Yuting He
AniGen: Unified $S^3$ Fields for Animatable 3D Asset Generation
Apr 14, 2026Abstract:Animatable 3D assets, defined as geometry equipped with an articulated skeleton and skinning weights, are fundamental to interactive graphics, embodied agents, and animation production. While recent 3D generative models can synthesize visually plausible shapes from images, the results are typically static. Obtaining usable rigs via post-hoc auto-rigging is brittle and often produces skeletons that are topologically inconsistent with the generated geometry. We present AniGen, a unified framework that directly generates animate-ready 3D assets conditioned on a single image. Our key insight is to represent shape, skeleton, and skinning as mutually consistent $S^3$ Fields (Shape, Skeleton, Skin) defined over a shared spatial domain. To enable the robust learning of these fields, we introduce two technical innovations: (i) a confidence-decaying skeleton field that explicitly handles the geometric ambiguity of bone prediction at Voronoi boundaries, and (ii) a dual skin feature field that decouples skinning weights from specific joint counts, allowing a fixed-architecture network to predict rigs of arbitrary complexity. Built upon a two-stage flow-matching pipeline, AniGen first synthesizes a sparse structural scaffold and then generates dense geometry and articulation in a structured latent space. Extensive experiments demonstrate that AniGen substantially outperforms state-of-the-art sequential baselines in rig validity and animation quality, generalizing effectively to in-the-wild images across diverse categories including animals, humanoids, and machinery. Homepage: https://yihua7.github.io/AniGen-web/
Generative Data-engine Foundation Model for Universal Few-shot 2D Vascular Image Segmentation
Apr 12, 2026Abstract:The segmentation of 2D vascular structures via deep learning holds significant clinical value but is hindered by the scarcity of annotated data, severely limiting its widespread application. Developing a universal few-shot vascular segmentation model is highly desirable, yet remains challenging due to the need for extensive training and the inherent complexities of vascular imaging. In this work, we propose UniVG (Generative Data-engine Foundation Model for Universal Few-shot 2D Vascular Image Segmentation), a novel approach that learns the compositionality of vascular images and constructing a generative foundation model for robust vascular segmentation. UniVG enables the synthesis and learning of diverse and realistic vascular images through two key innovations: 1) Compositional learning for flexible and diverse vascular synthesis: It decomposes and recombines vascular structures with varying morphological features and diverse foreground-background configurations to generate richly diverse synthetic image-label pairs. 2) Few-shot generative adaptation for transferable segmentation: It fine-tunes pre-trained models with minimal annotated data to bridge the gap between synthetic and real vascular domains, synthesizing authentic and diverse vessel images for downstream few-shot vascular segmentation learning. To support our approach, we develop UniVG-58K, a large dataset comprising 58,689 vascular images across five imaging modalities, facilitating robust large-scale generative pre-training. Extensive experiments on 11 vessel segmentation tasks cross 5 modalties (only with 5 labeled images on each task) demonstrate that UniVG achieves performance comparable to fully supervised models, significantly reducing data collection and annotation costs. All code and datasets will be made publicly available at https://github.com/XinAloha/UniVG.
Human Gaze-based Dual Teacher Guidance Learning for Semi-Supervised Medical Image Segmentation
Apr 12, 2026Abstract:In the field of medical image segmentation, the scarcity of labeled data poses a major challenge for existing models to accurately perceive target regions. Compared with manual annotation, gaze data is easier and cheaper to obtain. As a classical semi-supervised learning framework, mean-teacher can effectively use a large number of unlabeled medical images for stable training through self-teaching and collaborative optimization. Our study is based on the mean-teacher framework. By combining gaze data, it aims to address two crucial issues in semi-supervised medical image segmentation: 1) expand the scale and diversity of the dataset with limited labeled data; 2) enhance the network's perception ability. We propose the Human Gaze-based Dual Teacher Guidance Learning model (HG-DTGL). In this model, human gaze serves as an additional hidden `teacher' in the mean-teacher architecture. We introduce the GazeMix to generate reliable mixed data to expand the diversity and scale of the dataset, and the Multi-scale Gaze Perception (MGP) module is used to extract the multi-scale perception of the network. A Gaze Loss is designed to align the model's perception with human gaze. We have verified HG-DTGL on multiple datasets of different modalities and achieved superior performance on a total of ten different organs/tissues, with extensive experiments. This demonstrates that our method has strong generalization ability for medical images of different modalities, and shows the great application potential of gaze data in semi-supervised medical image segmentation.
Predictive Reasoning with Augmented Anomaly Contrastive Learning for Compositional Visual Relations
Mar 01, 2026Abstract:While visual reasoning for simple analogies has received significant attention, compositional visual relations (CVR) remain relatively unexplored due to their greater complexity. To solve CVR tasks, we propose Predictive Reasoning with Augmented Anomaly Contrastive Learning (PR-A$^2$CL), \ie, to identify an outlier image given three other images that follow the same compositional rules. To address the challenge of modelling abundant compositional rules, an Augmented Anomaly Contrastive Learning is designed to distil discriminative and generalizable features by maximizing similarity among normal instances while minimizing similarity between normal and anomalous outliers. More importantly, a predict-and-verify paradigm is introduced for rule-based reasoning, in which a series of Predictive Anomaly Reasoning Blocks (PARBs) iteratively leverage features from three out of the four images to predict those of the remaining one. Throughout the subsequent verification stage, the PARBs progressively pinpoint the specific discrepancies attributable to the underlying rules. Experimental results on SVRT, CVR and MC$^2$R datasets show that PR-A$^2$CL significantly outperforms state-of-the-art reasoning models.
MRI Contrast Enhancement Kinetics World Model
Feb 22, 2026Abstract:Clinical MRI contrast acquisition suffers from inefficient information yield, which presents as a mismatch between the risky and costly acquisition protocol and the fixed and sparse acquisition sequence. Applying world models to simulate the contrast enhancement kinetics in the human body enables continuous contrast-free dynamics. However, the low temporal resolution in MRI acquisition restricts the training of world models, leading to a sparsely sampled dataset. Directly training a generative model to capture the kinetics leads to two limitations: (a) Due to the absence of data on missing time, the model tends to overfit to irrelevant features, leading to content distortion. (b) Due to the lack of continuous temporal supervision, the model fails to learn the continuous kinetics law over time, causing temporal discontinuities. For the first time, we propose MRI Contrast Enhancement Kinetics World model (MRI CEKWorld) with SpatioTemporal Consistency Learning (STCL). For (a), guided by the spatial law that patient-level structures remain consistent during enhancement, we propose Latent Alignment Learning (LAL) that constructs a patient-specific template to constrain contents to align with this template. For (b), guided by the temporal law that the kinetics follow a consistent smooth trend, we propose Latent Difference Learning (LDL) which extends the unobserved intervals by interpolation and constrains smooth variations in the latent space among interpolated sequences. Extensive experiments on two datasets show our MRI CEKWorld achieves better realistic contents and kinetics. Codes will be available at https://github.com/DD0922/MRI-Contrast-Enhancement-Kinetics-World-Model.
Free Lunch in Medical Image Foundation Model Pre-training via Randomized Synthesis and Disentanglement
Feb 12, 2026Abstract:Medical image foundation models (MIFMs) have demonstrated remarkable potential for a wide range of clinical tasks, yet their development is constrained by the scarcity, heterogeneity, and high cost of large-scale annotated datasets. Here, we propose RaSD (Randomized Synthesis and Disentanglement), a scalable framework for pre-training MIFMs entirely on synthetic data. By modeling anatomical structures and appearance variations with randomized Gaussian distributions, RaSD exposes models to sufficient multi-scale structural and appearance perturbations, forcing them to rely on invariant and task-relevant anatomical cues rather than dataset-specific textures, thereby enabling robust and transferable representation learning. We pre-trained RaSD on 1.2 million 3D volumes and 9.6 million 2D images, and extensively evaluated the resulting models across 6 imaging modalities, 48 datasets, and 56 downstream tasks. Across all evaluated downstream tasks, RaSD consistently outperforms training-from-scratch models, achieves the best performance on 17 tasks, and remains comparable to models pre-trained on large real datasets in most others. These results demonstrate that the capacity of synthetic data alone to drive robust representation learning. Our findings establish a paradigm shift in medical AI, demonstrating that synthetic data can serve as a "free lunch" for scalable, privacy-preserving, and clinically generalizable foundation models.
Dynamic Stream Network for Combinatorial Explosion Problem in Deformable Medical Image Registration
Dec 22, 2025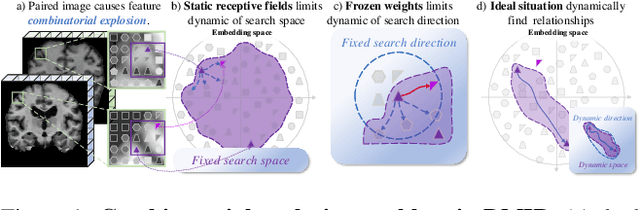
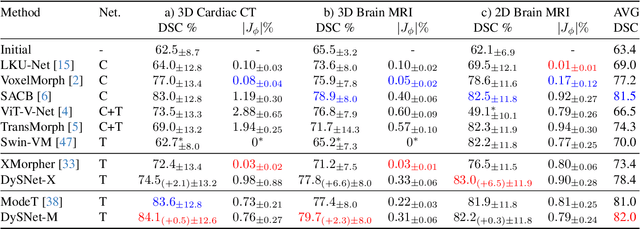
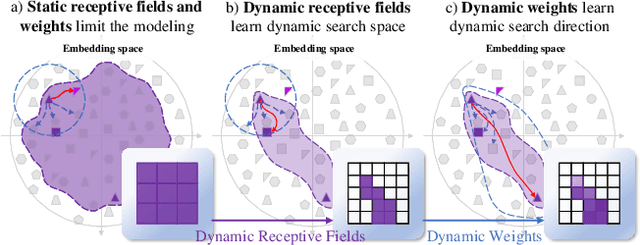
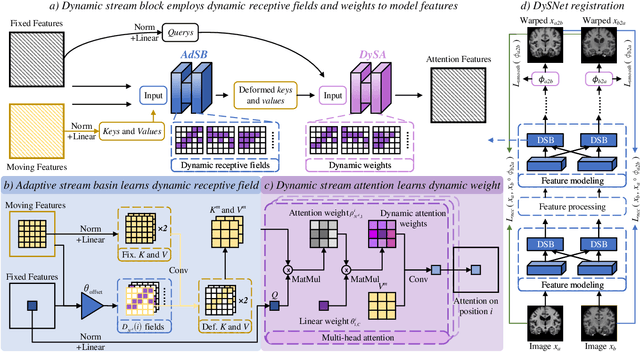
Abstract:Combinatorial explosion problem caused by dual inputs presents a critical challenge in Deformable Medical Image Registration (DMIR). Since DMIR processes two images simultaneously as input, the combination relationships between features has grown exponentially, ultimately the model considers more interfering features during the feature modeling process. Introducing dynamics in the receptive fields and weights of the network enable the model to eliminate the interfering features combination and model the potential feature combination relationships. In this paper, we propose the Dynamic Stream Network (DySNet), which enables the receptive fields and weights to be dynamically adjusted. This ultimately enables the model to ignore interfering feature combinations and model the potential feature relationships. With two key innovations: 1) Adaptive Stream Basin (AdSB) module dynamically adjusts the shape of the receptive field, thereby enabling the model to focus on the feature relationships with greater correlation. 2) Dynamic Stream Attention (DySA) mechanism generates dynamic weights to search for more valuable feature relationships. Extensive experiments have shown that DySNet consistently outperforms the most advanced DMIR methods, highlighting its outstanding generalization ability. Our code will be released on the website: https://github.com/ShaochenBi/DySNet.
DeepFeature: Iterative Context-aware Feature Generation for Wearable Biosignals
Dec 09, 2025Abstract:Biosignals collected from wearable devices are widely utilized in healthcare applications. Machine learning models used in these applications often rely on features extracted from biosignals due to their effectiveness, lower data dimensionality, and wide compatibility across various model architectures. However, existing feature extraction methods often lack task-specific contextual knowledge, struggle to identify optimal feature extraction settings in high-dimensional feature space, and are prone to code generation and automation errors. In this paper, we propose DeepFeature, the first LLM-empowered, context-aware feature generation framework for wearable biosignals. DeepFeature introduces a multi-source feature generation mechanism that integrates expert knowledge with task settings. It also employs an iterative feature refinement process that uses feature assessment-based feedback for feature re-selection. Additionally, DeepFeature utilizes a robust multi-layer filtering and verification approach for robust feature-to-code translation to ensure that the extraction functions run without crashing. Experimental evaluation results show that DeepFeature achieves an average AUROC improvement of 4.21-9.67% across eight diverse tasks compared to baseline methods. It outperforms state-of-the-art approaches on five tasks while maintaining comparable performance on the remaining tasks.
TimeSeriesScientist: A General-Purpose AI Agent for Time Series Analysis
Oct 02, 2025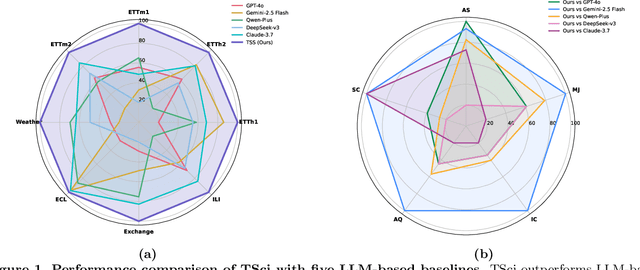
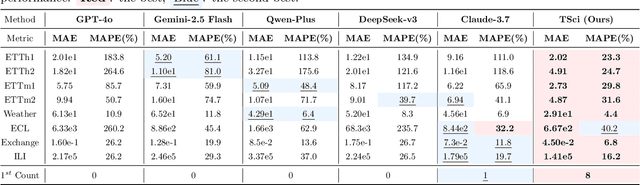
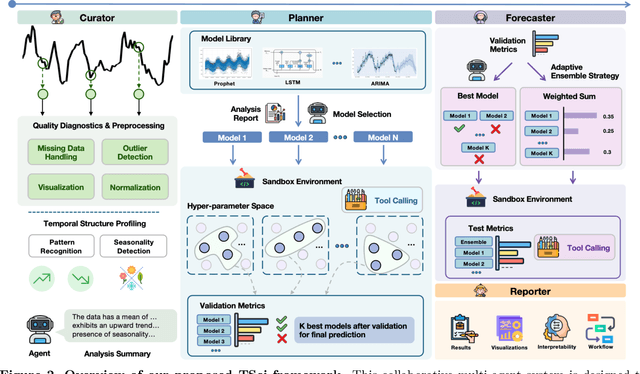
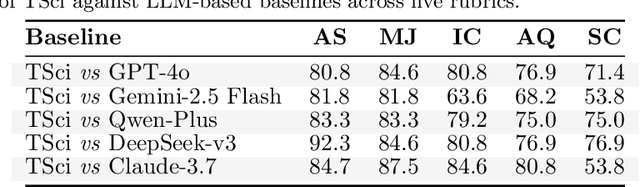
Abstract:Time series forecasting is central to decision-making in domains as diverse as energy, finance, climate, and public health. In practice, forecasters face thousands of short, noisy series that vary in frequency, quality, and horizon, where the dominant cost lies not in model fitting, but in the labor-intensive preprocessing, validation, and ensembling required to obtain reliable predictions. Prevailing statistical and deep learning models are tailored to specific datasets or domains and generalize poorly. A general, domain-agnostic framework that minimizes human intervention is urgently in demand. In this paper, we introduce TimeSeriesScientist (TSci), the first LLM-driven agentic framework for general time series forecasting. The framework comprises four specialized agents: Curator performs LLM-guided diagnostics augmented by external tools that reason over data statistics to choose targeted preprocessing; Planner narrows the hypothesis space of model choice by leveraging multi-modal diagnostics and self-planning over the input; Forecaster performs model fitting and validation and, based on the results, adaptively selects the best model configuration as well as ensemble strategy to make final predictions; and Reporter synthesizes the whole process into a comprehensive, transparent report. With transparent natural-language rationales and comprehensive reports, TSci transforms the forecasting workflow into a white-box system that is both interpretable and extensible across tasks. Empirical results on eight established benchmarks demonstrate that TSci consistently outperforms both statistical and LLM-based baselines, reducing forecast error by an average of 10.4% and 38.2%, respectively. Moreover, TSci produces a clear and rigorous report that makes the forecasting workflow more transparent and interpretable.
Cardiac-CLIP: A Vision-Language Foundation Model for 3D Cardiac CT Images
Jul 29, 2025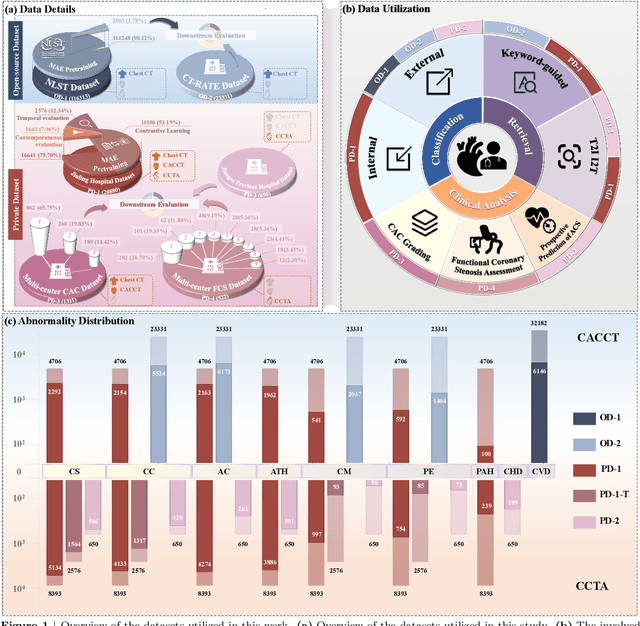
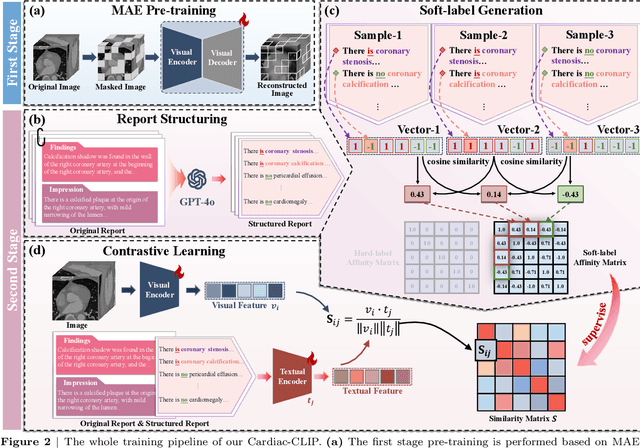
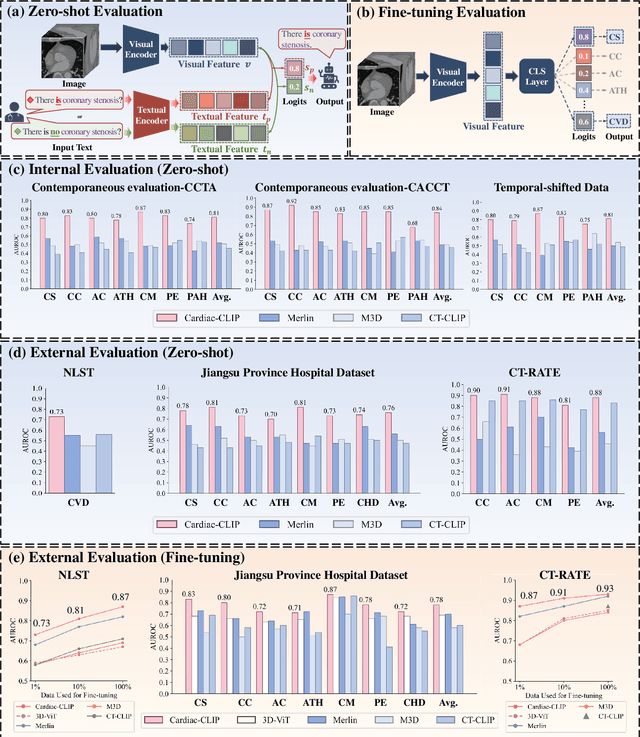
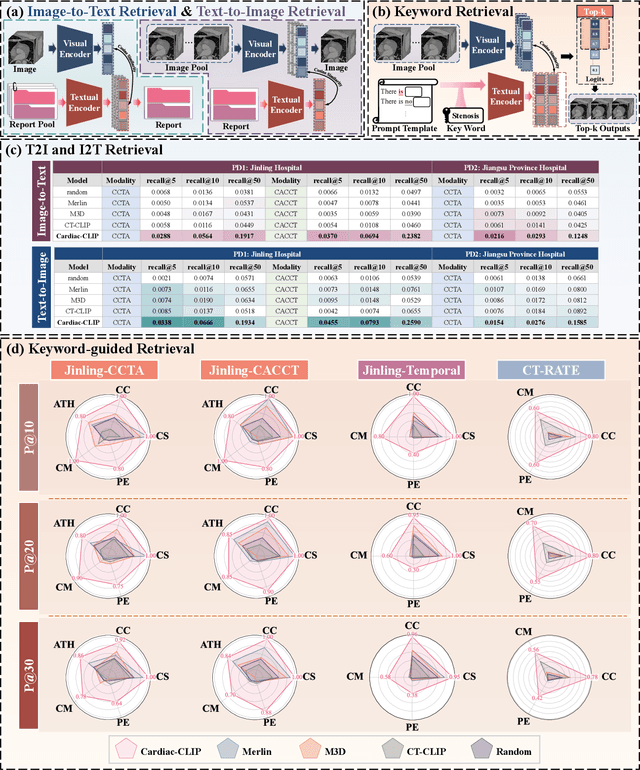
Abstract:Foundation models have demonstrated remarkable potential in medical domain. However, their application to complex cardiovascular diagnostics remains underexplored. In this paper, we present Cardiac-CLIP, a multi-modal foundation model designed for 3D cardiac CT images. Cardiac-CLIP is developed through a two-stage pre-training strategy. The first stage employs a 3D masked autoencoder (MAE) to perform self-supervised representation learning from large-scale unlabeled volumetric data, enabling the visual encoder to capture rich anatomical and contextual features. In the second stage, contrastive learning is introduced to align visual and textual representations, facilitating cross-modal understanding. To support the pre-training, we collect 16641 real clinical CT scans, supplemented by 114k publicly available data. Meanwhile, we standardize free-text radiology reports into unified templates and construct the pathology vectors according to diagnostic attributes, based on which the soft-label matrix is generated to supervise the contrastive learning process. On the other hand, to comprehensively evaluate the effectiveness of Cardiac-CLIP, we collect 6,722 real-clinical data from 12 independent institutions, along with the open-source data to construct the evaluation dataset. Specifically, Cardiac-CLIP is comprehensively evaluated across multiple tasks, including cardiovascular abnormality classification, information retrieval and clinical analysis. Experimental results demonstrate that Cardiac-CLIP achieves state-of-the-art performance across various downstream tasks in both internal and external data. Particularly, Cardiac-CLIP exhibits great effectiveness in supporting complex clinical tasks such as the prospective prediction of acute coronary syndrome, which is notoriously difficult in real-world scenarios.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge