Cheng Ouyang
Topology Optimization in Medical Image Segmentation with Fast Euler Characteristic
Jul 31, 2025Abstract:Deep learning-based medical image segmentation techniques have shown promising results when evaluated based on conventional metrics such as the Dice score or Intersection-over-Union. However, these fully automatic methods often fail to meet clinically acceptable accuracy, especially when topological constraints should be observed, e.g., continuous boundaries or closed surfaces. In medical image segmentation, the correctness of a segmentation in terms of the required topological genus sometimes is even more important than the pixel-wise accuracy. Existing topology-aware approaches commonly estimate and constrain the topological structure via the concept of persistent homology (PH). However, these methods are difficult to implement for high dimensional data due to their polynomial computational complexity. To overcome this problem, we propose a novel and fast approach for topology-aware segmentation based on the Euler Characteristic ($\chi$). First, we propose a fast formulation for $\chi$ computation in both 2D and 3D. The scalar $\chi$ error between the prediction and ground-truth serves as the topological evaluation metric. Then we estimate the spatial topology correctness of any segmentation network via a so-called topological violation map, i.e., a detailed map that highlights regions with $\chi$ errors. Finally, the segmentation results from the arbitrary network are refined based on the topological violation maps by a topology-aware correction network. Our experiments are conducted on both 2D and 3D datasets and show that our method can significantly improve topological correctness while preserving pixel-wise segmentation accuracy.
Dataset Distillation with Probabilistic Latent Features
May 10, 2025Abstract:As deep learning models grow in complexity and the volume of training data increases, reducing storage and computational costs becomes increasingly important. Dataset distillation addresses this challenge by synthesizing a compact set of synthetic data that can effectively replace the original dataset in downstream classification tasks. While existing methods typically rely on mapping data from pixel space to the latent space of a generative model, we propose a novel stochastic approach that models the joint distribution of latent features. This allows our method to better capture spatial structures and produce diverse synthetic samples, which benefits model training. Specifically, we introduce a low-rank multivariate normal distribution parameterized by a lightweight network. This design maintains low computational complexity and is compatible with various matching networks used in dataset distillation. After distillation, synthetic images are generated by feeding the learned latent features into a pretrained generator. These synthetic images are then used to train classification models, and performance is evaluated on real test set. We validate our method on several benchmarks, including ImageNet subsets, CIFAR-10, and the MedMNIST histopathological dataset. Our approach achieves state-of-the-art cross architecture performance across a range of backbone architectures, demonstrating its generality and effectiveness.
Federated Continual 3D Segmentation With Single-round Communication
Mar 19, 2025



Abstract:Federated learning seeks to foster collaboration among distributed clients while preserving the privacy of their local data. Traditionally, federated learning methods assume a fixed setting in which client data and learning objectives remain constant. However, in real-world scenarios, new clients may join, and existing clients may expand the segmentation label set as task requirements evolve. In such a dynamic federated analysis setup, the conventional federated communication strategy of model aggregation per communication round is suboptimal. As new clients join, this strategy requires retraining, linearly increasing communication and computation overhead. It also imposes requirements for synchronized communication, which is difficult to achieve among distributed clients. In this paper, we propose a federated continual learning strategy that employs a one-time model aggregation at the server through multi-model distillation. This approach builds and updates the global model while eliminating the need for frequent server communication. When integrating new data streams or onboarding new clients, this approach efficiently reuses previous client models, avoiding the need to retrain the global model across the entire federation. By minimizing communication load and bypassing the need to put unchanged clients online, our approach relaxes synchronization requirements among clients, providing an efficient and scalable federated analysis framework suited for real-world applications. Using multi-class 3D abdominal CT segmentation as an application task, we demonstrate the effectiveness of the proposed approach.
MedVLM-R1: Incentivizing Medical Reasoning Capability of Vision-Language Models (VLMs) via Reinforcement Learning
Feb 26, 2025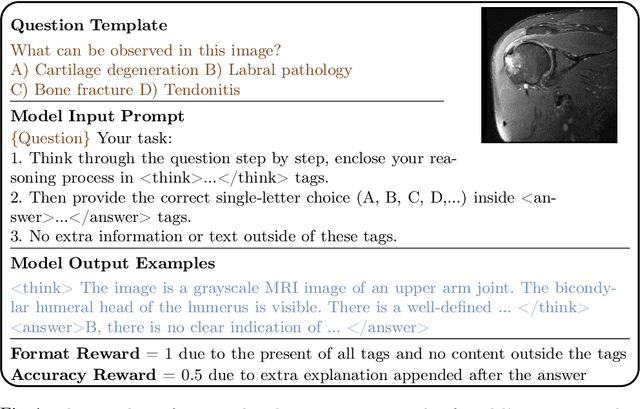
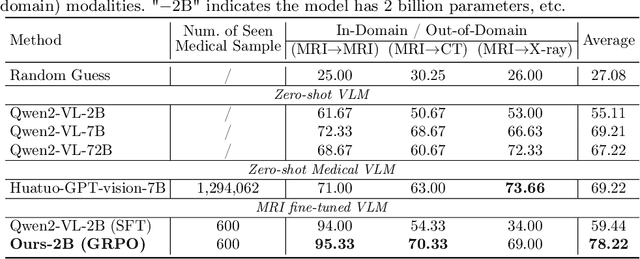
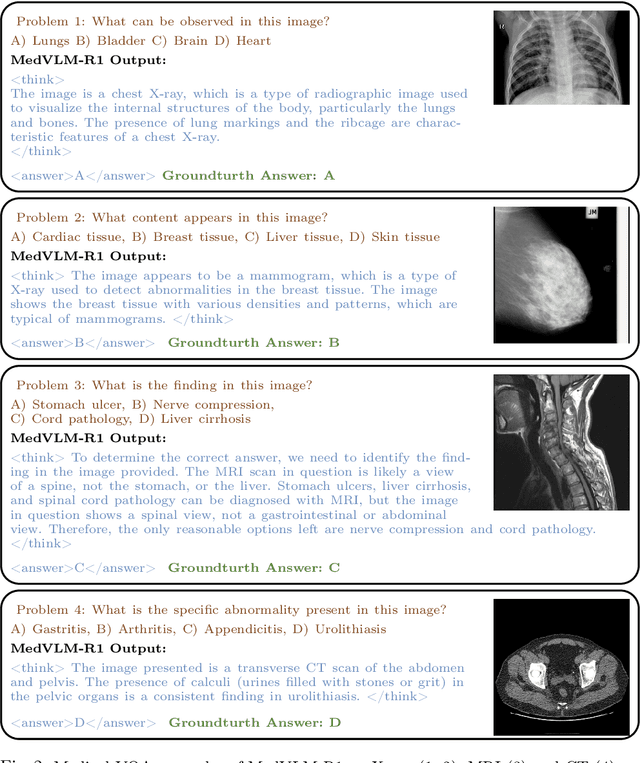
Abstract:Reasoning is a critical frontier for advancing medical image analysis, where transparency and trustworthiness play a central role in both clinician trust and regulatory approval. Although Medical Visual Language Models (VLMs) show promise for radiological tasks, most existing VLMs merely produce final answers without revealing the underlying reasoning. To address this gap, we introduce MedVLM-R1, a medical VLM that explicitly generates natural language reasoning to enhance transparency and trustworthiness. Instead of relying on supervised fine-tuning (SFT), which often suffers from overfitting to training distributions and fails to foster genuine reasoning, MedVLM-R1 employs a reinforcement learning framework that incentivizes the model to discover human-interpretable reasoning paths without using any reasoning references. Despite limited training data (600 visual question answering samples) and model parameters (2B), MedVLM-R1 boosts accuracy from 55.11% to 78.22% across MRI, CT, and X-ray benchmarks, outperforming larger models trained on over a million samples. It also demonstrates robust domain generalization under out-of-distribution tasks. By unifying medical image analysis with explicit reasoning, MedVLM-R1 marks a pivotal step toward trustworthy and interpretable AI in clinical practice.
Knowledge-enhanced Multimodal ECG Representation Learning with Arbitrary-Lead Inputs
Feb 25, 2025


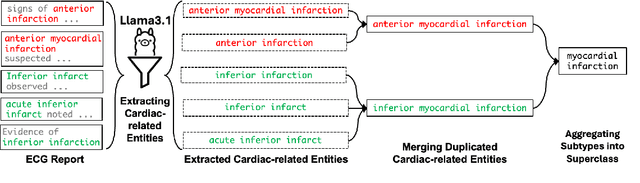
Abstract:Recent advances in multimodal ECG representation learning center on aligning ECG signals with paired free-text reports. However, suboptimal alignment persists due to the complexity of medical language and the reliance on a full 12-lead setup, which is often unavailable in under-resourced settings. To tackle these issues, we propose **K-MERL**, a knowledge-enhanced multimodal ECG representation learning framework. **K-MERL** leverages large language models to extract structured knowledge from free-text reports and employs a lead-aware ECG encoder with dynamic lead masking to accommodate arbitrary lead inputs. Evaluations on six external ECG datasets show that **K-MERL** achieves state-of-the-art performance in zero-shot classification and linear probing tasks, while delivering an average **16%** AUC improvement over existing methods in partial-lead zero-shot classification.
SPA: Efficient User-Preference Alignment against Uncertainty in Medical Image Segmentation
Nov 23, 2024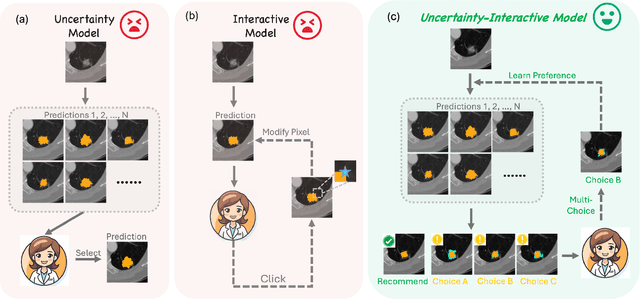
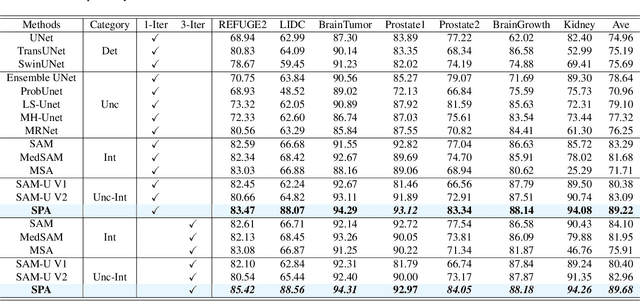
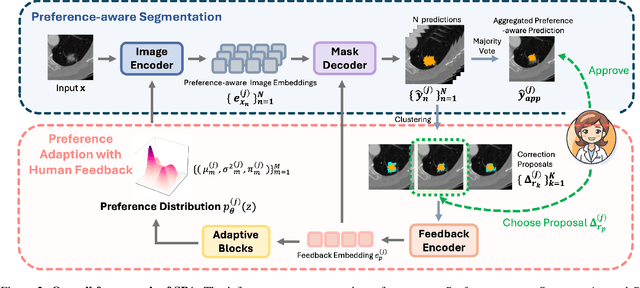
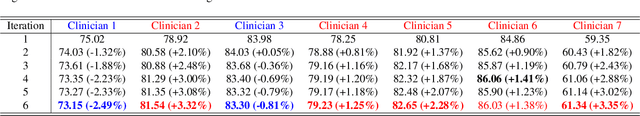
Abstract:Medical image segmentation data inherently contain uncertainty, often stemming from both imperfect image quality and variability in labeling preferences on ambiguous pixels, which depend on annotators' expertise and the clinical context of the annotations. For instance, a boundary pixel might be labeled as tumor in diagnosis to avoid under-assessment of severity, but as normal tissue in radiotherapy to prevent damage to sensitive structures. As segmentation preferences vary across downstream applications, it is often desirable for an image segmentation model to offer user-adaptable predictions rather than a fixed output. While prior uncertainty-aware and interactive methods offer adaptability, they are inefficient at test time: uncertainty-aware models require users to choose from numerous similar outputs, while interactive models demand significant user input through click or box prompts to refine segmentation. To address these challenges, we propose \textbf{SPA}, a segmentation framework that efficiently adapts to diverse test-time preferences with minimal human interaction. By presenting users a select few, distinct segmentation candidates that best capture uncertainties, it reduces clinician workload in reaching the preferred segmentation. To accommodate user preference, we introduce a probabilistic mechanism that leverages user feedback to adapt model's segmentation preference. The proposed framework is evaluated on a diverse range of medical image segmentation tasks: color fundus images, CT, and MRI. It demonstrates 1) a significant reduction in clinician time and effort compared with existing interactive segmentation approaches, 2) strong adaptability based on human feedback, and 3) state-of-the-art image segmentation performance across diverse modalities and semantic labels.
Universal Topology Refinement for Medical Image Segmentation with Polynomial Feature Synthesis
Sep 15, 2024Abstract:Although existing medical image segmentation methods provide impressive pixel-wise accuracy, they often neglect topological correctness, making their segmentations unusable for many downstream tasks. One option is to retrain such models whilst including a topology-driven loss component. However, this is computationally expensive and often impractical. A better solution would be to have a versatile plug-and-play topology refinement method that is compatible with any domain-specific segmentation pipeline. Directly training a post-processing model to mitigate topological errors often fails as such models tend to be biased towards the topological errors of a target segmentation network. The diversity of these errors is confined to the information provided by a labelled training set, which is especially problematic for small datasets. Our method solves this problem by training a model-agnostic topology refinement network with synthetic segmentations that cover a wide variety of topological errors. Inspired by the Stone-Weierstrass theorem, we synthesize topology-perturbation masks with randomly sampled coefficients of orthogonal polynomial bases, which ensures a complete and unbiased representation. Practically, we verified the efficiency and effectiveness of our methods as being compatible with multiple families of polynomial bases, and show evidence that our universal plug-and-play topology refinement network outperforms both existing topology-driven learning-based and post-processing methods. We also show that combining our method with learning-based models provides an effortless add-on, which can further improve the performance of existing approaches.
Stability and Generalizability in SDE Diffusion Models with Measure-Preserving Dynamics
Jun 19, 2024Abstract:Inverse problems describe the process of estimating the causal factors from a set of measurements or data. Mapping of often incomplete or degraded data to parameters is ill-posed, thus data-driven iterative solutions are required, for example when reconstructing clean images from poor signals. Diffusion models have shown promise as potent generative tools for solving inverse problems due to their superior reconstruction quality and their compatibility with iterative solvers. However, most existing approaches are limited to linear inverse problems represented as Stochastic Differential Equations (SDEs). This simplification falls short of addressing the challenging nature of real-world problems, leading to amplified cumulative errors and biases. We provide an explanation for this gap through the lens of measure-preserving dynamics of Random Dynamical Systems (RDS) with which we analyse Temporal Distribution Discrepancy and thus introduce a theoretical framework based on RDS for SDE diffusion models. We uncover several strategies that inherently enhance the stability and generalizability of diffusion models for inverse problems and introduce a novel score-based diffusion framework, the \textbf{D}ynamics-aware S\textbf{D}E \textbf{D}iffusion \textbf{G}enerative \textbf{M}odel (D$^3$GM). The \textit{Measure-preserving property} can return the degraded measurement to the original state despite complex degradation with the RDS concept of \textit{stability}. Our extensive experimental results corroborate the effectiveness of D$^3$GM across multiple benchmarks including a prominent application for inverse problems, magnetic resonance imaging. Code and data will be publicly available.
Probabilistic Contrastive Learning with Explicit Concentration on the Hypersphere
May 26, 2024Abstract:Self-supervised contrastive learning has predominantly adopted deterministic methods, which are not suited for environments characterized by uncertainty and noise. This paper introduces a new perspective on incorporating uncertainty into contrastive learning by embedding representations within a spherical space, inspired by the von Mises-Fisher distribution (vMF). We introduce an unnormalized form of vMF and leverage the concentration parameter, kappa, as a direct, interpretable measure to quantify uncertainty explicitly. This approach not only provides a probabilistic interpretation of the embedding space but also offers a method to calibrate model confidence against varying levels of data corruption and characteristics. Our empirical results demonstrate that the estimated concentration parameter correlates strongly with the degree of unforeseen data corruption encountered at test time, enables failure analysis, and enhances existing out-of-distribution detection methods.
A Foundation Model for Brain Lesion Segmentation with Mixture of Modality Experts
May 16, 2024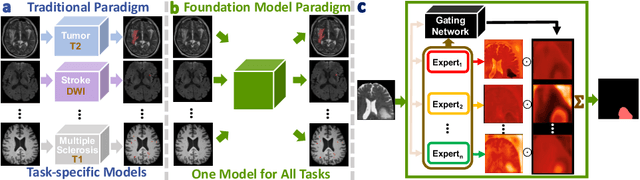
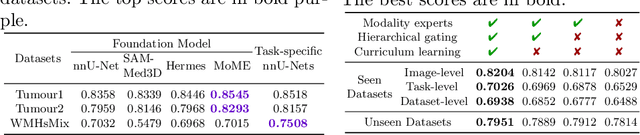
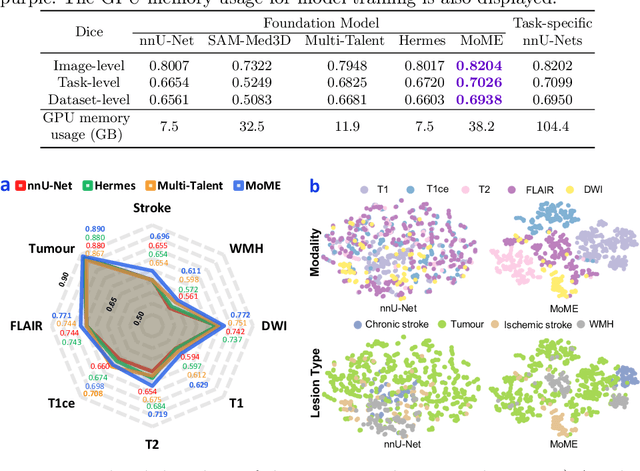

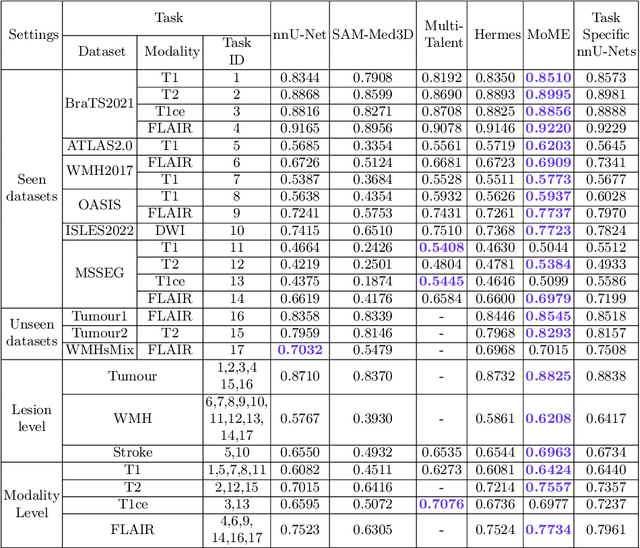
Abstract:Brain lesion segmentation plays an essential role in neurological research and diagnosis. As brain lesions can be caused by various pathological alterations, different types of brain lesions tend to manifest with different characteristics on different imaging modalities. Due to this complexity, brain lesion segmentation methods are often developed in a task-specific manner. A specific segmentation model is developed for a particular lesion type and imaging modality. However, the use of task-specific models requires predetermination of the lesion type and imaging modality, which complicates their deployment in real-world scenarios. In this work, we propose a universal foundation model for 3D brain lesion segmentation, which can automatically segment different types of brain lesions for input data of various imaging modalities. We formulate a novel Mixture of Modality Experts (MoME) framework with multiple expert networks attending to different imaging modalities. A hierarchical gating network combines the expert predictions and fosters expertise collaboration. Furthermore, we introduce a curriculum learning strategy during training to avoid the degeneration of each expert network and preserve their specialization. We evaluated the proposed method on nine brain lesion datasets, encompassing five imaging modalities and eight lesion types. The results show that our model outperforms state-of-the-art universal models and provides promising generalization to unseen datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge