Daniel Rubin
Towards trustworthy seizure onset detection using workflow notes
Jun 14, 2023



Abstract:A major barrier to deploying healthcare AI models is their trustworthiness. One form of trustworthiness is a model's robustness across different subgroups: while existing models may exhibit expert-level performance on aggregate metrics, they often rely on non-causal features, leading to errors in hidden subgroups. To take a step closer towards trustworthy seizure onset detection from EEG, we propose to leverage annotations that are produced by healthcare personnel in routine clinical workflows -- which we refer to as workflow notes -- that include multiple event descriptions beyond seizures. Using workflow notes, we first show that by scaling training data to an unprecedented level of 68,920 EEG hours, seizure onset detection performance significantly improves (+12.3 AUROC points) compared to relying on smaller training sets with expensive manual gold-standard labels. Second, we reveal that our binary seizure onset detection model underperforms on clinically relevant subgroups (e.g., up to a margin of 6.5 AUROC points between pediatrics and adults), while having significantly higher false positives on EEG clips showing non-epileptiform abnormalities compared to any EEG clip (+19 FPR points). To improve model robustness to hidden subgroups, we train a multilabel model that classifies 26 attributes other than seizures, such as spikes, slowing, and movement artifacts. We find that our multilabel model significantly improves overall seizure onset detection performance (+5.9 AUROC points) while greatly improving performance among subgroups (up to +8.3 AUROC points), and decreases false positives on non-epileptiform abnormalities by 8 FPR points. Finally, we propose a clinical utility metric based on false positives per 24 EEG hours and find that our multilabel model improves this clinical utility metric by a factor of 2x across different clinical settings.
Exploring Image Augmentations for Siamese Representation Learning with Chest X-Rays
Jan 30, 2023



Abstract:Image augmentations are quintessential for effective visual representation learning across self-supervised learning techniques. While augmentation strategies for natural imaging have been studied extensively, medical images are vastly different from their natural counterparts. Thus, it is unknown whether common augmentation strategies employed in Siamese representation learning generalize to medical images and to what extent. To address this challenge, in this study, we systematically assess the effect of various augmentations on the quality and robustness of the learned representations. We train and evaluate Siamese Networks for abnormality detection on chest X-Rays across three large datasets (MIMIC-CXR, CheXpert and VinDR-CXR). We investigate the efficacy of the learned representations through experiments involving linear probing, fine-tuning, zero-shot transfer, and data efficiency. Finally, we identify a set of augmentations that yield robust representations that generalize well to both out-of-distribution data and diseases, while outperforming supervised baselines using just zero-shot transfer and linear probes by up to 20%.
ATCON: Attention Consistency for Vision Models
Oct 18, 2022
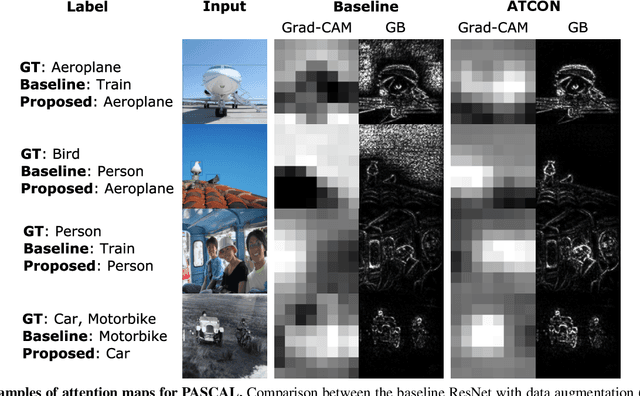
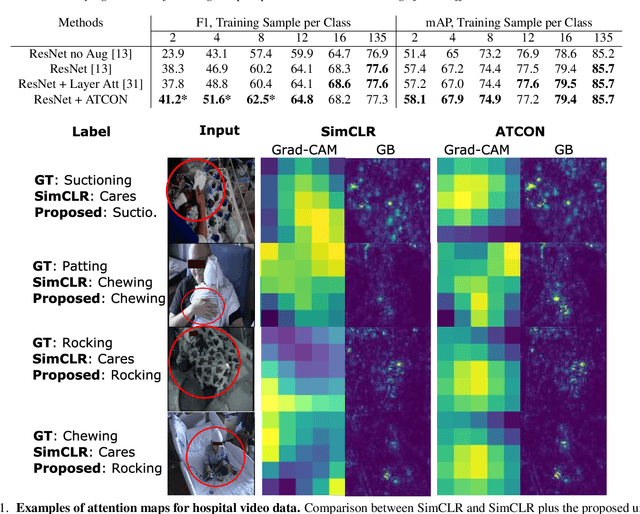

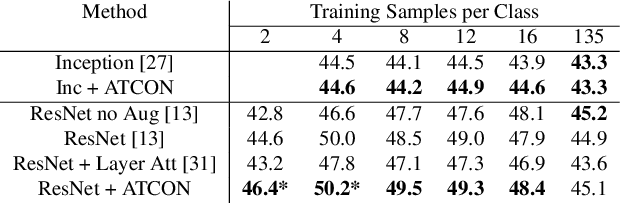
Abstract:Attention--or attribution--maps methods are methods designed to highlight regions of the model's input that were discriminative for its predictions. However, different attention maps methods can highlight different regions of the input, with sometimes contradictory explanations for a prediction. This effect is exacerbated when the training set is small. This indicates that either the model learned incorrect representations or that the attention maps methods did not accurately estimate the model's representations. We propose an unsupervised fine-tuning method that optimizes the consistency of attention maps and show that it improves both classification performance and the quality of attention maps. We propose an implementation for two state-of-the-art attention computation methods, Grad-CAM and Guided Backpropagation, which relies on an input masking technique. We also show results on Grad-CAM and Integrated Gradients in an ablation study. We evaluate this method on our own dataset of event detection in continuous video recordings of hospital patients aggregated and curated for this work. As a sanity check, we also evaluate the proposed method on PASCAL VOC and SVHN. With the proposed method, with small training sets, we achieve a 6.6 points lift of F1 score over the baselines on our video dataset, a 2.9 point lift of F1 score on PASCAL, and a 1.8 points lift of mean Intersection over Union over Grad-CAM for weakly supervised detection on PASCAL. Those improved attention maps may help clinicians better understand vision model predictions and ease the deployment of machine learning systems into clinical care. We share part of the code for this article at the following repository: https://github.com/alimirzazadeh/SemisupervisedAttention.
Contrastive learning-based pretraining improves representation and transferability of diabetic retinopathy classification models
Aug 24, 2022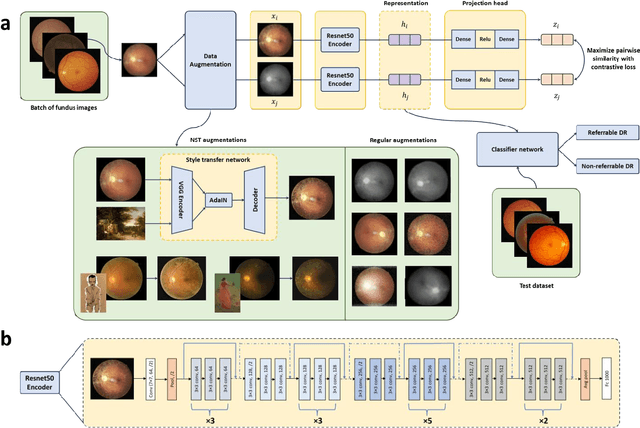
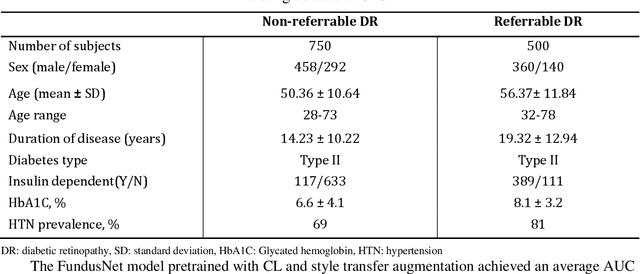

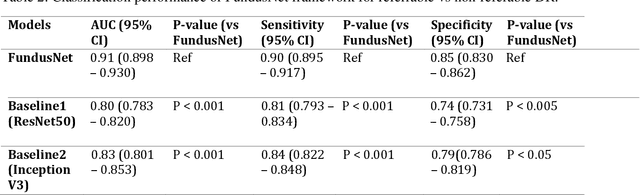
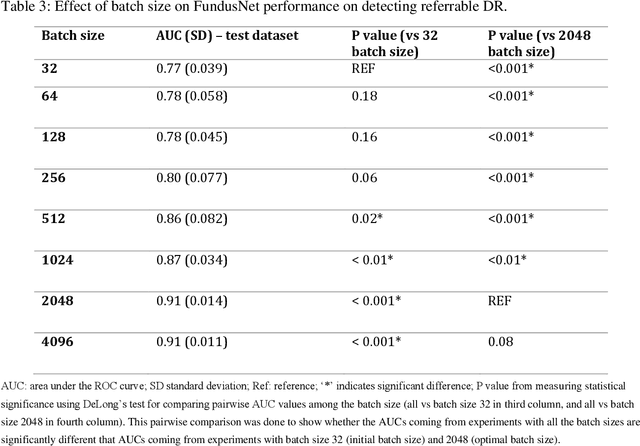
Abstract:Self supervised contrastive learning based pretraining allows development of robust and generalized deep learning models with small, labeled datasets, reducing the burden of label generation. This paper aims to evaluate the effect of CL based pretraining on the performance of referrable vs non referrable diabetic retinopathy (DR) classification. We have developed a CL based framework with neural style transfer (NST) augmentation to produce models with better representations and initializations for the detection of DR in color fundus images. We compare our CL pretrained model performance with two state of the art baseline models pretrained with Imagenet weights. We further investigate the model performance with reduced labeled training data (down to 10 percent) to test the robustness of the model when trained with small, labeled datasets. The model is trained and validated on the EyePACS dataset and tested independently on clinical data from the University of Illinois, Chicago (UIC). Compared to baseline models, our CL pretrained FundusNet model had higher AUC (CI) values (0.91 (0.898 to 0.930) vs 0.80 (0.783 to 0.820) and 0.83 (0.801 to 0.853) on UIC data). At 10 percent labeled training data, the FundusNet AUC was 0.81 (0.78 to 0.84) vs 0.58 (0.56 to 0.64) and 0.63 (0.60 to 0.66) in baseline models, when tested on the UIC dataset. CL based pretraining with NST significantly improves DL classification performance, helps the model generalize well (transferable from EyePACS to UIC data), and allows training with small, annotated datasets, therefore reducing ground truth annotation burden of the clinicians.
The Importance of Background Information for Out of Distribution Generalization
Jun 17, 2022



Abstract:Domain generalization in medical image classification is an important problem for trustworthy machine learning to be deployed in healthcare. We find that existing approaches for domain generalization which utilize ground-truth abnormality segmentations to control feature attributions have poor out-of-distribution (OOD) performance relative to the standard baseline of empirical risk minimization (ERM). We investigate what regions of an image are important for medical image classification and show that parts of the background, that which is not contained in the abnormality segmentation, provides helpful signal. We then develop a new task-specific mask which covers all relevant regions. Utilizing this new segmentation mask significantly improves the performance of the existing methods on the OOD test sets. To obtain better generalization results than ERM, we find it necessary to scale up the training data size in addition to the usage of these task-specific masks.
Label-Efficient Self-Supervised Federated Learning for Tackling Data Heterogeneity in Medical Imaging
May 17, 2022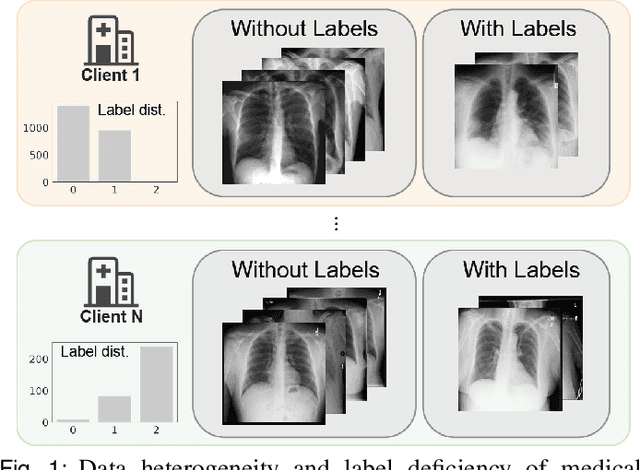

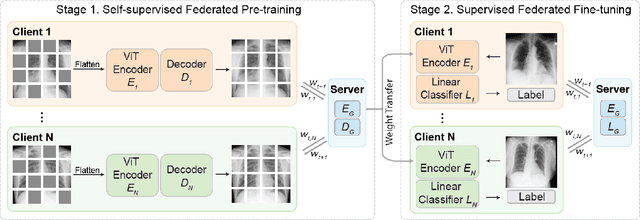
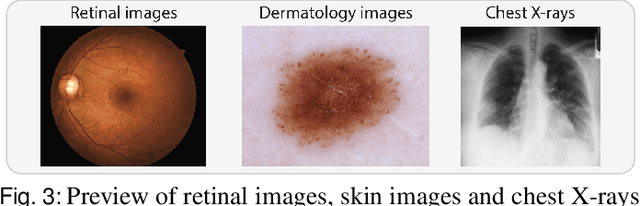
Abstract:The curation of large-scale medical datasets from multiple institutions necessary for training deep learning models is challenged by the difficulty in sharing patient data with privacy-preserving. Federated learning (FL), a paradigm that enables privacy-protected collaborative learning among different institutions, is a promising solution to this challenge. However, FL generally suffers from performance deterioration due to heterogeneous data distributions across institutions and the lack of quality labeled data. In this paper, we present a robust and label-efficient self-supervised FL framework for medical image analysis. Specifically, we introduce a novel distributed self-supervised pre-training paradigm into the existing FL pipeline (i.e., pre-training the models directly on the decentralized target task datasets). Built upon the recent success of Vision Transformers, we employ masked image encoding tasks for self-supervised pre-training, to facilitate more effective knowledge transfer to downstream federated models. Extensive empirical results on simulated and real-world medical imaging federated datasets show that self-supervised pre-training largely benefits the robustness of federated models against various degrees of data heterogeneity. Notably, under severe data heterogeneity, our method, without relying on any additional pre-training data, achieves an improvement of 5.06%, 1.53% and 4.58% in test accuracy on retinal, dermatology and chest X-ray classification compared with the supervised baseline with ImageNet pre-training. Moreover, we show that our self-supervised FL algorithm generalizes well to out-of-distribution data and learns federated models more effectively in limited label scenarios, surpassing the supervised baseline by 10.36% and the semi-supervised FL method by 8.3% in test accuracy.
Masked Co-attentional Transformer reconstructs 100x ultra-fast/low-dose whole-body PET from longitudinal images and anatomically guided MRI
May 09, 2022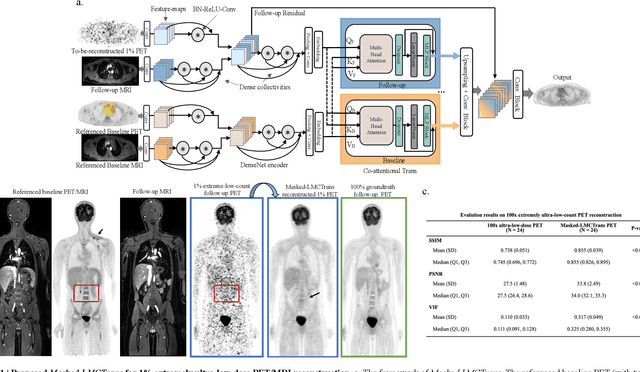
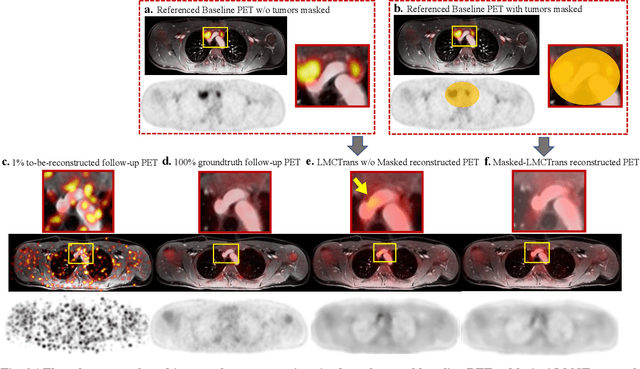
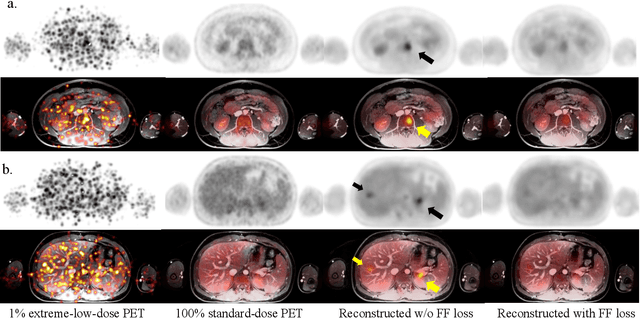
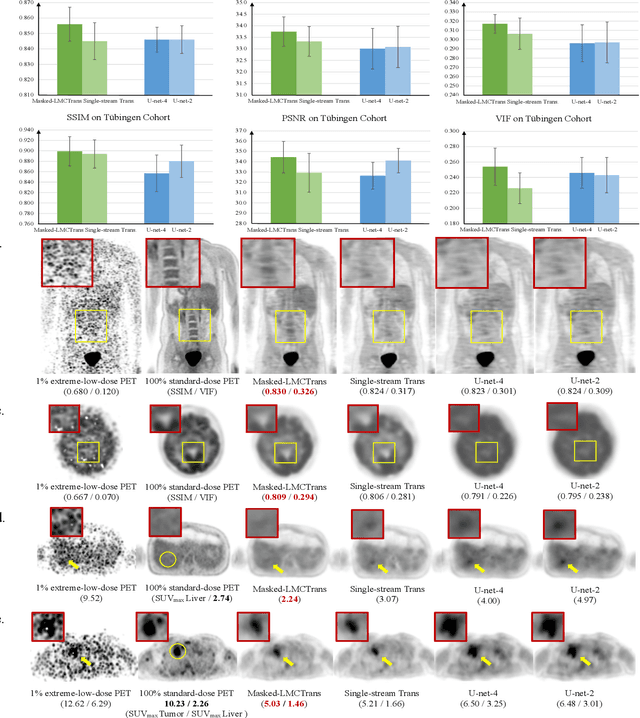
Abstract:Despite its tremendous value for the diagnosis, treatment monitoring and surveillance of children with cancer, whole body staging with positron emission tomography (PET) is time consuming and associated with considerable radiation exposure. 100x (1% of the standard clinical dosage) ultra-low-dose/ultra-fast whole-body PET reconstruction has the potential for cancer imaging with unprecedented speed and improved safety, but it cannot be achieved by the naive use of machine learning techniques. In this study, we utilize the global similarity between baseline and follow-up PET and magnetic resonance (MR) images to develop Masked-LMCTrans, a longitudinal multi-modality co-attentional CNN-Transformer that provides interaction and joint reasoning between serial PET/MRs of the same patient. We mask the tumor area in the referenced baseline PET and reconstruct the follow-up PET scans. In this manner, Masked-LMCTrans reconstructs 100x almost-zero radio-exposure whole-body PET that was not possible before. The technique also opens a new pathway for longitudinal radiology imaging reconstruction, a significantly under-explored area to date. Our model was trained and tested with Stanford PET/MRI scans of pediatric lymphoma patients and evaluated externally on PET/MRI images from T\"ubingen University. The high image quality of the reconstructed 100x whole-body PET images resulting from the application of Masked-LMCTrans will substantially advance the development of safer imaging approaches and shorter exam-durations for pediatric patients, as well as expand the possibilities for frequent longitudinal monitoring of these patients by PET.
Multimodal spatiotemporal graph neural networks for improved prediction of 30-day all-cause hospital readmission
Apr 14, 2022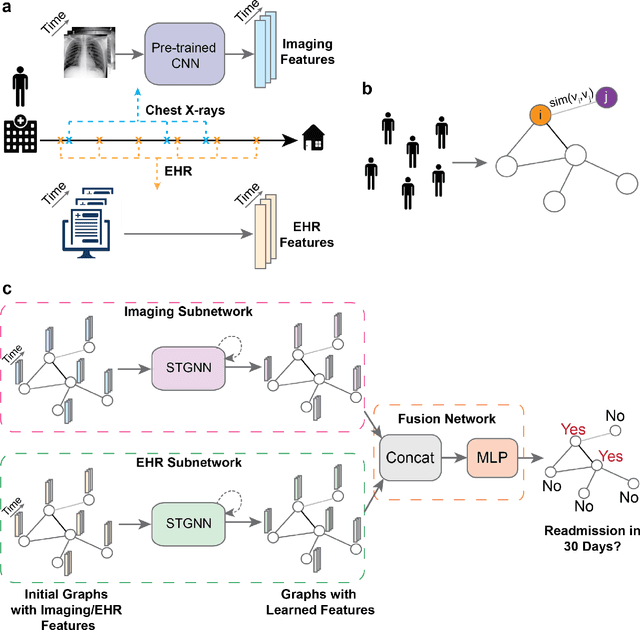
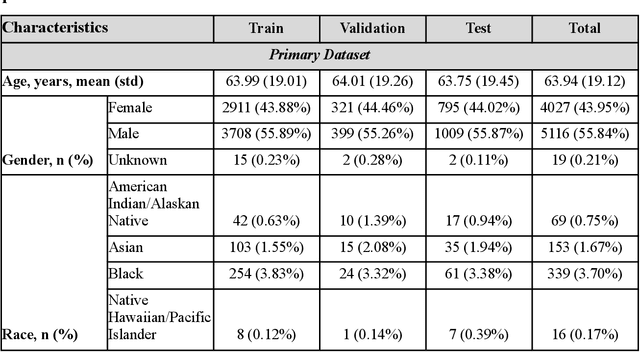
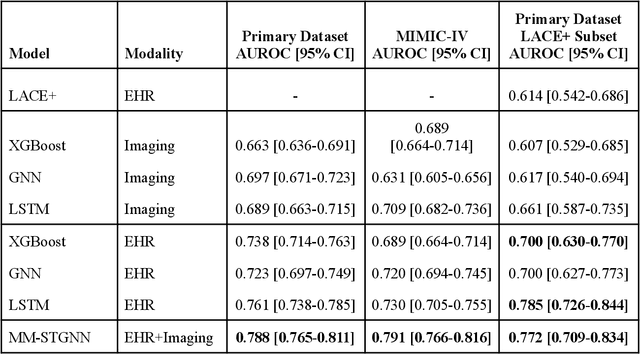
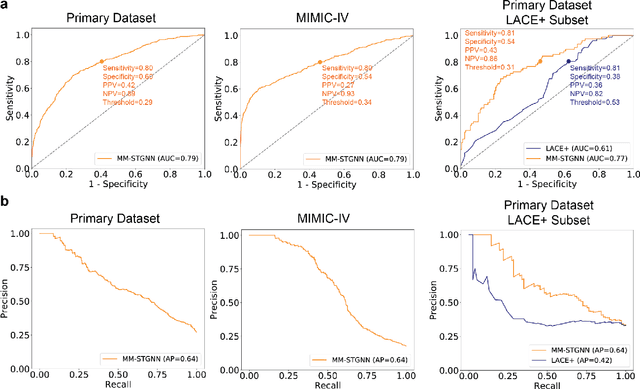
Abstract:Measures to predict 30-day readmission are considered an important quality factor for hospitals as accurate predictions can reduce the overall cost of care by identifying high risk patients before they are discharged. While recent deep learning-based studies have shown promising empirical results on readmission prediction, several limitations exist that may hinder widespread clinical utility, such as (a) only patients with certain conditions are considered, (b) existing approaches do not leverage data temporality, (c) individual admissions are assumed independent of each other, which is unrealistic, (d) prior studies are usually limited to single source of data and single center data. To address these limitations, we propose a multimodal, modality-agnostic spatiotemporal graph neural network (MM-STGNN) for prediction of 30-day all-cause hospital readmission that fuses multimodal in-patient longitudinal data. By training and evaluating our methods using longitudinal chest radiographs and electronic health records from two independent centers, we demonstrate that MM-STGNN achieves AUROC of 0.79 on both primary and external datasets. Furthermore, MM-STGNN significantly outperforms the current clinical reference standard, LACE+ score (AUROC=0.61), on the primary dataset. For subset populations of patients with heart and vascular disease, our model also outperforms baselines on predicting 30-day readmission (e.g., 3.7 point improvement in AUROC in patients with heart disease). Lastly, qualitative model interpretability analysis indicates that while patients' primary diagnoses were not explicitly used to train the model, node features crucial for model prediction directly reflect patients' primary diagnoses. Importantly, our MM-STGNN is agnostic to node feature modalities and could be utilized to integrate multimodal data for triaging patients in various downstream resource allocation tasks.
Automated Detection of Patients in Hospital Video Recordings
Nov 28, 2021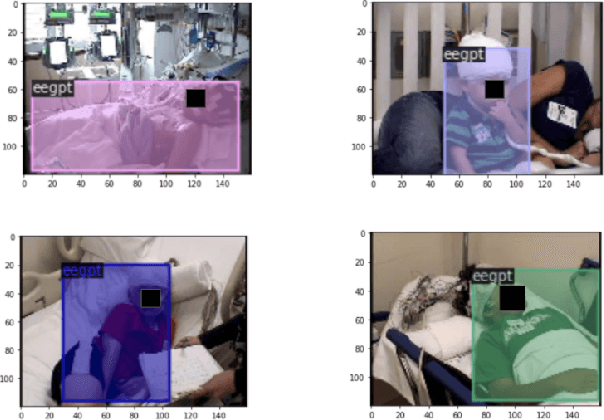
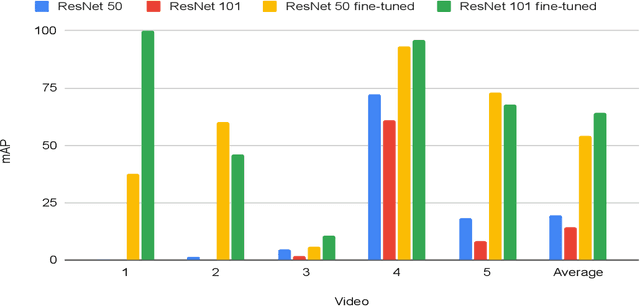
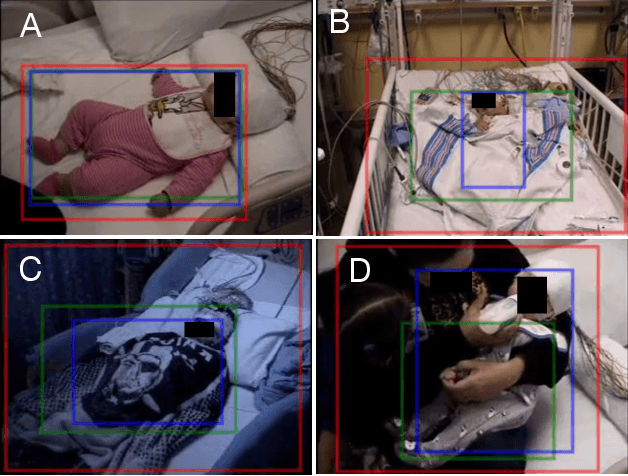
Abstract:In a clinical setting, epilepsy patients are monitored via video electroencephalogram (EEG) tests. A video EEG records what the patient experiences on videotape while an EEG device records their brainwaves. Currently, there are no existing automated methods for tracking the patient's location during a seizure, and video recordings of hospital patients are substantially different from publicly available video benchmark datasets. For example, the camera angle can be unusual, and patients can be partially covered with bedding sheets and electrode sets. Being able to track a patient in real-time with video EEG would be a promising innovation towards improving the quality of healthcare. Specifically, an automated patient detection system could supplement clinical oversight and reduce the resource-intensive efforts of nurses and doctors who need to continuously monitor patients. We evaluate an ImageNet pre-trained Mask R-CNN, a standard deep learning model for object detection, on the task of patient detection using our own curated dataset of 45 videos of hospital patients. The dataset was aggregated and curated for this work. We show that without fine-tuning, ImageNet pre-trained Mask R-CNN models perform poorly on such data. By fine-tuning the models with a subset of our dataset, we observe a substantial improvement in patient detection performance, with a mean average precision of 0.64. We show that the results vary substantially depending on the video clip.
RadFusion: Benchmarking Performance and Fairness for Multimodal Pulmonary Embolism Detection from CT and EHR
Nov 27, 2021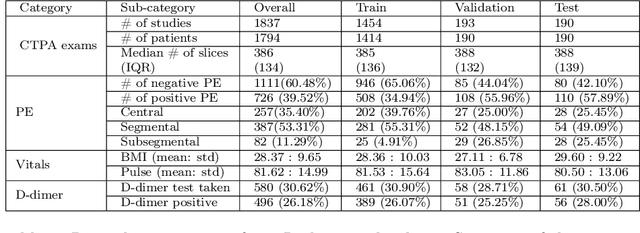
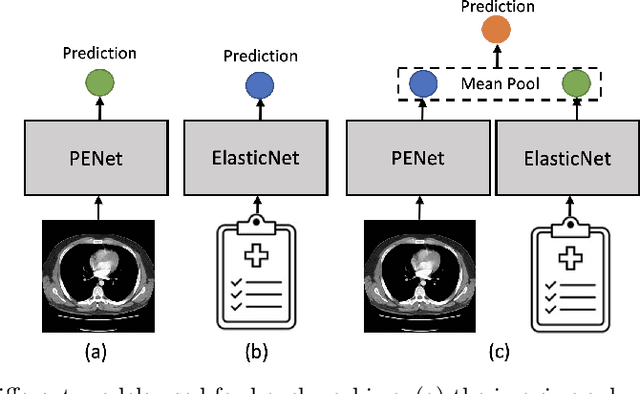
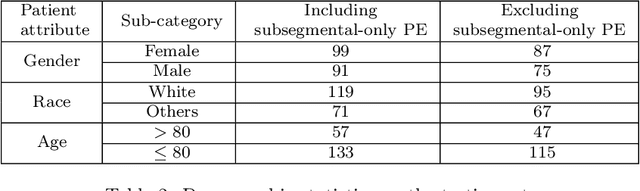
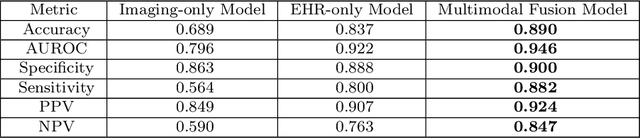
Abstract:Despite the routine use of electronic health record (EHR) data by radiologists to contextualize clinical history and inform image interpretation, the majority of deep learning architectures for medical imaging are unimodal, i.e., they only learn features from pixel-level information. Recent research revealing how race can be recovered from pixel data alone highlights the potential for serious biases in models which fail to account for demographics and other key patient attributes. Yet the lack of imaging datasets which capture clinical context, inclusive of demographics and longitudinal medical history, has left multimodal medical imaging underexplored. To better assess these challenges, we present RadFusion, a multimodal, benchmark dataset of 1794 patients with corresponding EHR data and high-resolution computed tomography (CT) scans labeled for pulmonary embolism. We evaluate several representative multimodal fusion models and benchmark their fairness properties across protected subgroups, e.g., gender, race/ethnicity, age. Our results suggest that integrating imaging and EHR data can improve classification performance and robustness without introducing large disparities in the true positive rate between population groups.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge