented an extensive k-space dataset comprising cine and mapping raw data, accompanied by detailed annotations of cardiac anatomical structures. With overwhelming participation, the challenge attracted more than 285 teams and over 600 participants. Among them, 22 teams successfully submitted Docker containers for the testing phase, with 7 teams submitted for both cine and mapping tasks. All teams use deep learning based approaches, indicating that deep learning has predominately become a promising solution for the problem. The first-place winner of both tasks utilizes the E2E-VarNet architecture as backbones. In contrast, U-Net is still the most popular backbone for both multi-coil and single-coil reconstructions. This paper provides a comprehensive overview of the challenge design, presents a summary of the submitted results, reviews the employed methods, and offers an in-depth discussion that aims to inspire future advancements in cardiac MRI reconstruction models. The summary emphasizes the effective strategies observed in Cardiac MRI reconstruction, including backbone architecture, loss function, pre-processing techniques, physical modeling, and model complexity, thereby providing valuable insights for further developments in this field.
CMRxRecon: An open cardiac MRI dataset for the competition of accelerated image reconstruction
Sep 19, 2023



Abstract:Cardiac magnetic resonance imaging (CMR) has emerged as a valuable diagnostic tool for cardiac diseases. However, a limitation of CMR is its slow imaging speed, which causes patient discomfort and introduces artifacts in the images. There has been growing interest in deep learning-based CMR imaging algorithms that can reconstruct high-quality images from highly under-sampled k-space data. However, the development of deep learning methods requires large training datasets, which have not been publicly available for CMR. To address this gap, we released a dataset that includes multi-contrast, multi-view, multi-slice and multi-coil CMR imaging data from 300 subjects. Imaging studies include cardiac cine and mapping sequences. Manual segmentations of the myocardium and chambers of all the subjects are also provided within the dataset. Scripts of state-of-the-art reconstruction algorithms were also provided as a point of reference. Our aim is to facilitate the advancement of state-of-the-art CMR image reconstruction by introducing standardized evaluation criteria and making the dataset freely accessible to the research community. Researchers can access the dataset at https://www.synapse.org/#!Synapse:syn51471091/wiki/.
Dual Arbitrary Scale Super-Resolution for Multi-Contrast MRI
Jul 10, 2023Abstract:Limited by imaging systems, the reconstruction of Magnetic Resonance Imaging (MRI) images from partial measurement is essential to medical imaging research. Benefiting from the diverse and complementary information of multi-contrast MR images in different imaging modalities, multi-contrast Super-Resolution (SR) reconstruction is promising to yield SR images with higher quality. In the medical scenario, to fully visualize the lesion, radiologists are accustomed to zooming the MR images at arbitrary scales rather than using a fixed scale, as used by most MRI SR methods. In addition, existing multi-contrast MRI SR methods often require a fixed resolution for the reference image, which makes acquiring reference images difficult and imposes limitations on arbitrary scale SR tasks. To address these issues, we proposed an implicit neural representations based dual-arbitrary multi-contrast MRI super-resolution method, called Dual-ArbNet. First, we decouple the resolution of the target and reference images by a feature encoder, enabling the network to input target and reference images at arbitrary scales. Then, an implicit fusion decoder fuses the multi-contrast features and uses an Implicit Decoding Function~(IDF) to obtain the final MRI SR results. Furthermore, we introduce a curriculum learning strategy to train our network, which improves the generalization and performance of our Dual-ArbNet. Extensive experiments in two public MRI datasets demonstrate that our method outperforms state-of-the-art approaches under different scale factors and has great potential in clinical practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge