Chenxi Zhang
HUAWEI
FinMTM: A Multi-Turn Multimodal Benchmark for Financial Reasoning and Agent Evaluation
Feb 03, 2026Abstract:The financial domain poses substantial challenges for vision-language models (VLMs) due to specialized chart formats and knowledge-intensive reasoning requirements. However, existing financial benchmarks are largely single-turn and rely on a narrow set of question formats, limiting comprehensive evaluation in realistic application scenarios. To address this gap, we propose FinMTM, a multi-turn multimodal benchmark that expands diversity along both data and task dimensions. On the data side, we curate and annotate 11{,}133 bilingual (Chinese and English) financial QA pairs grounded in financial visuals, including candlestick charts, statistical plots, and report figures. On the task side, FinMTM covers single- and multiple-choice questions, multi-turn open-ended dialogues, and agent-based tasks. We further design task-specific evaluation protocols, including a set-overlap scoring rule for multiple-choice questions, a weighted combination of turn-level and session-level scores for multi-turn dialogues, and a composite metric that integrates planning quality with final outcomes for agent tasks. Extensive experimental evaluation of 22 VLMs reveal their limitations in fine-grained visual perception, long-context reasoning, and complex agent workflows.
Retrospective Memory for Camouflaged Object Detection
Jun 18, 2025Abstract:Camouflaged object detection (COD) primarily focuses on learning subtle yet discriminative representations from complex scenes. Existing methods predominantly follow the parametric feedforward architecture based on static visual representation modeling. However, they lack explicit mechanisms for acquiring historical context, limiting their adaptation and effectiveness in handling challenging camouflage scenes. In this paper, we propose a recall-augmented COD architecture, namely RetroMem, which dynamically modulates camouflage pattern perception and inference by integrating relevant historical knowledge into the process. Specifically, RetroMem employs a two-stage training paradigm consisting of a learning stage and a recall stage to construct, update, and utilize memory representations effectively. During the learning stage, we design a dense multi-scale adapter (DMA) to improve the pretrained encoder's capability to capture rich multi-scale visual information with very few trainable parameters, thereby providing foundational inferences. In the recall stage, we propose a dynamic memory mechanism (DMM) and an inference pattern reconstruction (IPR). These components fully leverage the latent relationships between learned knowledge and current sample context to reconstruct the inference of camouflage patterns, thereby significantly improving the model's understanding of camouflage scenes. Extensive experiments on several widely used datasets demonstrate that our RetroMem significantly outperforms existing state-of-the-art methods.
Automatically Planning Optimal Parallel Strategy for Large Language Models
Dec 31, 2024



Abstract:The number of parameters in large-scale language models based on transformers is gradually increasing, and the scale of computing clusters is also growing. The technology of quickly mobilizing large amounts of computing resources for parallel computing is becoming increasingly important. In this paper, we propose an automatic parallel algorithm that automatically plans the parallel strategy with maximum throughput based on model and hardware information. By decoupling the training time into computation, communication, and overlap, we established a training duration simulation model. Based on this simulation model, we prune the parallel solution space to shorten the search time required. The multi-node experiment results show that the algorithm can estimate the parallel training duration in real time with an average accuracy of 96%. In our test, the recommendation strategy provided by the algorithm is always globally optimal.
CGCOD: Class-Guided Camouflaged Object Detection
Dec 25, 2024


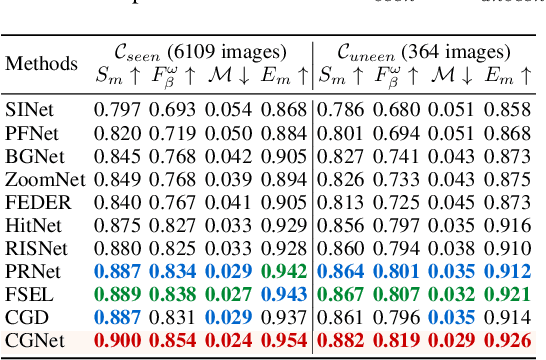
Abstract:Camouflaged Object Detection (COD) is designed to identify objects that blend seamlessly with their surroundings. Due to the complexity of camouflaged objects (such as shape, color, and texture), their semantic cues are often blurred or completely lost, posing a significant challenge for COD. Existing COD methods often rely on visual features, which are not stable enough in changeable camouflage environments. This instability leads to false positives and false negatives, resulting in incomplete or inaccurate segmentation results. In this paper, to solve this problem, we propose a new task, Class-Guided Camouflaged Object Detection (CG-COD), which extends the traditional COD task by introducing object class knowledge, significantly improving the robustness and segmentation accuracy of the model in complex environments. Toward this end, we construct a dataset, CamoClass, containing the camouflaged objects in the real scenes and their corresponding class annotation. Based on this, we propose a multi-stage framework CGNet which consists of a plug-and-play class prompt generator and a class-guided detector. Under the guidance of textual information, CGNet enables efficient segmentation. It is worth emphasizing that for the first time, we extend the object class annotations on existing COD benchmark datasets, and introduce a flexible framework to improve the performance of the existing COD model under text guidance.
SP${ }^3$ : Superpixel-propagated pseudo-label learning for weakly semi-supervised medical image segmentation
Nov 18, 2024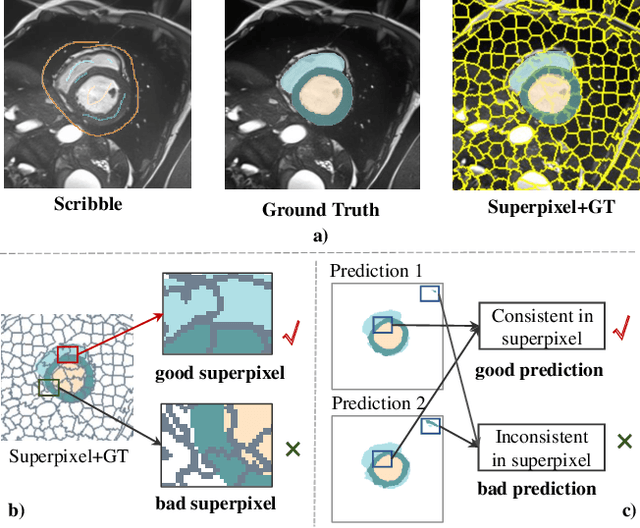



Abstract:Deep learning-based medical image segmentation helps assist diagnosis and accelerate the treatment process while the model training usually requires large-scale dense annotation datasets. Weakly semi-supervised medical image segmentation is an essential application because it only requires a small amount of scribbles and a large number of unlabeled data to train the model, which greatly reduces the clinician's effort to fully annotate images. To handle the inadequate supervisory information challenge in weakly semi-supervised segmentation (WSSS), a SuperPixel-Propagated Pseudo-label (SP${}^3$) learning method is proposed, using the structural information contained in superpixel for supplemental information. Specifically, the annotation of scribbles is propagated to superpixels and thus obtains a dense annotation for supervised training. Since the quality of pseudo-labels is limited by the low-quality annotation, the beneficial superpixels selected by dynamic thresholding are used to refine pseudo-labels. Furthermore, aiming to alleviate the negative impact of noise in pseudo-label, superpixel-level uncertainty is incorporated to guide the pseudo-label supervision for stable learning. Our method achieves state-of-the-art performance on both tumor and organ segmentation datasets under the WSSS setting, using only 3\% of the annotation workload compared to fully supervised methods and attaining approximately 80\% Dice score. Additionally, our method outperforms eight weakly and semi-supervised methods under both weakly supervised and semi-supervised settings. Results of extensive experiments validate the effectiveness and annotation efficiency of our weakly semi-supervised segmentation, which can assist clinicians in achieving automated segmentation for organs or tumors quickly and ultimately benefit patients.
Towards Arbitrary-Scale Histopathology Image Super-resolution: An Efficient Dual-branch Framework via Implicit Self-texture Enhancement
Jan 28, 2024Abstract:High-quality whole-slide scanners are expensive, complex, and time-consuming, thus limiting the acquisition and utilization of high-resolution pathology whole-slide images in daily clinical work. Deep learning-based single-image super-resolution techniques are an effective way to solve this problem by synthesizing high-resolution images from low-resolution ones. However, the existing super-resolution models applied in pathology images can only work in fixed integer magnifications, significantly decreasing their applicability. Though methods based on implicit neural representation have shown promising results in arbitrary-scale super-resolution of natural images, applying them directly to pathology images is inadequate because they have unique fine-grained image textures different from natural images. Thus, we propose an Implicit Self-Texture Enhancement-based dual-branch framework (ISTE) for arbitrary-scale super-resolution of pathology images to address this challenge. ISTE contains a pixel learning branch and a texture learning branch, which first learn pixel features and texture features, respectively. Then, we design a two-stage texture enhancement strategy to fuse the features from the two branches to obtain the super-resolution results, where the first stage is feature-based texture enhancement, and the second stage is spatial-domain-based texture enhancement. Extensive experiments on three public datasets show that ISTE outperforms existing fixed-scale and arbitrary-scale algorithms at multiple magnifications and helps to improve downstream task performance. To the best of our knowledge, this is the first work to achieve arbitrary-scale super-resolution in pathology images. Codes will be available.
O-PRESS: Boosting OCT axial resolution with Prior guidance, Recurrence, and Equivariant Self-Supervision
Jan 06, 2024



Abstract:Optical coherence tomography (OCT) is a noninvasive technology that enables real-time imaging of tissue microanatomies. The axial resolution of OCT is intrinsically constrained by the spectral bandwidth of the employed light source while maintaining a fixed center wavelength for a specific application. Physically extending this bandwidth faces strong limitations and requires a substantial cost. We present a novel computational approach, called as O-PRESS, for boosting the axial resolution of OCT with Prior Guidance, a Recurrent mechanism, and Equivariant Self-Supervision. Diverging from conventional superresolution methods that rely on physical models or data-driven techniques, our method seamlessly integrates OCT modeling and deep learning, enabling us to achieve real-time axial-resolution enhancement exclusively from measurements without a need for paired images. Our approach solves two primary tasks of resolution enhancement and noise reduction with one treatment. Both tasks are executed in a self-supervised manner, with equivariance imaging and free space priors guiding their respective processes. Experimental evaluations, encompassing both quantitative metrics and visual assessments, consistently verify the efficacy and superiority of our approach, which exhibits performance on par with fully supervised methods. Importantly, the robustness of our model is affirmed, showcasing its dual capability to enhance axial resolution while concurrently improving the signal-to-noise ratio.
keqing: knowledge-based question answering is a nature chain-of-thought mentor of LLM
Dec 31, 2023



Abstract:Large language models (LLMs) have exhibited remarkable performance on various natural language processing (NLP) tasks, especially for question answering. However, in the face of problems beyond the scope of knowledge, these LLMs tend to talk nonsense with a straight face, where the potential solution could be incorporating an Information Retrieval (IR) module and generating response based on these retrieved knowledge. In this paper, we present a novel framework to assist LLMs, such as ChatGPT, to retrieve question-related structured information on the knowledge graph, and demonstrate that Knowledge-based question answering (Keqing) could be a nature Chain-of-Thought (CoT) mentor to guide the LLM to sequentially find the answer entities of a complex question through interpretable logical chains. Specifically, the workflow of Keqing will execute decomposing a complex question according to predefined templates, retrieving candidate entities on knowledge graph, reasoning answers of sub-questions, and finally generating response with reasoning paths, which greatly improves the reliability of LLM's response. The experimental results on KBQA datasets show that Keqing can achieve competitive performance and illustrate the logic of answering each question.
Multi-organ segmentation: a progressive exploration of learning paradigms under scarce annotation
Feb 09, 2023Abstract:Precise delineation of multiple organs or abnormal regions in the human body from medical images plays an essential role in computer-aided diagnosis, surgical simulation, image-guided interventions, and especially in radiotherapy treatment planning. Thus, it is of great significance to explore automatic segmentation approaches, among which deep learning-based approaches have evolved rapidly and witnessed remarkable progress in multi-organ segmentation. However, obtaining an appropriately sized and fine-grained annotated dataset of multiple organs is extremely hard and expensive. Such scarce annotation limits the development of high-performance multi-organ segmentation models but promotes many annotation-efficient learning paradigms. Among these, studies on transfer learning leveraging external datasets, semi-supervised learning using unannotated datasets and partially-supervised learning integrating partially-labeled datasets have led the dominant way to break such dilemma in multi-organ segmentation. We first review the traditional fully supervised method, then present a comprehensive and systematic elaboration of the 3 abovementioned learning paradigms in the context of multi-organ segmentation from both technical and methodological perspectives, and finally summarize their challenges and future trends.
Robust Point Cloud Registration Framework Based on Deep Graph Matching(TPAMI Version)
Nov 09, 2022



Abstract:3D point cloud registration is a fundamental problem in computer vision and robotics. Recently, learning-based point cloud registration methods have made great progress. However, these methods are sensitive to outliers, which lead to more incorrect correspondences. In this paper, we propose a novel deep graph matching-based framework for point cloud registration. Specifically, we first transform point clouds into graphs and extract deep features for each point. Then, we develop a module based on deep graph matching to calculate a soft correspondence matrix. By using graph matching, not only the local geometry of each point but also its structure and topology in a larger range are considered in establishing correspondences, so that more correct correspondences are found. We train the network with a loss directly defined on the correspondences, and in the test stage the soft correspondences are transformed into hard one-to-one correspondences so that registration can be performed by a correspondence-based solver. Furthermore, we introduce a transformer-based method to generate edges for graph construction, which further improves the quality of the correspondences. Extensive experiments on object-level and scene-level benchmark datasets show that the proposed method achieves state-of-the-art performance. The code is available at: \href{https://github.com/fukexue/RGM}{https://github.com/fukexue/RGM}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge