Yaolei Qi
Rethinking the Detail-Preserved Completion of Complex Tubular Structures based on Point Cloud: a Dataset and a Benchmark
Aug 25, 2025



Abstract:Complex tubular structures are essential in medical imaging and computer-assisted diagnosis, where their integrity enhances anatomical visualization and lesion detection. However, existing segmentation algorithms struggle with structural discontinuities, particularly in severe clinical cases such as coronary artery stenosis and vessel occlusions, which leads to undesired discontinuity and compromising downstream diagnostic accuracy. Therefore, it is imperative to reconnect discontinuous structures to ensure their completeness. In this study, we explore the tubular structure completion based on point cloud for the first time and establish a Point Cloud-based Coronary Artery Completion (PC-CAC) dataset, which is derived from real clinical data. This dataset provides a novel benchmark for tubular structure completion. Additionally, we propose TSRNet, a Tubular Structure Reconnection Network that integrates a detail-preservated feature extractor, a multiple dense refinement strategy, and a global-to-local loss function to ensure accurate reconnection while maintaining structural integrity. Comprehensive experiments on our PC-CAC and two additional public datasets (PC-ImageCAS and PC-PTR) demonstrate that our method consistently outperforms state-of-the-art approaches across multiple evaluation metrics, setting a new benchmark for point cloud-based tubular structure reconstruction. Our benchmark is available at https://github.com/YaoleiQi/PCCAC.
Cardiac-CLIP: A Vision-Language Foundation Model for 3D Cardiac CT Images
Jul 29, 2025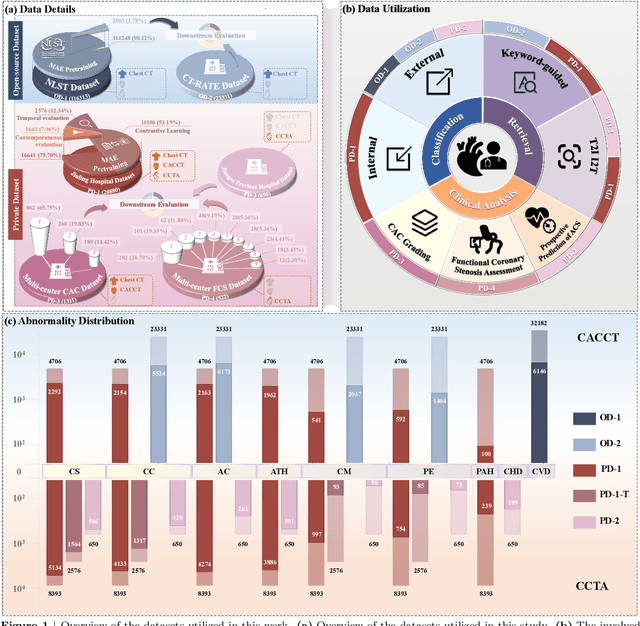
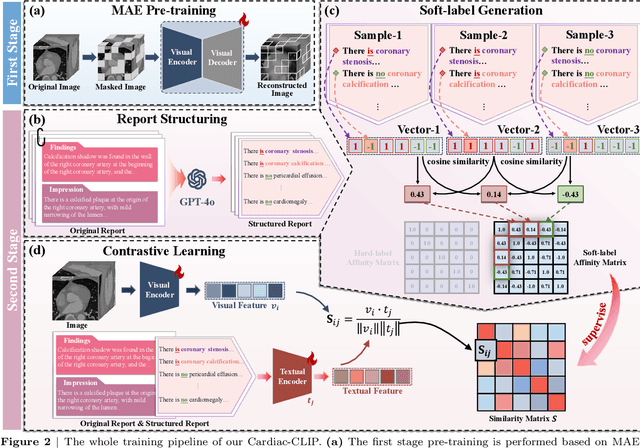
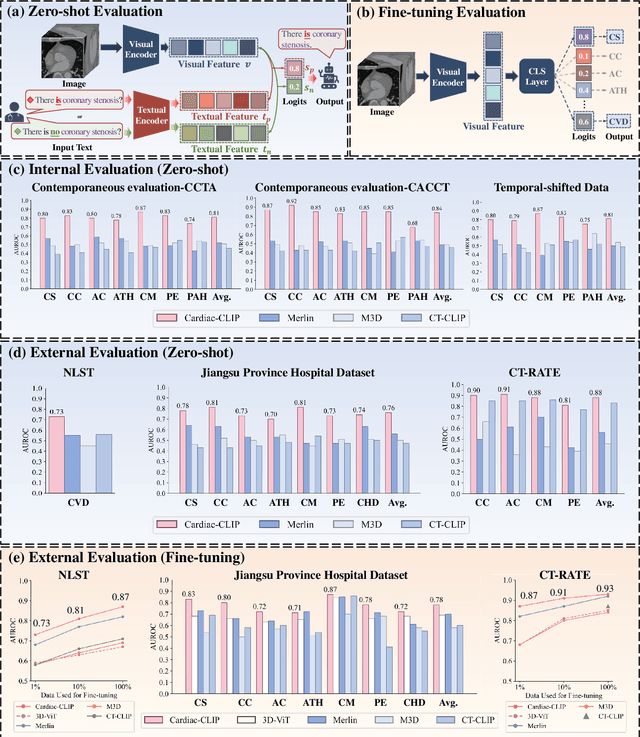
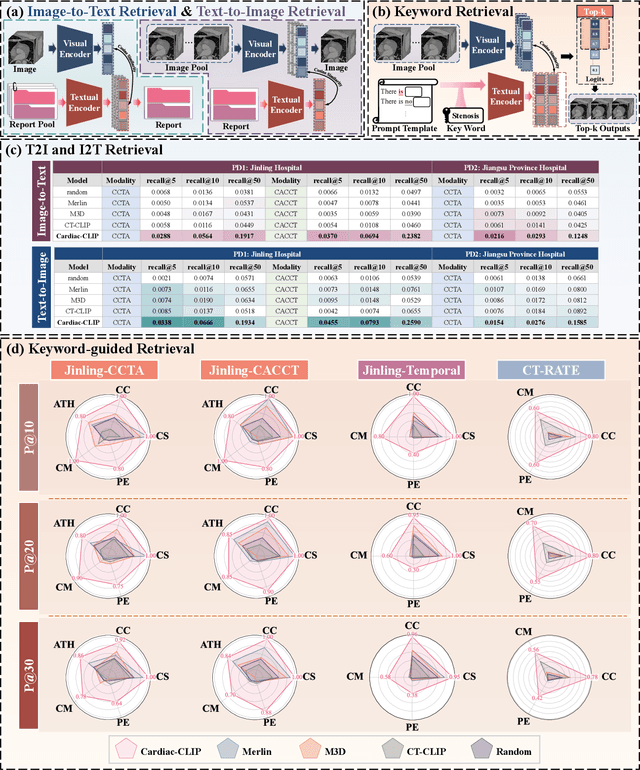
Abstract:Foundation models have demonstrated remarkable potential in medical domain. However, their application to complex cardiovascular diagnostics remains underexplored. In this paper, we present Cardiac-CLIP, a multi-modal foundation model designed for 3D cardiac CT images. Cardiac-CLIP is developed through a two-stage pre-training strategy. The first stage employs a 3D masked autoencoder (MAE) to perform self-supervised representation learning from large-scale unlabeled volumetric data, enabling the visual encoder to capture rich anatomical and contextual features. In the second stage, contrastive learning is introduced to align visual and textual representations, facilitating cross-modal understanding. To support the pre-training, we collect 16641 real clinical CT scans, supplemented by 114k publicly available data. Meanwhile, we standardize free-text radiology reports into unified templates and construct the pathology vectors according to diagnostic attributes, based on which the soft-label matrix is generated to supervise the contrastive learning process. On the other hand, to comprehensively evaluate the effectiveness of Cardiac-CLIP, we collect 6,722 real-clinical data from 12 independent institutions, along with the open-source data to construct the evaluation dataset. Specifically, Cardiac-CLIP is comprehensively evaluated across multiple tasks, including cardiovascular abnormality classification, information retrieval and clinical analysis. Experimental results demonstrate that Cardiac-CLIP achieves state-of-the-art performance across various downstream tasks in both internal and external data. Particularly, Cardiac-CLIP exhibits great effectiveness in supporting complex clinical tasks such as the prospective prediction of acute coronary syndrome, which is notoriously difficult in real-world scenarios.
PathFL: Multi-Alignment Federated Learning for Pathology Image Segmentation
May 28, 2025Abstract:Pathology image segmentation across multiple centers encounters significant challenges due to diverse sources of heterogeneity including imaging modalities, organs, and scanning equipment, whose variability brings representation bias and impedes the development of generalizable segmentation models. In this paper, we propose PathFL, a novel multi-alignment Federated Learning framework for pathology image segmentation that addresses these challenges through three-level alignment strategies of image, feature, and model aggregation. Firstly, at the image level, a collaborative style enhancement module aligns and diversifies local data by facilitating style information exchange across clients. Secondly, at the feature level, an adaptive feature alignment module ensures implicit alignment in the representation space by infusing local features with global insights, promoting consistency across heterogeneous client features learning. Finally, at the model aggregation level, a stratified similarity aggregation strategy hierarchically aligns and aggregates models on the server, using layer-specific similarity to account for client discrepancies and enhance global generalization. Comprehensive evaluations on four sets of heterogeneous pathology image datasets, encompassing cross-source, cross-modality, cross-organ, and cross-scanner variations, validate the effectiveness of our PathFL in achieving better performance and robustness against data heterogeneity.
FCaS: Fine-grained Cardiac Image Synthesis based on 3D Template Conditional Diffusion Model
Mar 12, 2025Abstract:Solving medical imaging data scarcity through semantic image generation has attracted significant attention in recent years. However, existing methods primarily focus on generating whole-organ or large-tissue structures, showing limited effectiveness for organs with fine-grained structure. Due to stringent topological consistency, fragile coronary features, and complex 3D morphological heterogeneity in cardiac imaging, accurately reconstructing fine-grained anatomical details of the heart remains a great challenge. To address this problem, in this paper, we propose the Fine-grained Cardiac image Synthesis(FCaS) framework, established on 3D template conditional diffusion model. FCaS achieves precise cardiac structure generation using Template-guided Conditional Diffusion Model (TCDM) through bidirectional mechanisms, which provides the fine-grained topological structure information of target image through the guidance of template. Meanwhile, we design a deformable Mask Generation Module (MGM) to mitigate the scarcity of high-quality and diverse reference mask in the generation process. Furthermore, to alleviate the confusion caused by imprecise synthetic images, we propose a Confidence-aware Adaptive Learning (CAL) strategy to facilitate the pre-training of downstream segmentation tasks. Specifically, we introduce the Skip-Sampling Variance (SSV) estimation to obtain confidence maps, which are subsequently employed to rectify the pre-training on downstream tasks. Experimental results demonstrate that images generated from FCaS achieves state-of-the-art performance in topological consistency and visual quality, which significantly facilitates the downstream tasks as well. Code will be released in the future.
DSCENet: Dynamic Screening and Clinical-Enhanced Multimodal Fusion for MPNs Subtype Classification
Jul 11, 2024
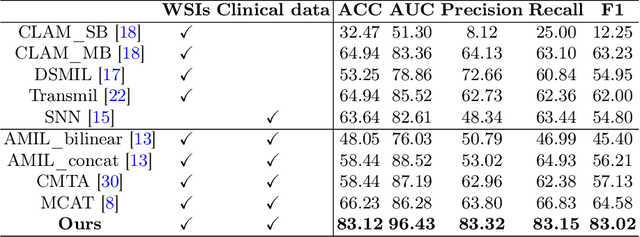


Abstract:The precise subtype classification of myeloproliferative neoplasms (MPNs) based on multimodal information, which assists clinicians in diagnosis and long-term treatment plans, is of great clinical significance. However, it remains a great challenging task due to the lack of diagnostic representativeness for local patches and the absence of diagnostic-relevant features from a single modality. In this paper, we propose a Dynamic Screening and Clinical-Enhanced Network (DSCENet) for the subtype classification of MPNs on the multimodal fusion of whole slide images (WSIs) and clinical information. (1) A dynamic screening module is proposed to flexibly adapt the feature learning of local patches, reducing the interference of irrelevant features and enhancing their diagnostic representativeness. (2) A clinical-enhanced fusion module is proposed to integrate clinical indicators to explore complementary features across modalities, providing comprehensive diagnostic information. Our approach has been validated on the real clinical data, achieving an increase of 7.91% AUC and 16.89% accuracy compared with the previous state-of-the-art (SOTA) methods. The code is available at https://github.com/yuanzhang7/DSCENet.
FedSODA: Federated Cross-assessment and Dynamic Aggregation for Histopathology Segmentation
Dec 20, 2023


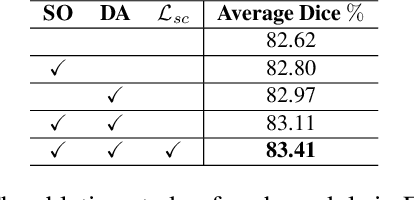
Abstract:Federated learning (FL) for histopathology image segmentation involving multiple medical sites plays a crucial role in advancing the field of accurate disease diagnosis and treatment. However, it is still a task of great challenges due to the sample imbalance across clients and large data heterogeneity from disparate organs, variable segmentation tasks, and diverse distribution. Thus, we propose a novel FL approach for histopathology nuclei and tissue segmentation, FedSODA, via synthetic-driven cross-assessment operation (SO) and dynamic stratified-layer aggregation (DA). Our SO constructs a cross-assessment strategy to connect clients and mitigate the representation bias under sample imbalance. Our DA utilizes layer-wise interaction and dynamic aggregation to diminish heterogeneity and enhance generalization. The effectiveness of our FedSODA has been evaluated on the most extensive histopathology image segmentation dataset from 7 independent datasets. The code is available at https://github.com/yuanzhang7/FedSODA.
Dynamic Snake Convolution based on Topological Geometric Constraints for Tubular Structure Segmentation
Jul 17, 2023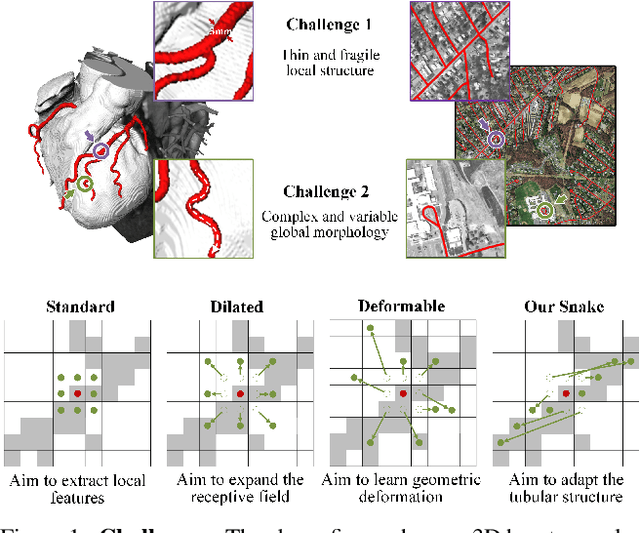
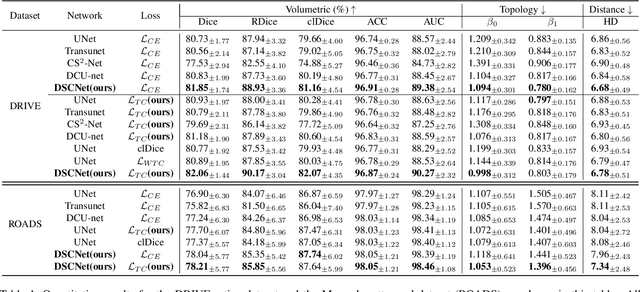

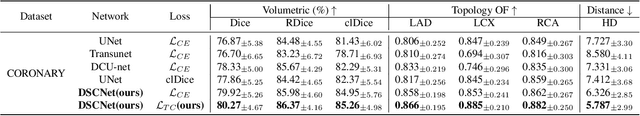
Abstract:Accurate segmentation of topological tubular structures, such as blood vessels and roads, is crucial in various fields, ensuring accuracy and efficiency in downstream tasks. However, many factors complicate the task, including thin local structures and variable global morphologies. In this work, we note the specificity of tubular structures and use this knowledge to guide our DSCNet to simultaneously enhance perception in three stages: feature extraction, feature fusion, and loss constraint. First, we propose a dynamic snake convolution to accurately capture the features of tubular structures by adaptively focusing on slender and tortuous local structures. Subsequently, we propose a multi-view feature fusion strategy to complement the attention to features from multiple perspectives during feature fusion, ensuring the retention of important information from different global morphologies. Finally, a continuity constraint loss function, based on persistent homology, is proposed to constrain the topological continuity of the segmentation better. Experiments on 2D and 3D datasets show that our DSCNet provides better accuracy and continuity on the tubular structure segmentation task compared with several methods. Our codes will be publicly available.
Partial Vessels Annotation-based Coronary Artery Segmentation with Self-training and Prototype Learning
Jul 10, 2023



Abstract:Coronary artery segmentation on coronary-computed tomography angiography (CCTA) images is crucial for clinical use. Due to the expertise-required and labor-intensive annotation process, there is a growing demand for the relevant label-efficient learning algorithms. To this end, we propose partial vessels annotation (PVA) based on the challenges of coronary artery segmentation and clinical diagnostic characteristics. Further, we propose a progressive weakly supervised learning framework to achieve accurate segmentation under PVA. First, our proposed framework learns the local features of vessels to propagate the knowledge to unlabeled regions. Subsequently, it learns the global structure by utilizing the propagated knowledge, and corrects the errors introduced in the propagation process. Finally, it leverages the similarity between feature embeddings and the feature prototype to enhance testing outputs. Experiments on clinical data reveals that our proposed framework outperforms the competing methods under PVA (24.29% vessels), and achieves comparable performance in trunk continuity with the baseline model using full annotation (100% vessels).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge