Quanzheng Li
CAMCA, Massachusetts General Hospital and Harvard Medical School
MAPLE: Elevating Medical Reasoning from Statistical Consensus to Process-Led Alignment
Mar 09, 2026Abstract:Recent advances in medical large language models have explored Test-Time Reinforcement Learning (TTRL) to enhance reasoning. However, standard TTRL often relies on majority voting (MV) as a heuristic supervision signal, which can be unreliable in complex medical scenarios where the most frequent reasoning path is not necessarily the clinically correct one. In this work, we propose a novel and unified training paradigm that integrates medical process reward models with TTRL to bridge the gap between test-time scaling (TTS) and parametric model optimization. Specifically, we advance the TTRL framework by replacing the conventional MV with a fine-grained, expert-aligned supervision paradigm using Med-RPM. This integration ensures that reinforcement learning is guided by medical correctness rather than mere consensus, effectively distilling search-based intelligence into the model's parametric memory. Extensive evaluations on four different benchmarks have demonstrated that our developed method consistently and significantly outperforms current TTRL and standalone PRM selection. Our findings establish that transitioning from stochastic heuristics to structured, step-wise rewards is essential for developing reliable and scalable medical AI systems
LiveMedBench: A Contamination-Free Medical Benchmark for LLMs with Automated Rubric Evaluation
Feb 10, 2026Abstract:The deployment of Large Language Models (LLMs) in high-stakes clinical settings demands rigorous and reliable evaluation. However, existing medical benchmarks remain static, suffering from two critical limitations: (1) data contamination, where test sets inadvertently leak into training corpora, leading to inflated performance estimates; and (2) temporal misalignment, failing to capture the rapid evolution of medical knowledge. Furthermore, current evaluation metrics for open-ended clinical reasoning often rely on either shallow lexical overlap (e.g., ROUGE) or subjective LLM-as-a-Judge scoring, both inadequate for verifying clinical correctness. To bridge these gaps, we introduce LiveMedBench, a continuously updated, contamination-free, and rubric-based benchmark that weekly harvests real-world clinical cases from online medical communities, ensuring strict temporal separation from model training data. We propose a Multi-Agent Clinical Curation Framework that filters raw data noise and validates clinical integrity against evidence-based medical principles. For evaluation, we develop an Automated Rubric-based Evaluation Framework that decomposes physician responses into granular, case-specific criteria, achieving substantially stronger alignment with expert physicians than LLM-as-a-Judge. To date, LiveMedBench comprises 2,756 real-world cases spanning 38 medical specialties and multiple languages, paired with 16,702 unique evaluation criteria. Extensive evaluation of 38 LLMs reveals that even the best-performing model achieves only 39.2%, and 84% of models exhibit performance degradation on post-cutoff cases, confirming pervasive data contamination risks. Error analysis further identifies contextual application-not factual knowledge-as the dominant bottleneck, with 35-48% of failures stemming from the inability to tailor medical knowledge to patient-specific constraints.
CoMMa: Contribution-Aware Medical Multi-Agents From A Game-Theoretic Perspective
Feb 09, 2026Abstract:Recent multi-agent frameworks have broadened the ability to tackle oncology decision support tasks that require reasoning over dynamic, heterogeneous patient data. We propose Contribution-Aware Medical Multi-Agents (CoMMa), a decentralized LLM-agent framework in which specialists operate on partitioned evidence and coordinate through a game-theoretic objective for robust decision-making. In contrast to most agent architectures relying on stochastic narrative-based reasoning, CoMMa utilizes deterministic embedding projections to approximate contribution-aware credit assignment. This yields explicit evidence attribution by estimating each agent's marginal utility, producing interpretable and mathematically grounded decision pathways with improved stability. Evaluated on diverse oncology benchmarks, including a real-world multidisciplinary tumor board dataset, CoMMa achieves higher accuracy and more stable performance than data-centralized and role-based multi-agents baselines.
Adaptation of Agentic AI
Dec 22, 2025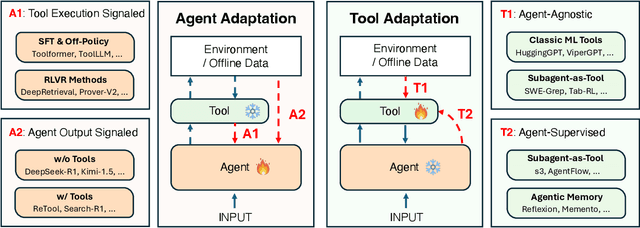
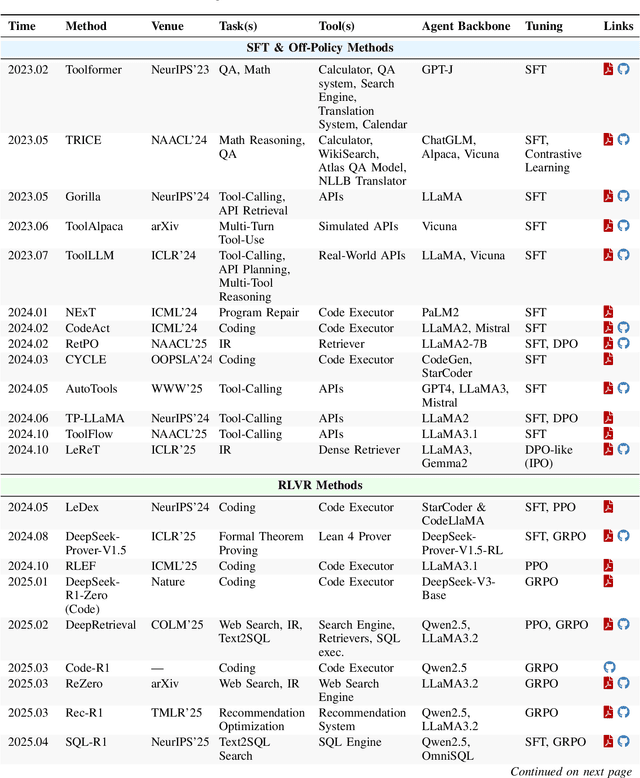
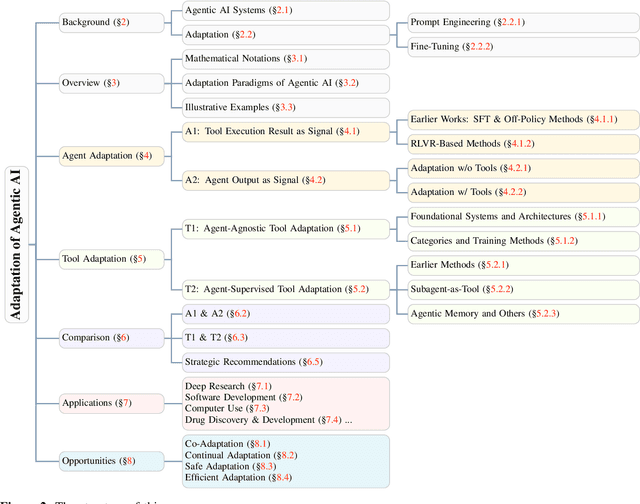
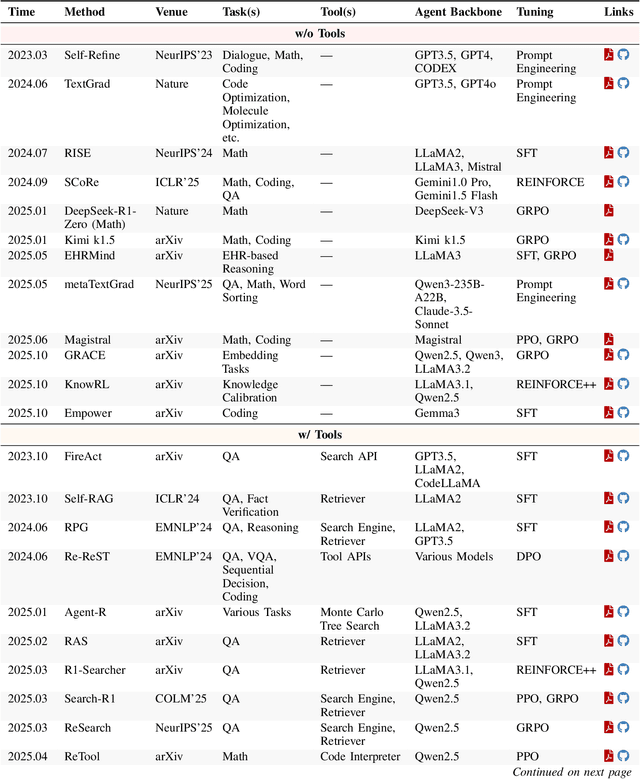
Abstract:Cutting-edge agentic AI systems are built on foundation models that can be adapted to plan, reason, and interact with external tools to perform increasingly complex and specialized tasks. As these systems grow in capability and scope, adaptation becomes a central mechanism for improving performance, reliability, and generalization. In this paper, we unify the rapidly expanding research landscape into a systematic framework that spans both agent adaptations and tool adaptations. We further decompose these into tool-execution-signaled and agent-output-signaled forms of agent adaptation, as well as agent-agnostic and agent-supervised forms of tool adaptation. We demonstrate that this framework helps clarify the design space of adaptation strategies in agentic AI, makes their trade-offs explicit, and provides practical guidance for selecting or switching among strategies during system design. We then review the representative approaches in each category, analyze their strengths and limitations, and highlight key open challenges and future opportunities. Overall, this paper aims to offer a conceptual foundation and practical roadmap for researchers and practitioners seeking to build more capable, efficient, and reliable agentic AI systems.
Projection Embedded Diffusion Bridge for CT Reconstruction from Incomplete Data
Oct 26, 2025Abstract:Reconstructing CT images from incomplete projection data remains challenging due to the ill-posed nature of the problem. Diffusion bridge models have recently shown promise in restoring clean images from their corresponding Filtered Back Projection (FBP) reconstructions, but incorporating data consistency into these models remains largely underexplored. Incorporating data consistency can improve reconstruction fidelity by aligning the reconstructed image with the observed projection data, and can enhance detail recovery by integrating structural information contained in the projections. In this work, we propose the Projection Embedded Diffusion Bridge (PEDB). PEDB introduces a novel reverse stochastic differential equation (SDE) to sample from the distribution of clean images conditioned on both the FBP reconstruction and the incomplete projection data. By explicitly conditioning on the projection data in sampling the clean images, PEDB naturally incorporates data consistency. We embed the projection data into the score function of the reverse SDE. Under certain assumptions, we derive a tractable expression for the posterior score. In addition, we introduce a free parameter to control the level of stochasticity in the reverse process. We also design a discretization scheme for the reverse SDE to mitigate discretization error. Extensive experiments demonstrate that PEDB achieves strong performance in CT reconstruction from three types of incomplete data, including sparse-view, limited-angle, and truncated projections. For each of these types, PEDB outperforms evaluated state-of-the-art diffusion bridge models across standard, noisy, and domain-shift evaluations.
RODS: Robust Optimization Inspired Diffusion Sampling for Detecting and Reducing Hallucination in Generative Models
Jul 16, 2025Abstract:Diffusion models have achieved state-of-the-art performance in generative modeling, yet their sampling procedures remain vulnerable to hallucinations, often stemming from inaccuracies in score approximation. In this work, we reinterpret diffusion sampling through the lens of optimization and introduce RODS (Robust Optimization-inspired Diffusion Sampler), a novel method that detects and corrects high-risk sampling steps using geometric cues from the loss landscape. RODS enforces smoother sampling trajectories and adaptively adjusts perturbations, reducing hallucinations without retraining and at minimal additional inference cost. Experiments on AFHQv2, FFHQ, and 11k-hands demonstrate that RODS improves both sampling fidelity and robustness, detecting over 70% of hallucinated samples and correcting more than 25%, all while avoiding the introduction of new artifacts.
Towards a general-purpose foundation model for fMRI analysis
Jun 11, 2025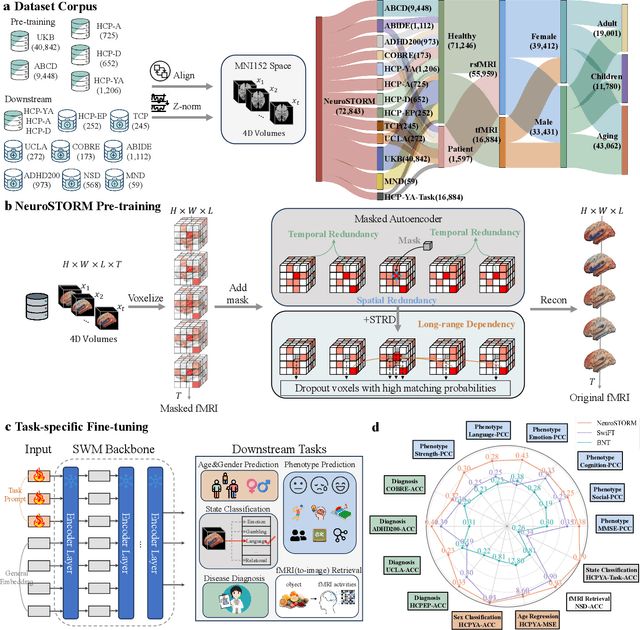

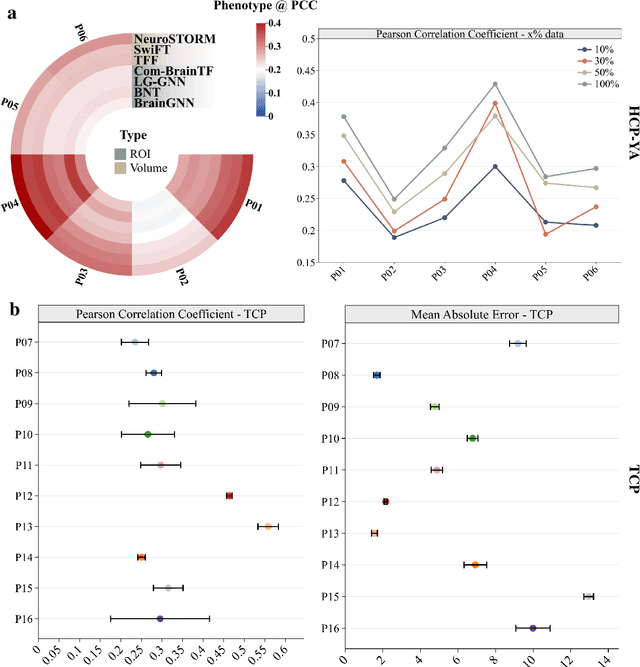
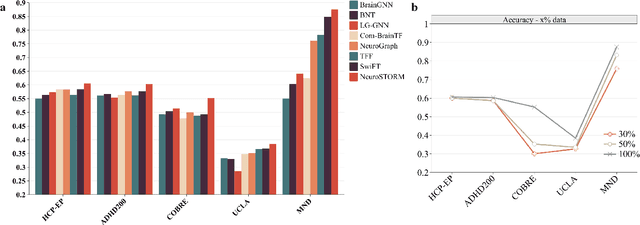
Abstract:Functional Magnetic Resonance Imaging (fMRI) is essential for studying brain function and diagnosing neurological disorders, but current analysis methods face reproducibility and transferability issues due to complex pre-processing and task-specific models. We introduce NeuroSTORM (Neuroimaging Foundation Model with Spatial-Temporal Optimized Representation Modeling), a generalizable framework that directly learns from 4D fMRI volumes and enables efficient knowledge transfer across diverse applications. NeuroSTORM is pre-trained on 28.65 million fMRI frames (>9,000 hours) from over 50,000 subjects across multiple centers and ages 5 to 100. Using a Mamba backbone and a shifted scanning strategy, it efficiently processes full 4D volumes. We also propose a spatial-temporal optimized pre-training approach and task-specific prompt tuning to improve transferability. NeuroSTORM outperforms existing methods across five tasks: age/gender prediction, phenotype prediction, disease diagnosis, fMRI-to-image retrieval, and task-based fMRI classification. It demonstrates strong clinical utility on datasets from hospitals in the U.S., South Korea, and Australia, achieving top performance in disease diagnosis and cognitive phenotype prediction. NeuroSTORM provides a standardized, open-source foundation model to improve reproducibility and transferability in fMRI-based clinical research.
Surf2CT: Cascaded 3D Flow Matching Models for Torso 3D CT Synthesis from Skin Surface
May 29, 2025Abstract:We present Surf2CT, a novel cascaded flow matching framework that synthesizes full 3D computed tomography (CT) volumes of the human torso from external surface scans and simple demographic data (age, sex, height, weight). This is the first approach capable of generating realistic volumetric internal anatomy images solely based on external body shape and demographics, without any internal imaging. Surf2CT proceeds through three sequential stages: (1) Surface Completion, reconstructing a complete signed distance function (SDF) from partial torso scans using conditional 3D flow matching; (2) Coarse CT Synthesis, generating a low-resolution CT volume from the completed SDF and demographic information; and (3) CT Super-Resolution, refining the coarse volume into a high-resolution CT via a patch-wise conditional flow model. Each stage utilizes a 3D-adapted EDM2 backbone trained via flow matching. We trained our model on a combined dataset of 3,198 torso CT scans (approximately 1.13 million axial slices) sourced from Massachusetts General Hospital (MGH) and the AutoPET challenge. Evaluation on 700 paired torso surface-CT cases demonstrated strong anatomical fidelity: organ volumes exhibited small mean percentage differences (range from -11.1% to 4.4%), and muscle/fat body composition metrics matched ground truth with strong correlation (range from 0.67 to 0.96). Lung localization had minimal bias (mean difference -2.5 mm), and surface completion significantly improved metrics (Chamfer distance: from 521.8 mm to 2.7 mm; Intersection-over-Union: from 0.87 to 0.98). Surf2CT establishes a new paradigm for non-invasive internal anatomical imaging using only external data, opening opportunities for home-based healthcare, preventive medicine, and personalized clinical assessments without the risks associated with conventional imaging techniques.
Cascaded 3D Diffusion Models for Whole-body 3D 18-F FDG PET/CT synthesis from Demographics
May 28, 2025Abstract:We propose a cascaded 3D diffusion model framework to synthesize high-fidelity 3D PET/CT volumes directly from demographic variables, addressing the growing need for realistic digital twins in oncologic imaging, virtual trials, and AI-driven data augmentation. Unlike deterministic phantoms, which rely on predefined anatomical and metabolic templates, our method employs a two-stage generative process. An initial score-based diffusion model synthesizes low-resolution PET/CT volumes from demographic variables alone, providing global anatomical structures and approximate metabolic activity. This is followed by a super-resolution residual diffusion model that refines spatial resolution. Our framework was trained on 18-F FDG PET/CT scans from the AutoPET dataset and evaluated using organ-wise volume and standardized uptake value (SUV) distributions, comparing synthetic and real data between demographic subgroups. The organ-wise comparison demonstrated strong concordance between synthetic and real images. In particular, most deviations in metabolic uptake values remained within 3-5% of the ground truth in subgroup analysis. These findings highlight the potential of cascaded 3D diffusion models to generate anatomically and metabolically accurate PET/CT images, offering a robust alternative to traditional phantoms and enabling scalable, population-informed synthetic imaging for clinical and research applications.
OWT: A Foundational Organ-Wise Tokenization Framework for Medical Imaging
May 08, 2025Abstract:Recent advances in representation learning often rely on holistic, black-box embeddings that entangle multiple semantic components, limiting interpretability and generalization. These issues are especially critical in medical imaging. To address these limitations, we propose an Organ-Wise Tokenization (OWT) framework with a Token Group-based Reconstruction (TGR) training paradigm. Unlike conventional approaches that produce holistic features, OWT explicitly disentangles an image into separable token groups, each corresponding to a distinct organ or semantic entity. Our design ensures each token group encapsulates organ-specific information, boosting interpretability, generalization, and efficiency while allowing fine-grained control in downstream tasks. Experiments on CT and MRI datasets demonstrate the effectiveness of OWT in not only achieving strong image reconstruction and segmentation performance, but also enabling novel semantic-level generation and retrieval applications that are out of reach for standard holistic embedding methods. These findings underscore the potential of OWT as a foundational framework for semantically disentangled representation learning, offering broad scalability and applicability to real-world medical imaging scenarios and beyond.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge