Zhiling Yan
Can MLLMs Read Students' Minds? Unpacking Multimodal Error Analysis in Handwritten Math
Mar 26, 2026Abstract:Assessing student handwritten scratchwork is crucial for personalized educational feedback but presents unique challenges due to diverse handwriting, complex layouts, and varied problem-solving approaches. Existing educational NLP primarily focuses on textual responses and neglects the complexity and multimodality inherent in authentic handwritten scratchwork. Current multimodal large language models (MLLMs) excel at visual reasoning but typically adopt an "examinee perspective", prioritizing generating correct answers rather than diagnosing student errors. To bridge these gaps, we introduce ScratchMath, a novel benchmark specifically designed for explaining and classifying errors in authentic handwritten mathematics scratchwork. Our dataset comprises 1,720 mathematics samples from Chinese primary and middle school students, supporting two key tasks: Error Cause Explanation (ECE) and Error Cause Classification (ECC), with seven defined error types. The dataset is meticulously annotated through rigorous human-machine collaborative approaches involving multiple stages of expert labeling, review, and verification. We systematically evaluate 16 leading MLLMs on ScratchMath, revealing significant performance gaps relative to human experts, especially in visual recognition and logical reasoning. Proprietary models notably outperform open-source models, with large reasoning models showing strong potential for error explanation. All evaluation data and frameworks are publicly available to facilitate further research.
LiveMedBench: A Contamination-Free Medical Benchmark for LLMs with Automated Rubric Evaluation
Feb 10, 2026Abstract:The deployment of Large Language Models (LLMs) in high-stakes clinical settings demands rigorous and reliable evaluation. However, existing medical benchmarks remain static, suffering from two critical limitations: (1) data contamination, where test sets inadvertently leak into training corpora, leading to inflated performance estimates; and (2) temporal misalignment, failing to capture the rapid evolution of medical knowledge. Furthermore, current evaluation metrics for open-ended clinical reasoning often rely on either shallow lexical overlap (e.g., ROUGE) or subjective LLM-as-a-Judge scoring, both inadequate for verifying clinical correctness. To bridge these gaps, we introduce LiveMedBench, a continuously updated, contamination-free, and rubric-based benchmark that weekly harvests real-world clinical cases from online medical communities, ensuring strict temporal separation from model training data. We propose a Multi-Agent Clinical Curation Framework that filters raw data noise and validates clinical integrity against evidence-based medical principles. For evaluation, we develop an Automated Rubric-based Evaluation Framework that decomposes physician responses into granular, case-specific criteria, achieving substantially stronger alignment with expert physicians than LLM-as-a-Judge. To date, LiveMedBench comprises 2,756 real-world cases spanning 38 medical specialties and multiple languages, paired with 16,702 unique evaluation criteria. Extensive evaluation of 38 LLMs reveals that even the best-performing model achieves only 39.2%, and 84% of models exhibit performance degradation on post-cutoff cases, confirming pervasive data contamination risks. Error analysis further identifies contextual application-not factual knowledge-as the dominant bottleneck, with 35-48% of failures stemming from the inability to tailor medical knowledge to patient-specific constraints.
TTT-Unet: Enhancing U-Net with Test-Time Training Layers for Biomedical Image Segmentation
Sep 18, 2024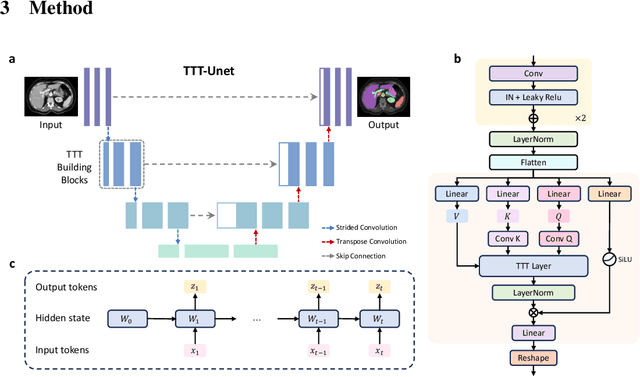

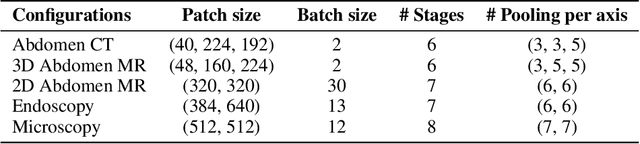
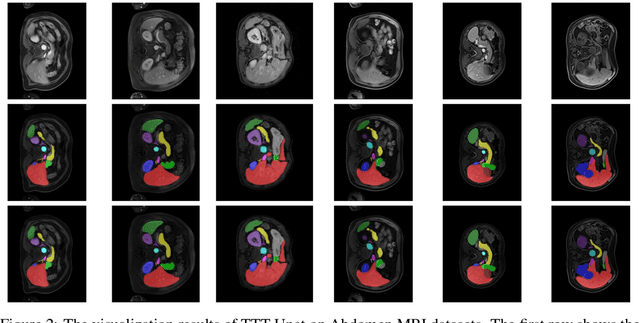
Abstract:Biomedical image segmentation is crucial for accurately diagnosing and analyzing various diseases. However, Convolutional Neural Networks (CNNs) and Transformers, the most commonly used architectures for this task, struggle to effectively capture long-range dependencies due to the inherent locality of CNNs and the computational complexity of Transformers. To address this limitation, we introduce TTT-Unet, a novel framework that integrates Test-Time Training (TTT) layers into the traditional U-Net architecture for biomedical image segmentation. TTT-Unet dynamically adjusts model parameters during the testing time, enhancing the model's ability to capture both local and long-range features. We evaluate TTT-Unet on multiple medical imaging datasets, including 3D abdominal organ segmentation in CT and MR images, instrument segmentation in endoscopy images, and cell segmentation in microscopy images. The results demonstrate that TTT-Unet consistently outperforms state-of-the-art CNN-based and Transformer-based segmentation models across all tasks. The code is available at https://github.com/rongzhou7/TTT-Unet.
Biomedical SAM 2: Segment Anything in Biomedical Images and Videos
Aug 06, 2024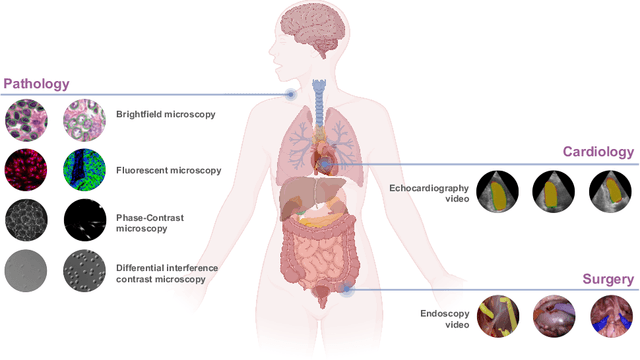
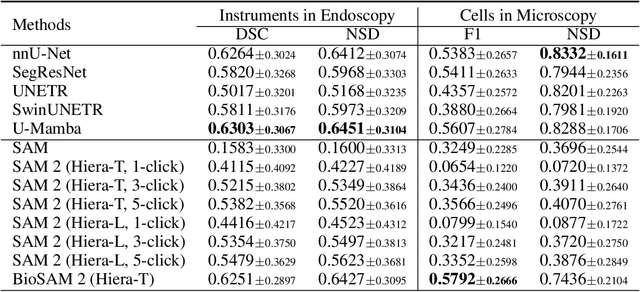
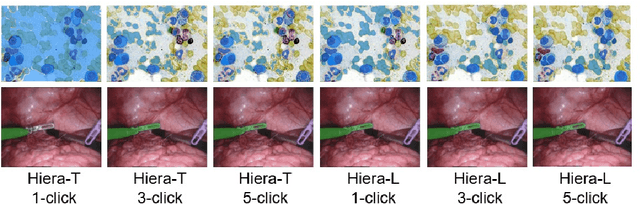
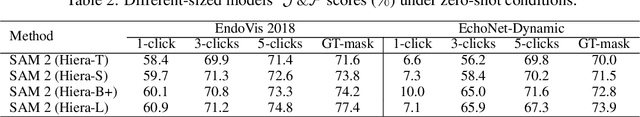
Abstract:Medical image segmentation and video object segmentation are essential for diagnosing and analyzing diseases by identifying and measuring biological structures. Recent advances in natural domain have been driven by foundation models like the Segment Anything Model 2 (SAM 2). To explore the performance of SAM 2 in biomedical applications, we designed two evaluation pipelines for single-frame image segmentation and multi-frame video segmentation with varied prompt designs, revealing SAM 2's limitations in medical contexts. Consequently, we developed BioSAM 2, an enhanced foundation model optimized for biomedical data based on SAM 2. Our experiments show that BioSAM 2 not only surpasses the performance of existing state-of-the-art foundation models but also matches or even exceeds specialist models, demonstrating its efficacy and potential in the medical domain.
Medical Unlearnable Examples: Securing Medical Data from Unauthorized Traning via Sparsity-Aware Local Masking
Mar 15, 2024Abstract:With the rapid growth of artificial intelligence (AI) in healthcare, there has been a significant increase in the generation and storage of sensitive medical data. This abundance of data, in turn, has propelled the advancement of medical AI technologies. However, concerns about unauthorized data exploitation, such as training commercial AI models, often deter researchers from making their invaluable datasets publicly available. In response to the need to protect this hard-to-collect data while still encouraging medical institutions to share it, one promising solution is to introduce imperceptible noise into the data. This method aims to safeguard the data against unauthorized training by inducing degradation in model generalization. Although existing methods have shown commendable data protection capabilities in general domains, they tend to fall short when applied to biomedical data, mainly due to their failure to account for the sparse nature of medical images. To address this problem, we propose the Sparsity-Aware Local Masking (SALM) method, a novel approach that selectively perturbs significant pixel regions rather than the entire image as previous strategies have done. This simple-yet-effective approach significantly reduces the perturbation search space by concentrating on local regions, thereby improving both the efficiency and effectiveness of data protection for biomedical datasets characterized by sparse features. Besides, we have demonstrated that SALM maintains the essential characteristics of the data, ensuring its clinical utility remains uncompromised. Our extensive experiments across various datasets and model architectures demonstrate that SALM effectively prevents unauthorized training of deep-learning models and outperforms previous state-of-the-art data protection methods.
Sora: A Review on Background, Technology, Limitations, and Opportunities of Large Vision Models
Feb 28, 2024



Abstract:Sora is a text-to-video generative AI model, released by OpenAI in February 2024. The model is trained to generate videos of realistic or imaginative scenes from text instructions and show potential in simulating the physical world. Based on public technical reports and reverse engineering, this paper presents a comprehensive review of the model's background, related technologies, applications, remaining challenges, and future directions of text-to-video AI models. We first trace Sora's development and investigate the underlying technologies used to build this "world simulator". Then, we describe in detail the applications and potential impact of Sora in multiple industries ranging from film-making and education to marketing. We discuss the main challenges and limitations that need to be addressed to widely deploy Sora, such as ensuring safe and unbiased video generation. Lastly, we discuss the future development of Sora and video generation models in general, and how advancements in the field could enable new ways of human-AI interaction, boosting productivity and creativity of video generation.
Multimodal ChatGPT for Medical Applications: an Experimental Study of GPT-4V
Oct 29, 2023



Abstract:In this paper, we critically evaluate the capabilities of the state-of-the-art multimodal large language model, i.e., GPT-4 with Vision (GPT-4V), on Visual Question Answering (VQA) task. Our experiments thoroughly assess GPT-4V's proficiency in answering questions paired with images using both pathology and radiology datasets from 11 modalities (e.g. Microscopy, Dermoscopy, X-ray, CT, etc.) and fifteen objects of interests (brain, liver, lung, etc.). Our datasets encompass a comprehensive range of medical inquiries, including sixteen distinct question types. Throughout our evaluations, we devised textual prompts for GPT-4V, directing it to synergize visual and textual information. The experiments with accuracy score conclude that the current version of GPT-4V is not recommended for real-world diagnostics due to its unreliable and suboptimal accuracy in responding to diagnostic medical questions. In addition, we delineate seven unique facets of GPT-4V's behavior in medical VQA, highlighting its constraints within this complex arena. The complete details of our evaluation cases are accessible at https://github.com/ZhilingYan/GPT4V-Medical-Report.
MA-SAM: Modality-agnostic SAM Adaptation for 3D Medical Image Segmentation
Sep 16, 2023



Abstract:The Segment Anything Model (SAM), a foundation model for general image segmentation, has demonstrated impressive zero-shot performance across numerous natural image segmentation tasks. However, SAM's performance significantly declines when applied to medical images, primarily due to the substantial disparity between natural and medical image domains. To effectively adapt SAM to medical images, it is important to incorporate critical third-dimensional information, i.e., volumetric or temporal knowledge, during fine-tuning. Simultaneously, we aim to harness SAM's pre-trained weights within its original 2D backbone to the fullest extent. In this paper, we introduce a modality-agnostic SAM adaptation framework, named as MA-SAM, that is applicable to various volumetric and video medical data. Our method roots in the parameter-efficient fine-tuning strategy to update only a small portion of weight increments while preserving the majority of SAM's pre-trained weights. By injecting a series of 3D adapters into the transformer blocks of the image encoder, our method enables the pre-trained 2D backbone to extract third-dimensional information from input data. The effectiveness of our method has been comprehensively evaluated on four medical image segmentation tasks, by using 10 public datasets across CT, MRI, and surgical video data. Remarkably, without using any prompt, our method consistently outperforms various state-of-the-art 3D approaches, surpassing nnU-Net by 0.9%, 2.6%, and 9.9% in Dice for CT multi-organ segmentation, MRI prostate segmentation, and surgical scene segmentation respectively. Our model also demonstrates strong generalization, and excels in challenging tumor segmentation when prompts are used. Our code is available at: https://github.com/cchen-cc/MA-SAM.
BiomedGPT: A Unified and Generalist Biomedical Generative Pre-trained Transformer for Vision, Language, and Multimodal Tasks
May 26, 2023Abstract:In this paper, we introduce a unified and generalist Biomedical Generative Pre-trained Transformer (BiomedGPT) model, which leverages self-supervision on large and diverse datasets to accept multi-modal inputs and perform a range of downstream tasks. Our experiments demonstrate that BiomedGPT delivers expansive and inclusive representations of biomedical data, outperforming the majority of preceding state-of-the-art models across five distinct tasks with 20 public datasets spanning over 15 unique biomedical modalities. Through the ablation study, we also showcase the efficacy of our multi-modal and multi-task pretraining approach in transferring knowledge to previously unseen data. Overall, our work presents a significant step forward in developing unified and generalist models for biomedicine, with far-reaching implications for improving healthcare outcomes.
Learning to Generate Poetic Chinese Landscape Painting with Calligraphy
May 08, 2023



Abstract:In this paper, we present a novel system (denoted as Polaca) to generate poetic Chinese landscape painting with calligraphy. Unlike previous single image-to-image painting generation, Polaca takes the classic poetry as input and outputs the artistic landscape painting image with the corresponding calligraphy. It is equipped with three different modules to complete the whole piece of landscape painting artwork: the first one is a text-to-image module to generate landscape painting image, the second one is an image-to-image module to generate stylistic calligraphy image, and the third one is an image fusion module to fuse the two images into a whole piece of aesthetic artwork.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge