Leo Anthony Celi
Laboratory for Computational Physiology, Massachusetts Institute of Technology, Cambridge, MA, USA, Division of Pulmonary, Critical Care and Sleep Medicine, Beth Israel Deaconess Medical Center, Boston, MA, USA, Department of Biostatistics, Harvard T.H. Chan School of Public Health, Boston, MA, USA
Quantum Kernel Advantage over Classical Collapse in Medical Foundation Model Embeddings
Apr 27, 2026Abstract:We provide evidence of quantum kernel advantage under noiseless simulation in binary insurance classification on MIMIC-CXR chest radiographs using quantum support vector machines (QSVM) with frozen embeddings from three medical foundation models (MedSigLIP-448, RAD-DINO, ViT-patch32). We propose a two-tier fair comparison framework in which both classifiers receive identical PCA-q features. At Tier 1 (untuned QSVM vs. untuned linear SVM, C = 1 both sides), QSVM wins minority-class F1 in all 18 tested configurations (17 at p < 0.001, 1 at p < 0.01). The classical linear kernel collapses to majority-class prediction on 90-100% of seeds at every qubit count, while QSVM maintains non-trivial recall. At q = 11 (MedSigLIP-448 plateau center), QSVM achieves mean F1 = 0.343 vs. classical F1 = 0.050 (F1 gain = +0.293, p < 0.001) without hyperparameter tuning. Under Tier 2 (untuned QSVM vs. C-tuned RBF SVM), QSVM wins all seven tested configurations (mean gain +0.068, max +0.112). Eigenspectrum analysis reveals quantum kernel effective rank reaches 69.80 at q = 11, far exceeding linear kernel rank, while classical collapse remains C-invariant. A full qubit sweep reveals architecture-dependent concentration onset across models. Code: https://github.com/sebasmos/qml-medimage
Surrogate modeling for interpreting black-box LLMs in medical predictions
Apr 22, 2026Abstract:Large language models (LLMs), trained on vast datasets, encode extensive real-world knowledge within their parameters, yet their black-box nature obscures the mechanisms and extent of this encoding. Surrogate modeling, which uses simplified models to approximate complex systems, can offer a path toward better interpretability of black-box models. We propose a surrogate modeling framework that quantitatively explains LLM-encoded knowledge. For a specific hypothesis derived from domain knowledge, this framework approximates the latent LLM knowledge space using observable elements (input-output pairs) through extensive prompting across a comprehensive range of simulated scenarios. Through proof-of-concept experiments in medical predictions, we demonstrate our framework's effectiveness in revealing the extent to which LLMs "perceive" each input variable in relation to the output. Particularly, given concerns that LLMs may perpetuate inaccuracies and societal biases embedded in their training data, our experiments using this framework quantitatively revealed both associations that contradict established medical knowledge and the persistence of scientifically refuted racial assumptions within LLM-encoded knowledge. By disclosing these issues, our framework can act as a red-flag indicator to support the safe and reliable application of these models.
Domain-Specific Latent Representations Improve the Fidelity of Diffusion-Based Medical Image Super-Resolution
Apr 14, 2026Abstract:Latent diffusion models for medical image super-resolution universally inherit variational autoencoders designed for natural photographs. We show that this default choice, not the diffusion architecture, is the dominant constraint on reconstruction quality. In a controlled experiment holding all other pipeline components fixed, replacing the generic Stable Diffusion VAE with MedVAE, a domain-specific autoencoder pretrained on more than 1.6 million medical images, yields +2.91 to +3.29 dB PSNR improvement across knee MRI, brain MRI, and chest X-ray (n = 1,820; Cohen's d = 1.37 to 1.86, all p < 10^{-20}, Wilcoxon signed-rank). Wavelet decomposition localises the advantage to the finest spatial frequency bands encoding anatomically relevant fine structure. Ablations across inference schedules, prediction targets, and generative architectures confirm the gap is stable within plus or minus 0.15 dB, while hallucination rates remain comparable between methods (Cohen's h < 0.02 across all datasets), establishing that reconstruction fidelity and generative hallucination are governed by independent pipeline components. These results provide a practical screening criterion: autoencoder reconstruction quality, measurable without diffusion training, predicts downstream SR performance (R^2 = 0.67), suggesting that domain-specific VAE selection should precede diffusion architecture search. Code and trained model weights are publicly available at https://github.com/sebasmos/latent-sr.
Learning Representations from Incomplete EHR Data with Dual-Masked Autoencoding
Feb 16, 2026Abstract:Learning from electronic health records (EHRs) time series is challenging due to irregular sam- pling, heterogeneous missingness, and the resulting sparsity of observations. Prior self-supervised meth- ods either impute before learning, represent missingness through a dedicated input signal, or optimize solely for imputation, reducing their capacity to efficiently learn representations that support clinical downstream tasks. We propose the Augmented-Intrinsic Dual-Masked Autoencoder (AID-MAE), which learns directly from incomplete time series by applying an intrinsic missing mask to represent naturally missing values and an augmented mask that hides a subset of observed values for reconstruction during training. AID-MAE processes only the unmasked subset of tokens and consistently outperforms strong baselines, including XGBoost and DuETT, across multiple clinical tasks on two datasets. In addition, the learned embeddings naturally stratify patient cohorts in the representation space.
Uncertainty Makes It Stable: Curiosity-Driven Quantized Mixture-of-Experts
Nov 19, 2025Abstract:Deploying deep neural networks on resource-constrained devices faces two critical challenges: maintaining accuracy under aggressive quantization while ensuring predictable inference latency. We present a curiosity-driven quantized Mixture-of-Experts framework that addresses both through Bayesian epistemic uncertainty-based routing across heterogeneous experts (BitNet ternary, 1-16 bit BitLinear, post-training quantization). Evaluated on audio classification benchmarks (ESC-50, Quinn, UrbanSound8K), our 4-bit quantization maintains 99.9 percent of 16-bit accuracy (0.858 vs 0.859 F1) with 4x compression and 41 percent energy savings versus 8-bit. Crucially, curiosity-driven routing reduces MoE latency variance by 82 percent (p = 0.008, Levene's test) from 230 ms to 29 ms standard deviation, enabling stable inference for battery-constrained devices. Statistical analysis confirms 4-bit/8-bit achieve practical equivalence with full precision (p > 0.05), while MoE architectures introduce 11 percent latency overhead (p < 0.001) without accuracy gains. At scale, deployment emissions dominate training by 10000x for models serving more than 1,000 inferences, making inference efficiency critical. Our information-theoretic routing demonstrates that adaptive quantization yields accurate (0.858 F1, 1.2M params), energy-efficient (3.87 F1/mJ), and predictable edge models, with simple 4-bit quantized architectures outperforming complex MoE for most deployments.
Algorithms Trained on Normal Chest X-rays Can Predict Health Insurance Types
Nov 17, 2025Abstract:Artificial intelligence is revealing what medicine never intended to encode. Deep vision models, trained on chest X-rays, can now detect not only disease but also invisible traces of social inequality. In this study, we show that state-of-the-art architectures (DenseNet121, SwinV2-B, MedMamba) can predict a patient's health insurance type, a strong proxy for socioeconomic status, from normal chest X-rays with significant accuracy (AUC around 0.67 on MIMIC-CXR-JPG, 0.68 on CheXpert). The signal persists even when age, race, and sex are controlled for, and remains detectable when the model is trained exclusively on a single racial group. Patch-based occlusion reveals that the signal is diffuse rather than localized, embedded in the upper and mid-thoracic regions. This suggests that deep networks may be internalizing subtle traces of clinical environments, equipment differences, or care pathways; learning socioeconomic segregation itself. These findings challenge the assumption that medical images are neutral biological data. By uncovering how models perceive and exploit these hidden social signatures, this work reframes fairness in medical AI: the goal is no longer only to balance datasets or adjust thresholds, but to interrogate and disentangle the social fingerprints embedded in clinical data itself.
CXR-LT 2024: A MICCAI challenge on long-tailed, multi-label, and zero-shot disease classification from chest X-ray
Jun 09, 2025Abstract:The CXR-LT series is a community-driven initiative designed to enhance lung disease classification using chest X-rays (CXR). It tackles challenges in open long-tailed lung disease classification and enhances the measurability of state-of-the-art techniques. The first event, CXR-LT 2023, aimed to achieve these goals by providing high-quality benchmark CXR data for model development and conducting comprehensive evaluations to identify ongoing issues impacting lung disease classification performance. Building on the success of CXR-LT 2023, the CXR-LT 2024 expands the dataset to 377,110 chest X-rays (CXRs) and 45 disease labels, including 19 new rare disease findings. It also introduces a new focus on zero-shot learning to address limitations identified in the previous event. Specifically, CXR-LT 2024 features three tasks: (i) long-tailed classification on a large, noisy test set, (ii) long-tailed classification on a manually annotated "gold standard" subset, and (iii) zero-shot generalization to five previously unseen disease findings. This paper provides an overview of CXR-LT 2024, detailing the data curation process and consolidating state-of-the-art solutions, including the use of multimodal models for rare disease detection, advanced generative approaches to handle noisy labels, and zero-shot learning strategies for unseen diseases. Additionally, the expanded dataset enhances disease coverage to better represent real-world clinical settings, offering a valuable resource for future research. By synthesizing the insights and innovations of participating teams, we aim to advance the development of clinically realistic and generalizable diagnostic models for chest radiography.
Performance Gains of LLMs With Humans in a World of LLMs Versus Humans
May 13, 2025



Abstract:Currently, a considerable research effort is devoted to comparing LLMs to a group of human experts, where the term "expert" is often ill-defined or variable, at best, in a state of constantly updating LLM releases. Without proper safeguards in place, LLMs will threaten to cause harm to the established structure of safe delivery of patient care which has been carefully developed throughout history to keep the safety of the patient at the forefront. A key driver of LLM innovation is founded on community research efforts which, if continuing to operate under "humans versus LLMs" principles, will expedite this trend. Therefore, research efforts moving forward must focus on effectively characterizing the safe use of LLMs in clinical settings that persist across the rapid development of novel LLM models. In this communication, we demonstrate that rather than comparing LLMs to humans, there is a need to develop strategies enabling efficient work of humans with LLMs in an almost symbiotic manner.
BRIDGE: Benchmarking Large Language Models for Understanding Real-world Clinical Practice Text
May 01, 2025



Abstract:Large language models (LLMs) hold great promise for medical applications and are evolving rapidly, with new models being released at an accelerated pace. However, current evaluations of LLMs in clinical contexts remain limited. Most existing benchmarks rely on medical exam-style questions or PubMed-derived text, failing to capture the complexity of real-world electronic health record (EHR) data. Others focus narrowly on specific application scenarios, limiting their generalizability across broader clinical use. To address this gap, we present BRIDGE, a comprehensive multilingual benchmark comprising 87 tasks sourced from real-world clinical data sources across nine languages. We systematically evaluated 52 state-of-the-art LLMs (including DeepSeek-R1, GPT-4o, Gemini, and Llama 4) under various inference strategies. With a total of 13,572 experiments, our results reveal substantial performance variation across model sizes, languages, natural language processing tasks, and clinical specialties. Notably, we demonstrate that open-source LLMs can achieve performance comparable to proprietary models, while medically fine-tuned LLMs based on older architectures often underperform versus updated general-purpose models. The BRIDGE and its corresponding leaderboard serve as a foundational resource and a unique reference for the development and evaluation of new LLMs in real-world clinical text understanding. The BRIDGE leaderboard: https://huggingface.co/spaces/YLab-Open/BRIDGE-Medical-Leaderboard
An Algorithmic Approach for Causal Health Equity: A Look at Race Differentials in Intensive Care Unit (ICU) Outcomes
Jan 09, 2025
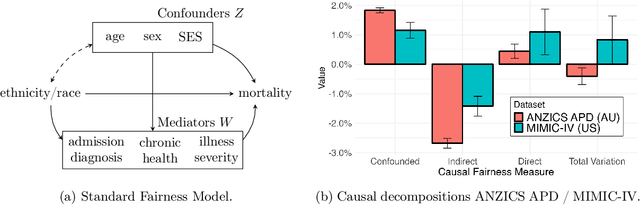
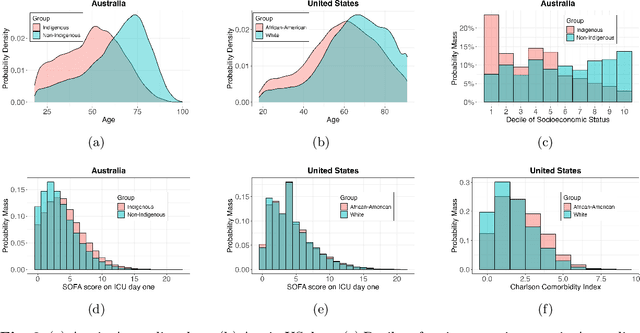

Abstract:The new era of large-scale data collection and analysis presents an opportunity for diagnosing and understanding the causes of health inequities. In this study, we describe a framework for systematically analyzing health disparities using causal inference. The framework is illustrated by investigating racial and ethnic disparities in intensive care unit (ICU) outcome between majority and minority groups in Australia (Indigenous vs. Non-Indigenous) and the United States (African-American vs. White). We demonstrate that commonly used statistical measures for quantifying inequity are insufficient, and focus on attributing the observed disparity to the causal mechanisms that generate it. We find that minority patients are younger at admission, have worse chronic health, are more likely to be admitted for urgent and non-elective reasons, and have higher illness severity. At the same time, however, we find a protective direct effect of belonging to a minority group, with minority patients showing improved survival compared to their majority counterparts, with all other variables kept equal. We demonstrate that this protective effect is related to the increased probability of being admitted to ICU, with minority patients having an increased risk of ICU admission. We also find that minority patients, while showing improved survival, are more likely to be readmitted to ICU. Thus, due to worse access to primary health care, minority patients are more likely to end up in ICU for preventable conditions, causing a reduction in the mortality rates and creating an effect that appears to be protective. Since the baseline risk of ICU admission may serve as proxy for lack of access to primary care, we developed the Indigenous Intensive Care Equity (IICE) Radar, a monitoring system for tracking the over-utilization of ICU resources by the Indigenous population of Australia across geographical areas.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge