TTT-Unet: Enhancing U-Net with Test-Time Training Layers for Biomedical Image Segmentation
Paper and Code
Sep 18, 2024

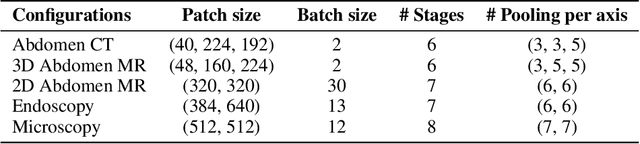
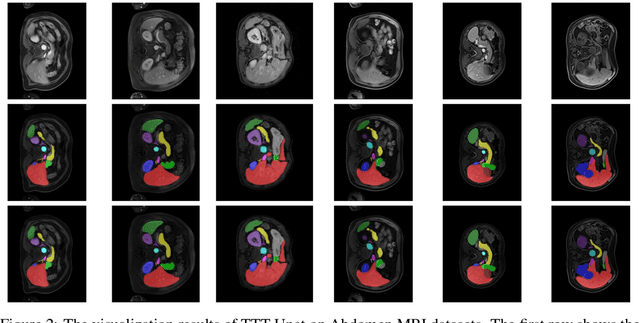
Biomedical image segmentation is crucial for accurately diagnosing and analyzing various diseases. However, Convolutional Neural Networks (CNNs) and Transformers, the most commonly used architectures for this task, struggle to effectively capture long-range dependencies due to the inherent locality of CNNs and the computational complexity of Transformers. To address this limitation, we introduce TTT-Unet, a novel framework that integrates Test-Time Training (TTT) layers into the traditional U-Net architecture for biomedical image segmentation. TTT-Unet dynamically adjusts model parameters during the testing time, enhancing the model's ability to capture both local and long-range features. We evaluate TTT-Unet on multiple medical imaging datasets, including 3D abdominal organ segmentation in CT and MR images, instrument segmentation in endoscopy images, and cell segmentation in microscopy images. The results demonstrate that TTT-Unet consistently outperforms state-of-the-art CNN-based and Transformer-based segmentation models across all tasks. The code is available at https://github.com/rongzhou7/TTT-Unet.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge